A Cancer Researcher Opens Up About His Astonishing Breakthrough
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
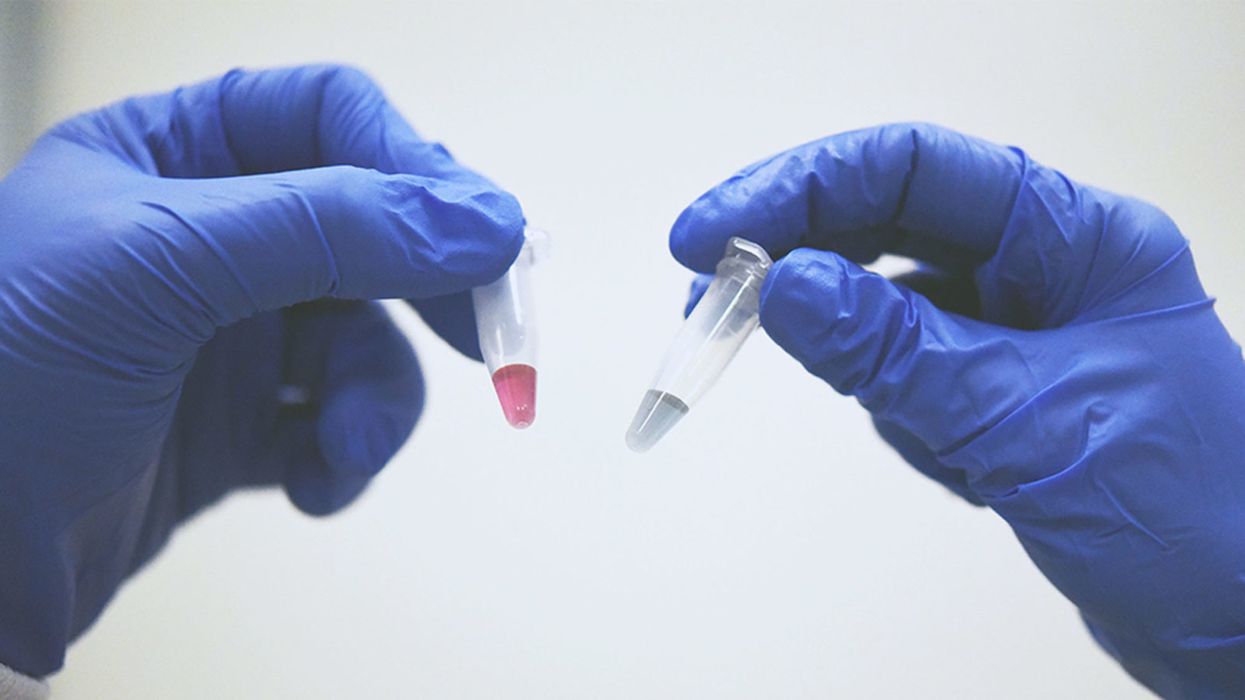
A simple ten-minute universal cancer test that can be detected by the human eye or an electronic device - published in Nature Communications (Dec 2018) by the Trau lab at the University of Queensland. Red indicates the presence of cancerous cells and blue doesn't.
Matt Trau, a professor of chemistry at the University of Queensland, stunned the science world back in December when the prestigious journal Nature Communications published his lab's discovery about a unique property of cancer DNA that could lead to a simple, cheap, and accurate test to detect any type of cancer in under 10 minutes.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke."
Trau granted very few interviews in the wake of the news, but he recently opened up to leapsmag about the significance of this promising early research. Here is his story in his own words, as told to Editor-in-Chief Kira Peikoff.
There's been an incredible explosion of knowledge over the past 20 years, particularly since the genome was sequenced. The area of diagnostics has a tremendous amount of promise and has caught our lab's interest. If you catch cancer early, you can improve survival rates to as high as 98 percent, sometimes even now surpassing that.
My lab is interested in devices to improve the trajectory of cancer patients. So, once people get diagnosed, can we get really sophisticated information about the molecular origins of the disease, and can we measure it in real time? And then can we match that with the best treatment and monitor it in real time, too?
I think those approaches, also coupled with immunotherapy, where one dreams of monitoring the immune system simultaneously with the disease progress, will be the future.
