7 New Insights about the Frontrunner U.S. Vaccine Candidate
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
A vaccine from Moderna could be available as early as January, Fauci recently said.
Earlier this year, biotech company Moderna broke world records for speed in vaccine development. Their researchers translated the genetic code of the coronavirus into a vaccine candidate in just 42 days.
We're about to expand our safety data in Phase II.
Phase I of the clinical trial started in Seattle on March 16th, with the already-iconic image of volunteer Jennifer Haller calmly receiving the very first dose.
Instead of traditional methods, this vaccine uses a new -- and so far unproven -- technology based on synthetic biology: It hijacks the software of life – messenger RNA – to deliver a copy of the virus's genetic sequence into cells, which, in theory, triggers the body to produce antibodies to fight off a coronavirus infection.
U.S. National Institute of Allergy and Infectious Diseases Director Anthony Fauci called the vaccine's preclinical data "impressive" and told National Geographic this week that a vaccine could be ready for general use as early as January.
The Phase I trial has dosed 45 healthy adults. Phase II trials are about to start, enrolling around 600 adults. Pivotal efficacy trials would follow soon thereafter, bankrolled in collaboration with the government office BARDA (Biomedical Advanced Research and Development Authority).
Today, the chief medical officer of Moderna, Tal Zaks, answered burning questions from the public in a webinar hosted by STAT. Here's an edited and condensed summary of his answers.
1) When will a vaccine become available?
We expect to have data in early summer about the antibody levels from our mRNA vaccine. At the same time, we can measure the antibody levels of people who have had the disease, and we should be able to measure the ability of those antibodies to prevent disease.
We will not yet know if the mRNA vaccine works to prevent disease, but we could soon talk about a potential for benefit. We don't yet know about risk. We're about to expand our safety data in Phase II.
In the summer, there is an expectation that we will be launching pivotal trials, in collaboration with government agencies that are helping fund the research. The trials would be launched with the vaccine vs. a placebo with the goal of establishing: How many cases can we show we prevented with the vaccine?
This is determined by two factors: How big is the trial? And what's the attack rate in the population we vaccinate? The challenge will be to vaccinate in the areas where the risk of infection is still high in the coming months, and we're able to vaccinate and demonstrate fewer infections compared to a placebo. If the disease is happening faster in a given area, you will be able to see an outcome faster. Potentially by the end of the year, we will have the data to say if the vaccine works.
Will that be enough for regulatory approval? The main question is: When will we cross the threshold for the anticipated benefit of a presumed vaccine to be worth the risk?
There is a distinction between approval for those who need it most, like the elderly. Their unmet need and risk/benefit is not the same as it is for younger adults.
My private opinion: I don't think it's a one-size-fits-all. It will be a more measured stance.
2) Can you speed up the testing process with challenge studies, where volunteers willingly get infected?
It's a great question and I applaud the people who ask it and I applaud those signing up to do it. I'm not sure I am a huge fan, for both practical and ethical reasons. The devil is in the details. A challenge study has to show us a vaccine can prevent not just infection but prevent disease. Otherwise, how do I know the dose in the challenge study is the right dose? If you take 100 young people, 90 of them will get mild or no disease. Ten may end up in hospital and one in the ICU.
Also, the timeline. Can it let you skip Phase II of large efficacy trial? The reality for us is that we are about to start Phase II anyway. It would be months before a challenge trial could be designed. And ethically: everybody agrees there is a risk that is not zero of having very serious disease. To justify the risk, we have to be sure the benefit is worth it - that it actually shrunk the timeline. To just give us another data point, I find it hard to accept.
This technology allows us to scale up manufacturing and production.
3) What was seen preclinically in the animal models with Moderna's mRNA vaccines?
We have taken vaccines using our technology against eight different viruses, including two flu strains. In every case, in the preclinical model, we showed we could prevent disease, and when we got to antibody levels, we got the data we wanted to see. In doses of 25-100 micrograms, that usually ends up being a sweet spot where we see an effect. It's a good place as to the expectation of what we will see in Phase I trials.
