Scientists redesign bacteria to tackle the antibiotic resistance crisis
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
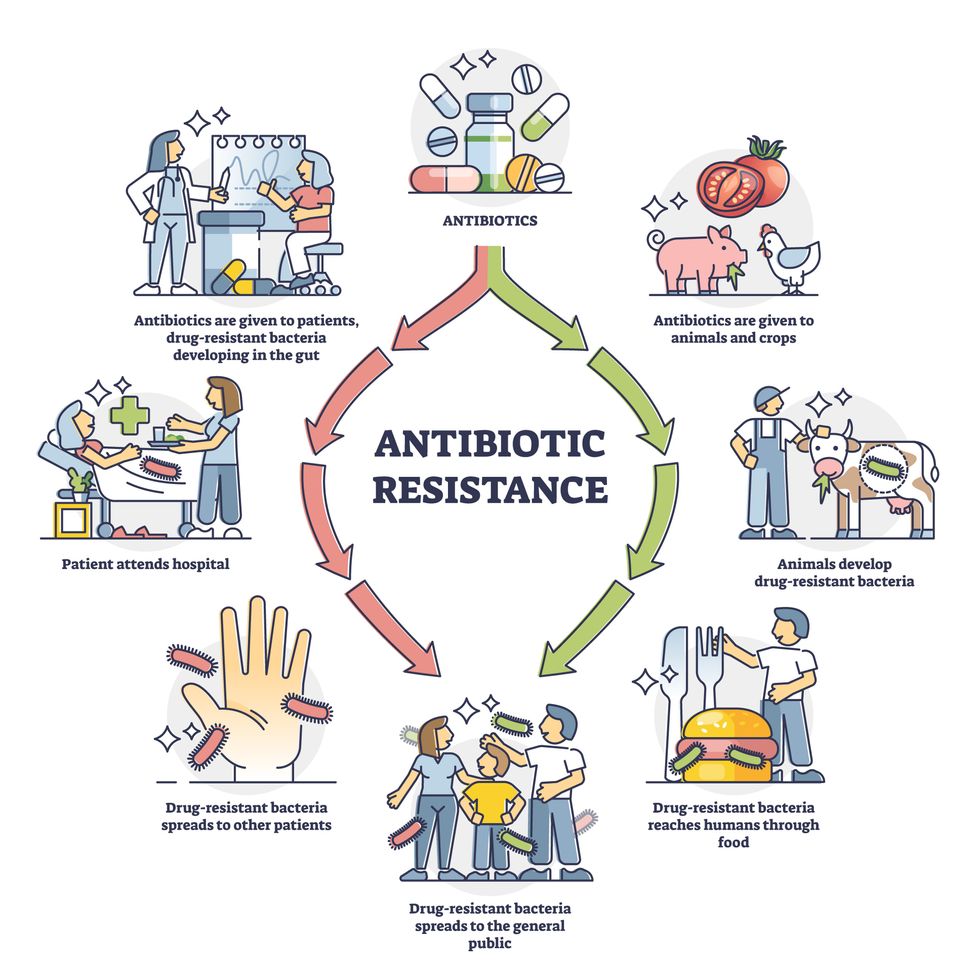
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
Fast for Longevity, with Less Hunger, with Dr. Valter Longo
Valter Longo, a biogerontologist at USC, and centenarian Rocco Longo (no relation) appear together in Italy in 2021. The elder Longo is from a part of Italy where people have fasted regularly and are enjoying long lifespans.
You’ve probably heard about intermittent fasting, where you don’t eat for about 16 hours each day and limit the window where you’re taking in food to the remaining eight hours.
But there’s another type of fasting, called a fasting-mimicking diet, with studies pointing to important benefits. For today’s podcast episode, I chatted with Dr. Valter Longo, a biogerontologist at the University of Southern California, about all kinds of fasting, and particularly the fasting-mimicking diet, which minimizes hunger as much as possible. Going without food for a period of time is an example of good stress: challenges that work at the cellular level to boost health and longevity.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
If you’ve ever spent more than a few minutes looking into fasting, you’ve almost certainly come upon Dr. Longo's name. He is the author of the bestselling book, The Longevity Diet, and the best known researcher of fasting-mimicking diets.
With intermittent fasting, your body might begin to switch up its fuel type. It's usually running on carbs you get from food, which gets turned into glucose, but without food, your liver starts making something called ketones, which are molecules that may benefit the body in a number of ways.
With the fasting-mimicking diet, you go for several days eating only types of food that, in a way, keep themselves secret from your body. So at the level of your cells, the body still thinks that it’s fasting. This is the best of both worlds – you’re not completely starving because you do take in some food, and you’re getting the boosts to health that come with letting a fast run longer than intermittent fasting. In this episode, Dr. Longo talks about the growing number of studies showing why this could be very advantageous for health, as long as you undertake the diet no more than a few times per year.
