With this new technology, hospitals and pharmacies could make vaccines and medicines onsite
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.

New research focuses on methods that could change medicine-making worldwide. The scientists propose bursting cells open, removing their DNA and using the cellular gears inside to make therapies.
Most modern biopharmaceutical medicines are produced by workhorse cells—typically bacterial but sometimes mammalian. The cells receive the synthesizing instructions on a snippet of a genetic code, which they incorporate into their DNA. The cellular machinery—ribosomes, RNAs, polymerases, and other compounds—read and use these instructions to build the medicinal molecules, which are harvested and administered to patients.
Although a staple of modern pharma, this process is complex and expensive. One must first insert the DNA instructions into the cells, which they may or may not uptake. One then must grow the cells, keeping them alive and well, so that they produce the required therapeutics, which then must be isolated and purified. To make this at scale requires massive bioreactors and big factories from where the drugs are distributed—and may take a while to arrive where they’re needed. “The pandemic showed us that this method is slow and cumbersome,” says Govind Rao, professor of biochemical engineering who directs the Center for Advanced Sensor Technology at the University of Maryland, Baltimore County (UMBC). “We need better methods that can work faster and can work locally where an outbreak is happening.”
Rao and his team of collaborators, which spans multiple research institutions, believe they have a better approach that may change medicine-making worldwide. They suggest forgoing the concept of using living cells as medicine-producers. Instead, they propose breaking the cells and using the remaining cellular gears for assembling the therapeutic compounds. Instead of inserting the DNA into living cells, the team burst them open, and removed their DNA altogether. Yet, the residual molecular machinery of ribosomes, polymerases and other cogwheels still functioned the way it would in a cell. “Now if you drop your DNA drug-making instructions into that soup, this machinery starts making what you need,” Rao explains. “And because you're no longer worrying about living cells, it becomes much simpler and more efficient.” The collaborators detail their cell-free protein synthesis or CFPS method in their recent paper published in preprint BioAxiv.
While CFPS does not use living cells, it still needs the basic building blocks to assemble proteins from—such as amino acids, nucleotides and certain types of enzymes. These are regularly added into this “soup” to keep the molecular factory chugging. “We just mix everything in as a batch and we let it integrate,” says James Robert Swartz, professor of chemical engineering and bioengineering at Stanford University and co-author of the paper. “And we make sure that we provide enough oxygen.” Rao likens the process to making milk from milk powder.
For a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology.
The idea of a cell-free protein synthesis is older than one might think. Swartz first experimented with it around 1997, when he was a chemical engineer at Genentech. While working on engineering bacteria to make pharmaceuticals, he discovered that there was a limit to what E. coli cells, the workhorse darling of pharma, could do. For example, it couldn’t grow and properly fold some complex proteins. “We tried many genetic engineering approaches, many fermentation, development, and environmental control approaches,” Swartz recalls—to no avail.
“The organism had its own agenda,” he quips. “And because everything was happening within the organism, we just couldn't really change those conditions very easily. Some of them we couldn’t change at all—we didn’t have control.”
It was out of frustration with the defiant bacteria that a new idea took hold. Could the cells be opened instead, so that the protein-forming reactions could be influenced more easily? “Obviously, we’d lose the ability for them to reproduce,” Swartz says. But that also meant that they no longer needed to keep the cells alive and could focus on making the specific reactions happen. “We could take the catalysts, the enzymes, and the more complex catalysts and activate them, make them work together, much as they would in a living cell, but the way we wanted.”
In 1998, Swartz joined Stanford, and began perfecting the biochemistry of the cell-free method, identifying the reactions he wanted to foster and stopping those he didn’t want. He managed to make the idea work, but for a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology. For their BioArxiv paper, the team tested the method by growing a specific antiviral protein called griffithsin.
First identified by Barry O’Keefe at National Cancer Institute over a decade ago, griffithsin is an antiviral known to interfere with many viruses’ ability to enter cells—including HIV, SARS, SARS-CoV-2, MERS and others. Originally isolated from the red algae Griffithsia, it works differently from antibodies and antibody cocktails.