4) Why is Moderna pursuing an mRNA virus instead of a traditional inactivated virus or recombinant one? This is an untried technology.
First, speed matters in a pandemic. If you have tech that can move much quicker, that makes a difference. The reason we have broken world records is that we have invested time and effort to be ready. We're starting from a platform where it's all based on synthetic biology.
Second, it's fundamental biology - we do not need to make an elaborate vaccine or stick a new virus in an old virus, or try to make a neutralizing but not binding virus. Our technology is basically mimicking the virus. All life works on making proteins through RNA. We have a biological advantage by teaching the immune system to do the right thing.
Third, this technology allows us to scale up manufacturing and production. We as a company have always seen this ahead of us. We invested in our own manufacturing facility two years ago. We have already envisioned scale up on two dimensions. Lot size and vaccines. Vaccines is the easier piece of it. If everybody gets 100 micrograms, it's not a heck of a lot. Prior to COVID, our lead program was a CMV (Cytomegalovirus) vaccine. We had envisioned launching Phase III next year. We had been already well on the path to scale up when COVID-19 caught us by surprise. This would be millions and millions of doses, but the train tracks have been laid.
5) People tend to think of vaccines as an on-off switch -- you get a vaccine and you're protected. But efficacy can be low or high (like the flu vs. measles vaccines). How good is good enough here for protection, and could we need several doses?
Probably around 50-60 percent efficacy is good enough for preventing a significant amount of disease and decreasing the R0. We will aim higher, but it's hard to estimate what degree of efficacy to prepare for until we do the trial. (For comparison, the average flu vaccine efficacy is around 50 percent.)
We anticipate a prime boost. If our immune system has never seen a virus, you can show you're getting to a certain antibody level and then remind the immune system (with another dose). A prime boost is optimal.
My only two competitors are the virus and the clock.
6) How would mutations affect a vaccine?
Coronaviruses tend to mutate the least compared to other viruses but it's entirely possible that it mutates. The report this week about those projected mutations on the spike protein have not been predicted to alter the critical antibodies.
As we scale up manufacturing, the ability to plug in a new genetic sequence and get a new vaccine out there will be very rapid.
For flu vaccine, we don't prove efficacy every year. If we get to the same place with an mRNA vaccine, we will just change the sequence and come out with a new vaccine. The path to approval would be much faster if we leverage the totality of efficacy data like we do for flu.
7) Will there be more than one vaccine and how will they be made available?
I hope so, I don't know. The path to making these available will go through a public-private partnership. It's not your typical commercial way of deploying a vaccine. But my only two competitors are the virus and the clock. We need everybody to be successful.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