But currently, the methodologies for cancer are still pretty old. So, for example, let's talk about biopsies in general. Liquid biopsy just means using a blood test or a urine test, rather than extracting out a piece of solid tissue. Now consider breast cancer. Still, the cutting-edge screening method is mammography or the physical interrogation for lumps. This has had a big impact in terms of early detection and awareness, but it's still primitive compared to interrogating, forensically, blood samples to look at traces of DNA.
Large machines like CAT scans, PET scans, MRIs, are very expensive and very subjective in terms of the operator. They don't look at the root causes of the cancer. Cancer is caused by changes in DNA. These can be changes in the hard drive of the DNA (the genomic changes) or changes in the apps that the DNA are running (the epigenetics and the transcriptomics).
We don't look at that now, even though we have, emerging, all of these technologies to do it, and those technologies are getting so much cheaper. I saw some statistics at a conference just a few months ago that, in the United States, less than 1 percent of cancer patients have their DNA interrogated. That's the current state-of-the-art in the modern medical system.

Professor Matt Trau, a cancer researcher at the University of Queensland in Australia.
(Courtesy)
Blood, as the highway of the body, is carrying all of this information. Cancer cells, if they are present in the body, are constantly getting turned over. When they die, they release their contents into the blood. Many of these cells end up in the urine and saliva. Having technologies that can forensically scan the highways looking for evidence of cancer is little bit like looking for explosives at the airport. That's very valuable as a security tool.
The trouble is that there are thousands of different types of cancer. Going back to breast cancer, there's at least a dozen different types, probably more, and each of them change the DNA (the hard drive of the disease) and the epigenetics (or the RAM memory). So one of the problems for diagnostics in cancer is to find something that is a signature of all cancers. That's been a really, really, really difficult problem.
Ours was a completely serendipitous discovery. What we found in the lab was this one marker that just kept coming up in all of the types of breast cancers we were studying.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke, maybe it works just for breast cancer." So we went on to test it in prostate cancer, which is also many different types of diseases, and it seemed to be working in all of those. We then tested it further in lymphoma. Again, many different types of lymphoma. It worked across all of those. We tested it in gastrointestinal cancer. Again, many different types, and still, it worked, but we were skeptical.
Then we looked at cell lines, which are cells that have come from previous cancer patients, that we grow in the lab, but are used as model experimental systems. We have many of those cell lines, both ones that are cancerous, and ones that are healthy. It was quite remarkable that the marker worked in all of the cancer cell lines and didn't work in the healthy cell lines.
What could possibly be going on?
Well, imagine DNA as a piece of string, that's your hard drive. Epigenetics is like the beads that you put on that string. Those beads you can take on and off as you wish and they control which apps are run, meaning which genetic programs the cell runs. We hypothesized that for cancer, those beads cluster together, rather than being randomly distributed across the string.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office.
The implications of this are profound. It means that DNA from cancer folds in water into three-dimensional structures that are very different from healthy cells' DNA. It's quite literally the needle in a haystack. Because when you do a liquid biopsy for early detection of cancer, most of the DNA from blood contains a vast abundance of healthy DNA. And that's not of interest. What's of interest is to find the cancerous DNA. That's there only in trace.
Once we figured out what was going on, we could easily set up a system to detect the trace cancerous DNA. It binds to gold nanoparticles in water and changes color. The test takes 10 minutes, and you can detect it by eye. Red indicates cancer and blue doesn't.
We're very, very excited about where we go from here. We're starting to test the test on a greater number of cancers, in thousands of patient samples. We're looking to the scientific community to engage with us, and we're getting a really good response from groups around the world who are supplying more samples to us so we can test this more broadly.
We also are very interested in testing how early can we go with this test. Can we detect cancer through a simple blood test even before there are any symptoms whatsoever? If so, we might be able to convert a cancer diagnosis to something almost as good as a vaccine.
Of course, we have to watch what are called false positives. We don't want to be detecting people as positives when they don't have cancer, and so the technology needs to improve there. We see this version as the iPhone 1. We're interested in the iPhone 2, 3, 4, getting better and better.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office. If it came back positive, your doctor could say, "Look, there's some news here, but actually, it's not bad news, it's good news. We've caught this so early that we will be able to manage this, and this won't be a problem for you."