Most antiviral medicines tend to target the specific receptors that viruses use to gain entry to the cells they infect. For example, SARS-CoV-2 uses the infamous spike protein to latch onto the ACE2 receptor of mammalian cells. The antibodies or other antiviral molecules stick to the spike protein, shutting off its ability to cling onto the ACE2 receptors. Unfortunately, the spike proteins mutate very often, so the medicines lose their potency. On the contrary, griffithsin has the ability to cling to the different parts of viral shells called capsids—namely to the molecules of mannose, a type of sugar. That extra stuff, glued all around the capsid like dead weight, makes it impossible for the virus to squeeze into the cell.
“Every time we have a vaccine or an antibody against a specific SARS-CoV-2 strain, that strain then mutates and so you lose efficacy,” Rao explains. “But griffithsin molecules glom onto the viral capsid, so the capsid essentially becomes a sticky mess and can’t enter the cell.” Mannose molecules also don’t mutate as easily as viruses’ receptors, so griffithsin-based antivirals do not have to be constantly updated. And because mannose molecules are found on many viruses’ capsids, it makes griffithsin “a universal neutralizer,” Rao explains.
“When griffithsin was discovered, we recognized that it held a lot of promise as a potential antiviral agent,” O’Keefe says. In 2010, he published a paper about griffithsin efficacy in neutralizing viruses of the corona family—after the first SARS outbreak in the early 2000s, the scientific community was interested in such antivirals. Yet, griffithsin is still not available as an off-the-shelf product. So during the Covid pandemic, the team experimented with synthesizing griffithsin using the cell-free production method. They were able to generate potent griffithsin in less than 24 hours without having to grow living cells.
The antiviral protein isn't the only type of medicine that can be made cell-free. The proteins needed for vaccine production could also be made the same way. “Such portable, on-demand drug manufacturing platforms can produce antiviral proteins within hours, making them ideal for combating future pandemics,” Rao says. “We would be able to stop the pandemic before it spreads.”
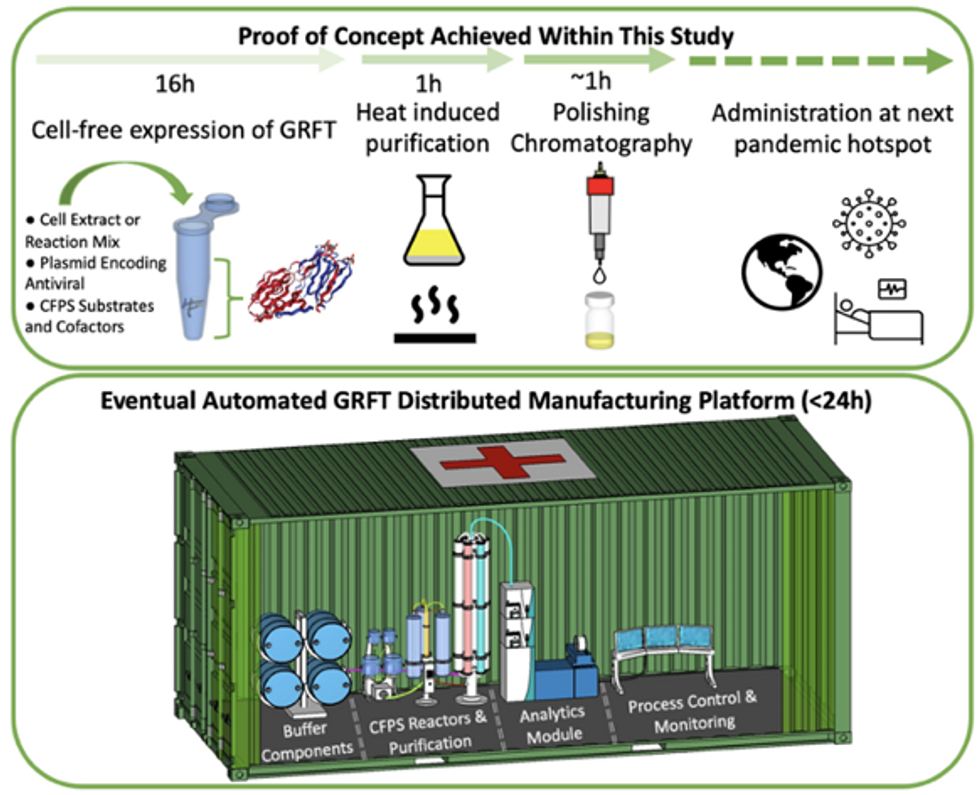
Top: Describes the process used in the study. Bottom: Describes how the new medicines and vaccines could be made at the site of a future viral outbreak.