DNA- and RNA-based electronic implants may revolutionize healthcare
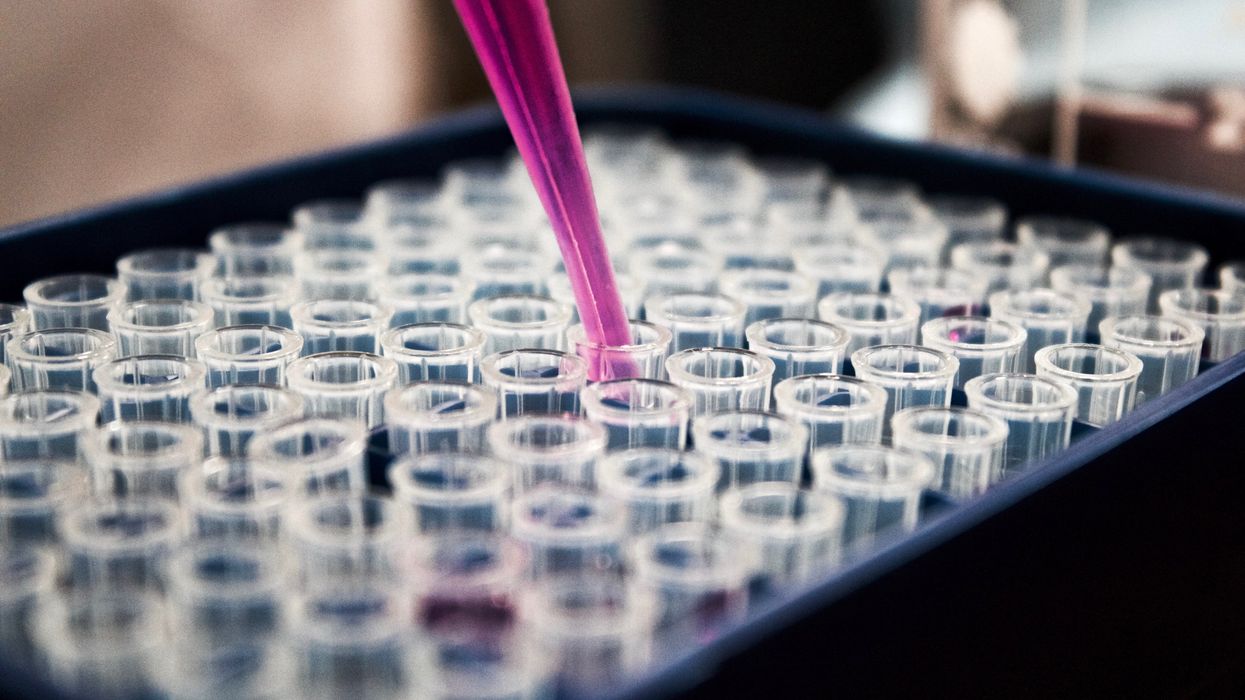
The test tubes contain tiny DNA/enzyme-based circuits, which comprise TRUMPET, a new type of electronic device, smaller than a cell.
Implantable electronic devices can significantly improve patients’ quality of life. A pacemaker can encourage the heart to beat more regularly. A neural implant, usually placed at the back of the skull, can help brain function and encourage higher neural activity. Current research on neural implants finds them helpful to patients with Parkinson’s disease, vision loss, hearing loss, and other nerve damage problems. Several of these implants, such as Elon Musk’s Neuralink, have already been approved by the FDA for human use.
Yet, pacemakers, neural implants, and other such electronic devices are not without problems. They require constant electricity, limited through batteries that need replacements. They also cause scarring. “The problem with doing this with electronics is that scar tissue forms,” explains Kate Adamala, an assistant professor of cell biology at the University of Minnesota Twin Cities. “Anytime you have something hard interacting with something soft [like muscle, skin, or tissue], the soft thing will scar. That's why there are no long-term neural implants right now.” To overcome these challenges, scientists are turning to biocomputing processes that use organic materials like DNA and RNA. Other promised benefits include “diagnostics and possibly therapeutic action, operating as nanorobots in living organisms,” writes Evgeny Katz, a professor of bioelectronics at Clarkson University, in his book DNA- And RNA-Based Computing Systems.
While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output.
Adamala’s research focuses on developing such biocomputing systems using DNA, RNA, proteins, and lipids. Using these molecules in the biocomputing systems allows the latter to be biocompatible with the human body, resulting in a natural healing process. In a recent Nature Communications study, Adamala and her team created a new biocomputing platform called TRUMPET (Transcriptional RNA Universal Multi-Purpose GatE PlaTform) which acts like a DNA-powered computer chip. “These biological systems can heal if you design them correctly,” adds Adamala. “So you can imagine a computer that will eventually heal itself.”
The basics of biocomputing
Biocomputing and regular computing have many similarities. Like regular computing, biocomputing works by running information through a series of gates, usually logic gates. A logic gate works as a fork in the road for an electronic circuit. The input will travel one way or another, giving two different outputs. An example logic gate is the AND gate, which has two inputs (A and B) and two different results. If both A and B are 1, the AND gate output will be 1. If only A is 1 and B is 0, the output will be 0 and vice versa. If both A and B are 0, the result will be 0. While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output. In this case, the DNA enters the logic gate as a single or double strand.