If this were to be in routine use in the medical system, countless lives could be saved. Cancer is now becoming one of the biggest killers in the world. We're talking millions upon millions upon millions of people who are affected. This really motivates our work. We might make a difference there.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
Regenerative medicine has come a long way, baby
After a cloned baby sheep, what started as one of the most controversial areas in medicine is now promising to transform it.
The field of regenerative medicine had a shaky start. In 2002, when news spread about the first cloned animal, Dolly the sheep, a raucous debate ensued. Scary headlines and organized opposition groups put pressure on government leaders, who responded by tightening restrictions on this type of research.
Fast forward to today, and regenerative medicine, which focuses on making unhealthy tissues and organs healthy again, is rewriting the code to healing many disorders, though it’s still young enough to be considered nascent. What started as one of the most controversial areas in medicine is now promising to transform it.
Progress in the lab has addressed previous concerns. Back in the early 2000s, some of the most fervent controversy centered around somatic cell nuclear transfer (SCNT), the process used by scientists to produce Dolly. There was fear that this technique could be used in humans, with possibly adverse effects, considering the many medical problems of the animals who had been cloned.
But today, scientists have discovered better approaches with fewer risks. Pioneers in the field are embracing new possibilities for cellular reprogramming, 3D organ printing, AI collaboration, and even growing organs in space. It could bring a new era of personalized medicine for longer, healthier lives - while potentially sparking new controversies.
Engineering tissues from amniotic fluids
Work in regenerative medicine seeks to reverse damage to organs and tissues by culling, modifying and replacing cells in the human body. Scientists in this field reach deep into the mechanisms of diseases and the breakdowns of cells, the little workhorses that perform all life-giving processes. If cells can’t do their jobs, they take whole organs and systems down with them. Regenerative medicine seeks to harness the power of healthy cells derived from stem cells to do the work that can literally restore patients to a state of health—by giving them healthy, functioning tissues and organs.
Modern-day regenerative medicine takes its origin from the 1998 isolation of human embryonic stem cells, first achieved by John Gearhart at Johns Hopkins University. Gearhart isolated the pluripotent cells that can differentiate into virtually every kind of cell in the human body. There was a raging controversy about the use of these cells in research because at that time they came exclusively from early-stage embryos or fetal tissue.
Back then, the highly controversial SCNT cells were the only way to produce genetically matched stem cells to treat patients. Since then, the picture has changed radically because other sources of highly versatile stem cells have been developed. Today, scientists can derive stem cells from amniotic fluid or reprogram patients’ skin cells back to an immature state, so they can differentiate into whatever types of cells the patient needs.
In the context of medical history, the field of regenerative medicine is progressing at a dizzying speed. But for those living with aggressive or chronic illnesses, it can seem that the wheels of medical progress grind slowly.
The ethical debate has been dialed back and, in the last few decades, the field has produced important innovations, spurring the development of whole new FDA processes and categories, says Anthony Atala, a bioengineer and director of the Wake Forest Institute for Regenerative Medicine. Atala and a large team of researchers have pioneered many of the first applications of 3D printed tissues and organs using cells developed from patients or those obtained from amniotic fluid or placentas.
His lab, considered to be the largest devoted to translational regenerative medicine, is currently working with 40 different engineered human tissues. Sixteen of them have been transplanted into patients. That includes skin, bladders, urethras, muscles, kidneys and vaginal organs, to name just a few.
These achievements are made possible by converging disciplines and technologies, such as cell therapies, bioengineering, gene editing, nanotechnology and 3D printing, to create living tissues and organs for human transplants. Atala is currently overseeing clinical trials to test the safety of tissues and organs engineered in the Wake Forest lab, a significant step toward FDA approval.
In the context of medical history, the field of regenerative medicine is progressing at a dizzying speed. But for those living with aggressive or chronic illnesses, it can seem that the wheels of medical progress grind slowly.
“It’s never fast enough,” Atala says. “We want to get new treatments into the clinic faster, but the reality is that you have to dot all your i’s and cross all your t’s—and rightly so, for the sake of patient safety. People want predictions, but you can never predict how much work it will take to go from conceptualization to utilization.”