Image courtesy of Rao and team, sourced from An approach to rapid distributed manufacturing of broad spectrumanti-viral griffithsin using cell-free systems to mitigate pandemics.
Rao’s idea is to perfect the technology to the point that any hospital or pharmacy can load up the media containing molecular factories, mix up the required amino acids, nucleotides and enzymes, and harvest the meds within hours. That will allow making medicines onsite and on demand. “That would be a self-contained production unit, so that you could just ship the production wherever the pandemic is breaking out,” says Swartz.
These units and the meds they produce, will, of course, have to undergo rigorous testing. “The biggest hurdles will be validating these against conventional technology,” Rao says. The biotech industry is risk-averse and prefers the familiar methods. But if this approach works, it may go beyond emergency situations and revolutionize the medicine-making paradigm even outside hospitals and pharmacies. Rao hopes that someday the method might become so mainstream that people may be able to buy and operate such reactors at home. “You can imagine a diabetic patient making insulin that way, or some other drugs,” Rao says. It would work not unlike making baby formula from the mere white powder. Just add water—and some oxygen, too.
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.
Did Anton the AI find a new treatment for a deadly cancer?
Researchers used a supercomputer to learn about the subtle movement of a cancer-causing molecule, and then they found the precise drug that can recognize that motion.
Bile duct cancer is a rare and aggressive form of cancer that is often difficult to diagnose. Patients with advanced forms of the disease have an average life expectancy of less than two years.
Many patients who get cancer in their bile ducts – the tubes that carry digestive fluid from the liver to the small intestine – have mutations in the protein FGFR2, which leads cells to grow uncontrollably. One treatment option is chemotherapy, but it’s toxic to both cancer cells and healthy cells, failing to distinguish between the two. Increasingly, cancer researchers are focusing on biomarker directed therapy, or making drugs that target a particular molecule that causes the disease – FGFR2, in the case of bile duct cancer.
A problem is that in targeting FGFR2, these drugs inadvertently inhibit the FGFR1 protein, which looks almost identical. This causes elevated phosphate levels, which is a sign of kidney damage, so doses are often limited to prevent complications.
In recent years, though, a company called Relay has taken a unique approach to picking out FGFR2, using a powerful supercomputer to simulate how proteins move and change shape. The team, leveraging this AI capability, discovered that FGFR2 and FGFR1 move differently, which enabled them to create a more precise drug.
Preliminary studies have shown robust activity of this drug, called RLY-4008, in FGFR2 altered tumors, especially in bile duct cancer. The drug did not inhibit FGFR1 or cause significant side effects. “RLY-4008 is a prime example of a precision oncology therapeutic with its highly selective and potent targeting of FGFR2 genetic alterations and resistance mutations,” says Lipika Goyal, assistant professor of medicine at Harvard Medical School. She is a principal investigator of Relay’s phase 1-2 clinical trial.
Boosts from AI and a billionaire
Traditional drug design has been very much a case of trial and error, as scientists investigate many molecules to see which ones bind to the intended target and bind less to other targets.
“It’s being done almost blindly, without really being guided by structure, so it fails very often,” says Olivier Elemento, associate director of the Institute for Computational Biomedicine at Cornell. “The issue is that they are not sampling enough molecules to cover some of the chemical space that would be specific to the target of interest and not specific to others.”
Relay’s unique hardware and software allow simulations that could never be achieved through traditional experiments, Elemento says.
Some scientists have tried to use X-rays of crystallized proteins to look at the structure of proteins and design better drugs. But they have failed to account for an important factor: proteins are moving and constantly folding into different shapes.
David Shaw, a hedge fund billionaire, wanted to help improve drug discovery and understood that a key obstacle was that computer models of molecular dynamics were limited; they simulated motion for less than 10 millionths of a second.