If the DNA is double-stranded, the system “digests” the DNA or destroys it, which results in non-fluorescence or “0” output. Conversely, if the DNA is single-stranded, it won’t be digested and instead will be copied by several enzymes in the biocomputing system, resulting in fluorescent RNA or a “1” output. And the output for this type of binary system can be expanded beyond fluorescence or not. For example, a “1” output might be the production of the enzyme insulin, while a “0” may be that no insulin is produced. “This kind of synergy between biology and computation is the essence of biocomputing,” says Stephanie Forrest, a professor and the director of the Biodesign Center for Biocomputing, Security and Society at Arizona State University.
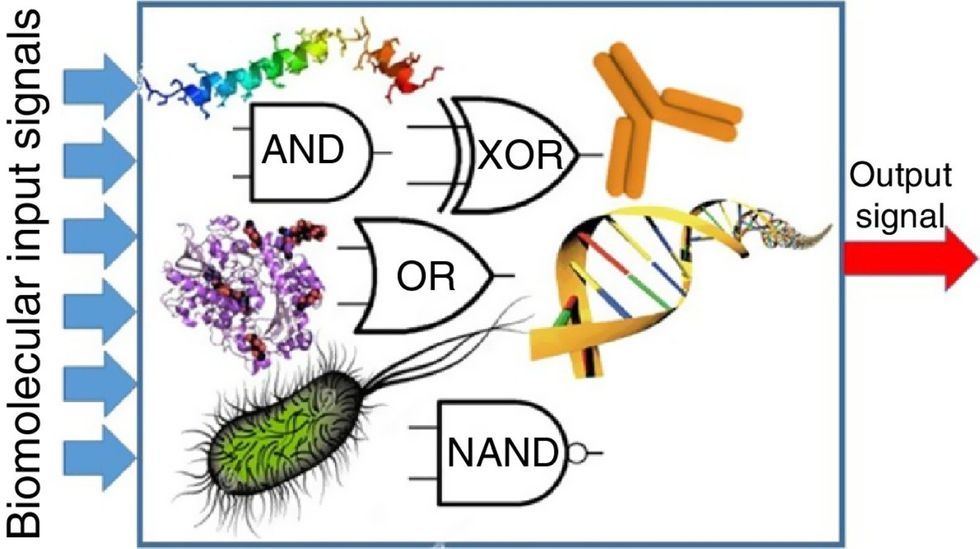
Biocomputing circles are made of DNA, RNA, proteins and even bacteria.
Evgeny Katz
The TRUMPET’s promise
Depending on whether the biocomputing system is placed directly inside a cell within the human body, or run in a test-tube, different environmental factors play a role. When an output is produced inside a cell, the cell's natural processes can amplify this output (for example, a specific protein or DNA strand), creating a solid signal. However, these cells can also be very leaky. “You want the cells to do the thing you ask them to do before they finish whatever their businesses, which is to grow, replicate, metabolize,” Adamala explains. “However, often the gate may be triggered without the right inputs, creating a false positive signal. So that's why natural logic gates are often leaky." While biocomputing outside a cell in a test tube can allow for tighter control over the logic gates, the outputs or signals cannot be amplified by a cell and are less potent.
TRUMPET, which is smaller than a cell, taps into both cellular and non-cellular biocomputing benefits. “At its core, it is a nonliving logic gate system,” Adamala states, “It's a DNA-based logic gate system. But because we use enzymes, and the readout is enzymatic [where an enzyme replicates the fluorescent RNA], we end up with signal amplification." This readout means that the output from the TRUMPET system, a fluorescent RNA strand, can be replicated by nearby enzymes in the platform, making the light signal stronger. "So it combines the best of both worlds,” Adamala adds.
These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body.