As a surgeon, he also treats patients and is able to follow transplant recipients. “At the end of the day, the goal is to get these technologies into patients, and working with the patients is a very rewarding experience,” he says. Will the 3D printed organs ever outrun the shortage of donated organs? “That’s the hope,” Atala says, “but this technology won’t eliminate the need for them in our lifetime.”
New methods are out of this world
Jeanne Loring, another pioneer in the field and director of the Center for Regenerative Medicine at Scripps Research Institute in San Diego, says that investment in regenerative medicine is not only paying off, but is leading to truly personalized medicine, one of the holy grails of modern science.
This is because a patient’s own skin cells can be reprogrammed to become replacements for various malfunctioning cells causing incurable diseases, such as diabetes, heart disease, macular degeneration and Parkinson’s. If the cells are obtained from a source other than the patient, they can be rejected by the immune system. This means that patients need lifelong immunosuppression, which isn’t ideal. “With Covid,” says Loring, “I became acutely aware of the dangers of immunosuppression.” Using the patient’s own cells eliminates that problem.
Microgravity conditions make it easier for the cells to form three-dimensional structures, which could more easily lead to the growing of whole organs. In fact, Loring's own cells have been sent to the ISS for study.
Loring has a special interest in neurons, or brain cells that can be developed by manipulating cells found in the skin. She is looking to eventually treat Parkinson’s disease using them. The manipulated cells produce dopamine, the critical hormone or neurotransmitter lacking in the brains of patients. A company she founded plans to start a Phase I clinical trial using cell therapies for Parkinson’s soon, she says.
This is the culmination of many years of basic research on her part, some of it on her own cells. In 2007, Loring had her own cells reprogrammed, so there’s a cell line that carries her DNA. “They’re just like embryonic stem cells, but personal,” she said.
Loring has another special interest—sending immature cells into space to be studied at the International Space Station. There, microgravity conditions make it easier for the cells to form three-dimensional structures, which could more easily lead to the growing of whole organs. In fact, her own cells have been sent to the ISS for study. “My colleagues and I have completed four missions at the space station,” she says. “The last cells came down last August. They were my own cells reprogrammed into pluripotent cells in 2009. No one else can say that,” she adds.
Future controversies and tipping points
Although the original SCNT debate has calmed down, more controversies may arise, Loring thinks.
One of them could concern growing synthetic embryos. The embryos are ultimately derived from embryonic stem cells, and it’s not clear to what stage these embryos can or will be grown in an artificial uterus—another recent invention. The science, so far done only in animals, is still new and has not been widely publicized but, eventually, “People will notice the production of synthetic embryos and growing them in an artificial uterus,” Loring says. It’s likely to incite many of the same reactions as the use of embryonic stem cells.
Bernard Siegel, the founder and director of the Regenerative Medicine Foundation and executive director of the newly formed Healthspan Action Coalition (HSAC), believes that stem cell science is rapidly approaching tipping point and changing all of medical science. (For disclosure, I do consulting work for HSAC). Siegel says that regenerative medicine has become a new pillar of medicine that has recently been fast-tracked by new technology.
Artificial intelligence is speeding up discoveries and the convergence of key disciplines, as demonstrated in Atala’s lab, which is creating complex new medical products that replace the body’s natural parts. Just as importantly, those parts are genetically matched and pose no risk of rejection.
These new technologies must be regulated, which can be a challenge, Siegel notes. “Cell therapies represent a challenge to the existing regulatory structure, including payment, reimbursement and infrastructure issues that 20 years ago, didn’t exist.” Now the FDA and other agencies are faced with this revolution, and they’re just beginning to adapt.
Siegel cited the 2021 FDA Modernization Act as a major step. The Act allows drug developers to use alternatives to animal testing in investigating the safety and efficacy of new compounds, loosening the agency’s requirement for extensive animal testing before a new drug can move into clinical trials. The Act is a recognition of the profound effect that cultured human cells are having on research. Being able to test drugs using actual human cells promises to be far safer and more accurate in predicting how they will act in the human body, and could accelerate drug development.
Siegel, a longtime veteran and founding father of several health advocacy organizations, believes this work helped bring cell therapies to people sooner rather than later. His new focus, through the HSAC, is to leverage regenerative medicine into extending not just the lifespan but the worldwide human healthspan, the period of life lived with health and vigor. “When you look at the HSAC as a tree,” asks Siegel, “what are the roots of that tree? Stem cell science and the huge ecosystem it has created.” The study of human aging is another root to the tree that has potential to lengthen healthspans.