In 2001, Shaw set up his own research facility, D.E. Shaw Research, to create a supercomputer that would be specifically designed to simulate protein motion. Seven years later, he succeeded in firing up a supercomputer that can now conduct high speed simulations roughly 100 times faster than others. Called Anton, it has special computer chips to enable this speed, and its software is powered by AI to conduct many simulations.
After creating the supercomputer, Shaw teamed up with leading scientists who were interested in molecular motion, and they founded Relay Therapeutics.
Elemento believes that Relay’s approach is highly beneficial in designing a better drug for bile duct cancer. “Relay Therapeutics has a cutting-edge approach for molecular dynamics that I don’t believe any other companies have, at least not as advanced.” Relay’s unique hardware and software allow simulations that could never be achieved through traditional experiments, Elemento says.
How it works
Relay used both experimental and computational approaches to design RLY-4008. The team started out by taking X-rays of crystallized versions of both their intended target, FGFR2, and the almost identical FGFR1. This enabled them to get a 3D snapshot of each of their structures. They then fed the X-rays into the Anton supercomputer to simulate how the proteins were likely to move.
Anton’s simulations showed that the FGFR1 protein had a flap that moved more frequently than FGFR2. Based on this distinct motion, the team tried to design a compound that would recognize this flap shifting around and bind to FGFR2 while steering away from its more active lookalike.
For that, they went back Anton, using the supercomputer to simulate the behavior of thousands of potential molecules for over a year, looking at what made a particular molecule selective to the target versus another molecule that wasn’t. These insights led them to determine the best compounds to make and test in the lab and, ultimately, they found that RLY-4008 was the most effective.
Promising results so far
Relay began phase 1-2 trials in 2020 and will continue until 2024. Preliminary results showed that, in the 17 patients taking a 70 mg dose of RLY-4008, the drug worked to shrink tumors in 88 percent of patients. This was a significant increase compared to other FGFR inhibitors. For instance, Futibatinib, which recently got FDA approval, had a response rate of only 42 percent.
Across all dose levels, RLY-4008 shrank tumors by 63 percent in 38 patients. In more good news, the drug didn’t elevate their phosphate levels, which suggests that it could be taken without increasing patients’ risk for kidney disease.
“Objectively, this is pretty remarkable,” says Elemento. “In a small patient study, you have a molecule that is able to shrink tumors in such a high fraction of patients. It is unusual to see such good results in a phase 1-2 trial.”
A simulated future
The research team is continuing to use molecular dynamic simulations to develop other new drug, such as one that is being studied in patients with solid tumors and breast cancer.
As for their bile duct cancer drug, RLY-4008, Relay plans by 2024 to have tested it in around 440 patients. “The mature results of the phase 1-2 trial are highly anticipated,” says Goyal, the principal investigator of the trial.
Sameek Roychowdhury, an oncologist and associate professor of internal medicine at Ohio State University, highlights the need for caution. “This has early signs of benefit, but we will look forward to seeing longer term results for benefit and side effect profiles. We need to think a few more steps ahead - these treatments are like the ’Whack-a-Mole game’ where cancer finds a way to become resistant to each subsequent drug.”
“I think the issue is going to be how durable are the responses to the drug and what are the mechanisms of resistance,” says Raymond Wadlow, an oncologist at the Inova Medical Group who specializes in gastrointestinal and haematological cancer. “But the results look promising. It is a much more selective inhibitor of the FGFR protein and less toxic. It’s been an exciting development.”
The Friday Five: How to exercise for cancer prevention
How to exercise for cancer prevention. Plus, a device that brings relief to back pain, ingredients for reducing Alzheimer's risk, the world's oldest disease could make you young again, and more.
The Friday Five covers five stories in research that you may have missed this week. There are plenty of controversies and troubling ethical issues in science – and we get into many of them in our online magazine – but this news roundup focuses on scientific creativity and progress to give you a therapeutic dose of inspiration headed into the weekend.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
Here are the promising studies covered in this week's Friday Five:
- How to exercise for cancer prevention
- A device that brings relief to back pain
- Ingredients for reducing Alzheimer's risk
- Is the world's oldest disease the fountain of youth?
- Scared of crossing bridges? Your phone can help