The TRUMPET biocomputing process is relatively straightforward. “If the DNA [input] shows up as single-stranded, it will not be digested [by the logic gate], and you get this nice fluorescent output as the RNA is made from the single-stranded DNA, and that's a 1,” Adamala explains. "And if the DNA input is double-stranded, it gets digested by the enzymes in the logic gate, and there is no RNA created from the DNA, so there is no fluorescence, and the output is 0." On the story's leading image above, if the tube is "lit" with a purple color, that is a binary 1 signal for computing. If it's "off" it is a 0.
While still in research, TRUMPET and other biocomputing systems promise significant benefits to personalized healthcare and medicine. These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body. The study’s lead author and graduate student Judee Sharon is already beginning to research TRUMPET's ability for earlier cancer diagnoses. Because the inputs for TRUMPET are single or double-stranded DNA, any mutated or cancerous DNA could theoretically be detected from the platform through the biocomputing process. Theoretically, devices like TRUMPET could be used to detect cancer and other diseases earlier.
Adamala sees TRUMPET not only as a detection system but also as a potential cancer drug delivery system. “Ideally, you would like the drug only to turn on when it senses the presence of a cancer cell. And that's how we use the logic gates, which work in response to inputs like cancerous DNA. Then the output can be the production of a small molecule or the release of a small molecule that can then go and kill what needs killing, in this case, a cancer cell. So we would like to develop applications that use this technology to control the logic gate response of a drug’s delivery to a cell.”
Although platforms like TRUMPET are making progress, a lot more work must be done before they can be used commercially. “The process of translating mechanisms and architecture from biology to computing and vice versa is still an art rather than a science,” says Forrest. “It requires deep computer science and biology knowledge,” she adds. “Some people have compared interdisciplinary science to fusion restaurants—not all combinations are successful, but when they are, the results are remarkable.”
Crickets are low on fat, high on protein, and can be farmed sustainably. They are also crunchy.
In today’s podcast episode, Leaps.org Deputy Editor Lina Zeldovich speaks about the health and ecological benefits of farming crickets for human consumption with Bicky Nguyen, who joins Lina from Vietnam. Bicky and her business partner Nam Dang operate an insect farm named CricketOne. Motivated by the idea of sustainable and healthy protein production, they started their unconventional endeavor a few years ago, despite numerous naysayers who didn’t believe that humans would ever consider munching on bugs.
Yet, making creepy crawlers part of our diet offers many health and planetary advantages. Food production needs to match the rise in global population, estimated to reach 10 billion by 2050. One challenge is that some of our current practices are inefficient, polluting and wasteful. According to nonprofit EarthSave.org, it takes 2,500 gallons of water, 12 pounds of grain, 35 pounds of topsoil and the energy equivalent of one gallon of gasoline to produce one pound of feedlot beef, although exact statistics vary between sources.
Meanwhile, insects are easy to grow, high on protein and low on fat. When roasted with salt, they make crunchy snacks. When chopped up, they transform into delicious pâtes, says Bicky, who invents her own cricket recipes and serves them at industry and public events. Maybe that’s why some research predicts that edible insects market may grow to almost $10 billion by 2030. Tune in for a delectable chat on this alternative and sustainable protein.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
Further reading:
More info on Bicky Nguyen
https://yseali.fulbright.edu.vn/en/faculty/bicky-n...
The environmental footprint of beef production
https://www.earthsave.org/environment.htm
https://www.watercalculator.org/news/articles/beef-king-big-water-footprints/
https://www.frontiersin.org/articles/10.3389/fsufs.2019.00005/full
https://ourworldindata.org/carbon-footprint-food-methane
Insect farming as a source of sustainable protein
https://www.insectgourmet.com/insect-farming-growing-bugs-for-protein/
https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/insect-farming
Cricket flour is taking the world by storm
https://www.cricketflours.com/
https://talk-commerce.com/blog/what-brands-use-cricket-flour-and-why/

Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.