The revolutionary science underlying the extension of the healthspan needs to be available to the whole world, Siegel says. “We need to take all these roots and come up with a way to improve the life of all mankind,” he says. “Everyone should be able to take advantage of this promising new world.”
Joy Milne's unusual sense of smell led Dr. Tilo Kunath, a neurobiologist at the Centre for Regenerative Medicine at the University of Edinburgh, and a host of other scientists, to develop a new diagnostic test for Parkinson's.
Forty years ago, Joy Milne, a nurse from Perth, Scotland, noticed a musky odor coming from her husband, Les. At first, Milne thought the smell was a result of bad hygiene and badgered her husband to take longer showers. But when the smell persisted, Milne learned to live with it, not wanting to hurt her husband's feelings.
Twelve years after she first noticed the "woodsy" smell, Les was diagnosed at the age of 44 with Parkinson's Disease, a neurodegenerative condition characterized by lack of dopamine production and loss of movement. Parkinson's Disease currently affects more than 10 million people worldwide.
Milne spent the next several years believing the strange smell was exclusive to her husband. But to her surprise, at a local support group meeting in 2012, she caught the familiar scent once again, hanging over the group like a cloud. Stunned, Milne started to wonder if the smell was the result of Parkinson's Disease itself.
Milne's discovery led her to Dr. Tilo Kunath, a neurobiologist at the Centre for Regenerative Medicine at the University of Edinburgh. Together, Milne, Kunath, and a host of other scientists would use Milne's unusual sense of smell to develop a new diagnostic test, now in development and poised to revolutionize the treatment of Parkinson's Disease.
"Joy was in the audience during a talk I was giving on my work, which has to do with Parkinson's and stem cell biology," Kunath says. "During the patient engagement portion of the talk, she asked me if Parkinson's had a smell to it." Confused, Kunath said he had never heard of this – but for months after his talk he continued to turn the question over in his mind.
Kunath knew from his research that the skin's microbiome changes during different disease processes, releasing metabolites that can give off odors. In the medical literature, diseases like melanoma and Type 2 diabetes have been known to carry a specific scent – but no such connection had been made with Parkinson's. If people could smell Parkinson's, he thought, then it stood to reason that those metabolites could be isolated, identified, and used to potentially diagnose Parkinson's by their presence alone.
First, Kunath and his colleagues decided to test Milne's sense of smell. "I got in touch with Joy again and we designed a protocol to test her sense of smell without her having to be around patients," says Kunath, which could have affected the validity of the test. In his spare time, Kunath collected t-shirt samples from people diagnosed with Parkinson's and from others without the diagnosis and gave them to Milne to smell. In 100 percent of the samples, Milne was able to detect whether a person had Parkinson's based on smell alone. Amazingly, Milne was even able to detect the "Parkinson's scent" in a shirt from the control group – someone who did not have a Parkinson's diagnosis, but would go on to be diagnosed nine months later.
From the initial study, the team discovered that Parkinson's did have a smell, that Milne – inexplicably – could detect it, and that she could detect it long before diagnosis like she had with her husband, Les. But the experiments revealed other things that the team hadn't been expecting.
"One surprising thing we learned from that experiment was that the odor was always located in the back of the shirt – never in the armpit, where we expected the smell to be," Kunath says. "I had a chance meeting with a dermatologist and he said the smell was due to the patient's sebum, which are greasy secretions that are really dense on your upper back. We have sweat glands, instead of sebum, in our armpits." Patients with Parkinson's are also known to have increased sebum production.
With the knowledge that a patient's sebum was the source of the unusual smell, researchers could go on to investigate exactly what metabolites were in the sebum and in what amounts. Kunath, along with his associate, Dr. Perdita Barran, collected and analyzed sebum samples from 64 participants across the United Kingdom. Once the samples were collected, Barran and others analyzed it using a method called gas chromatography mass spectrometry, or GS-MC, which separated, weighed and helped identify the individual compounds present in each sebum sample.
Barran's team can now correctly identify Parkinson's in nine out of 10 patients – a much quicker and more accurate way to diagnose than what clinicians do now.
"The compounds we've identified in the sebum are not unique to people with Parkinson's, but they are differently expressed," says Barran, a professor of mass spectrometry at the University of Manchester. "So this test we're developing now is not a black-and-white, do-you-have-something kind of test, but rather how much of these compounds do you have compared to other people and other compounds." The team identified over a dozen compounds that were present in the sebum of Parkinson's patients in much larger amounts than the control group.
Using only the GC-MS and a sebum swab test, Barran's team can now correctly identify Parkinson's in nine out of 10 patients – a much quicker and more accurate way to diagnose than what clinicians do now.
"At the moment, a clinical diagnosis is based on the patient's physical symptoms," Barran says, and determining whether a patient has Parkinson's is often a long and drawn-out process of elimination. "Doctors might say that a group of symptoms looks like Parkinson's, but there are other reasons people might have those symptoms, and it might take another year before they're certain," Barran says. "Some of those symptoms are just signs of aging, and other symptoms like tremor are present in recovering alcoholics or people with other kinds of dementia." People under the age of 40 with Parkinson's symptoms, who present with stiff arms, are often misdiagnosed with carpal tunnel syndrome, she adds.
Additionally, by the time physical symptoms are present, Parkinson's patients have already lost a substantial amount of dopamine receptors – about sixty percent -- in the brain's basal ganglia. Getting a diagnosis before physical symptoms appear would mean earlier interventions that could prevent dopamine loss and preserve regular movement, Barran says.
"Early diagnosis is good if it means there's a chance of early intervention," says Barran. "It stops the process of dopamine loss, which means that motor symptoms potentially will not happen, or the onset of symptoms will be substantially delayed." Barran's team is in the processing of streamlining the sebum test so that definitive results will be ready in just two minutes.
"What we're doing right now will be a very inexpensive test, a rapid-screen test, and that will encourage people to self-sample and test at home," says Barran. In addition to diagnosing Parkinson's, she says, this test could also be potentially useful to determine if medications were at a therapeutic dose in people who have the disease, since the odor is strongest in people whose symptoms are least controlled by medication.
"When symptoms are under control, the odor is lower," Barran says. "Potentially this would allow patients and clinicians to see whether their symptoms are being managed properly with medication, or perhaps if they're being overmedicated." Hypothetically, patients could also use the test to determine if interventions like diet and exercise are effective at keeping Parkinson's controlled.
"We hope within the next two to five years we will have a test available."
Barran is now running another clinical trial – one that determines whether they can diagnose at an earlier stage and whether they can identify a difference in sebum samples between different forms of Parkinson's or diseases that have Parkinson's-like symptoms, such as Lewy Body Dementia.
"Within the next one to two years, we hope to be running a trial in the Manchester area for those people who do not have motor symptoms but are at risk for developing dementia due to symptoms like loss of smell and sleep difficulty," Barran had said in 2019. "If we can establish that, we can roll out a test that determines if you have Parkinson's or not with those first pre-motor symptoms, and then at what stage. We hope within the next two to five years we will have a test available."
In a 2022 study, published in the American Chemical Society, researchers used mass spectrometry to analyze sebum from skin swabs for the presence of the specific molecules. They found that some specific molecules are present only in people who have Parkinson’s. Now they hope that the same method can be used in regular diagnostic labs. The test, many years in the making, is inching its way to the clinic.
"We would likely first give this test to people who are at risk due to a genetic predisposition, or who are at risk based on prodomal symptoms, like people who suffer from a REM sleep disorder who have a 50 to 70 percent chance of developing Parkinson's within a ten year period," Barran says. "Those would be people who would benefit from early therapeutic intervention. For the normal population, it isn't beneficial at the moment to know until we have therapeutic interventions that can be useful."
Milne's husband, Les, passed away from complications of Parkinson's Disease in 2015. But thanks to him and the dedication of his wife, Joy, science may have found a way to someday prolong the lives of others with this devastating disease. Sometimes she can smell people who have Parkinson’s while in the supermarket or walking down the street but has been told by medical ethicists she cannot tell them, Milne said in an interview with the Guardian. But once the test becomes available in the clinics, it will do the job for her.
[Ed. Note: A older version of this hit article originally ran on September 3, 2019.]