Dr. Longo is the director of the Longevity Institute at USC’s Leonard Davis School of Gerontology, and the director of the Longevity and Cancer program at the IFOM Institute of Molecular Oncology in Milan. In addition, he's the founder and president of the Create Cures Foundation in L.A., which focuses on nutrition for the prevention and treatment of major chronic illnesses. In 2016, he received the Glenn Award for Research on Aging for the discovery of genes and dietary interventions that regulate aging and prevent diseases. Dr. Longo received his PhD in biochemistry from UCLA and completed his postdoc in the neurobiology of aging and Alzheimer’s at USC.
Show links:
Create Cures Foundation, founded by Dr. Longo: www.createcures.org
Dr. Longo's Facebook: https://www.facebook.com/profvalterlongo/
Dr. Longo's Instagram: https://www.instagram.com/prof_valterlongo/
Dr. Longo's book: The Longevity Diet
The USC Longevity Institute: https://gero.usc.edu/longevity-institute/
Dr. Longo's research on nutrition, longevity and disease: https://pubmed.ncbi.nlm.nih.gov/35487190/
Dr. Longo's research on fasting mimicking diet and cancer: https://pubmed.ncbi.nlm.nih.gov/34707136/
Full list of Dr. Longo's studies: https://pubmed.ncbi.nlm.nih.gov/?term=Longo%2C+Valter%5BAuthor%5D&sort=date
Research on MCT oil and Alzheimer's: https://alz-journals.onlinelibrary.wiley.com/doi/f...
Keto Mojo device for measuring ketones
Silkworms with spider DNA spin silk stronger than Kevlar
Silkworm silk is fragile, which limits its uses, but a few extra genes can help.
Story by Freethink
The study and copying of nature’s models, systems, or elements to address complex human challenges is known as “biomimetics.” Five hundred years ago, an elderly Italian polymath spent months looking at the soaring flight of birds. The result was Leonardo da Vinci’s biomimetic Codex on the Flight of Birds, one of the foundational texts in the science of aerodynamics. It’s the science that elevated the Wright Brothers and has yet to peak.
Today, biomimetics is everywhere. Shark-inspired swimming trunks, gecko-inspired adhesives, and lotus-inspired water-repellents are all taken from observing the natural world. After millions of years of evolution, nature has quite a few tricks up its sleeve. They are tricks we can learn from. And now, thanks to some spider DNA and clever genetic engineering, we have another one to add to the list.
The elusive spider silk
We’ve known for a long time that spider silk is remarkable, in ways that synthetic fibers can’t emulate. Nylon is incredibly strong (it can support a lot of force), and Kevlar is incredibly tough (it can absorb a lot of force). But neither is both strong and tough. In all artificial polymeric fibers, strength and toughness are mutually exclusive, and so we pick the material best for the job and make do.
Spider silk, a natural polymeric fiber, breaks this rule. It is somehow both strong and tough. No surprise, then, that spider silk is a source of much study.The problem, though, is that spiders are incredibly hard to cultivate — let alone farm. If you put them together, they will attack and kill each other until only one or a few survive. If you put 100 spiders in an enclosed space, they will go about an aggressive, arachnocidal Hunger Games. You need to give each its own space and boundaries, and a spider hotel is hard and costly. Silkworms, on the other hand, are peaceful and productive. They’ll hang around all day to make the silk that has been used in textiles for centuries. But silkworm silk is fragile. It has very limited use.
The elusive – and lucrative – trick, then, would be to genetically engineer a silkworm to produce spider-quality silk. So far, efforts have been fruitless. That is, until now.
We can have silkworms creating silk six times as tough as Kevlar and ten times as strong as nylon.
Spider-silkworms
Junpeng Mi and his colleagues working at Donghua University, China, used CRISPR gene-editing technology to recode the silk-creating properties of a silkworm. First, they took genes from Araneus ventricosus, an East Asian orb-weaving spider known for its strong silk. Then they placed these complex genes – genes that involve more than 100 amino acids – into silkworm egg cells. (This description fails to capture how time-consuming, technical, and laborious this was; it’s a procedure that requires hundreds of thousands of microinjections.)
This had all been done before, and this had failed before. Where Mi and his team succeeded was using a concept called “localization.” Localization involves narrowing in on a very specific location in a genome. For this experiment, the team from Donghua University developed a “minimal basic structure model” of silkworm silk, which guided the genetic modifications. They wanted to make sure they had the exactly right transgenic spider silk proteins. Mi said that combining localization with this basic structure model “represents a significant departure from previous research.” And, judging only from the results, he might be right. Their “fibers exhibited impressive tensile strength (1,299 MPa) and toughness (319 MJ/m3), surpassing Kevlar’s toughness 6-fold.”
A world of super-materials
Mi’s research represents the bursting of a barrier. It opens up hugely important avenues for future biomimetic materials. As Mi puts it, “This groundbreaking achievement effectively resolves the scientific, technical, and engineering challenges that have hindered the commercialization of spider silk, positioning it as a viable alternative to commercially synthesized fibers like nylon and contributing to the advancement of ecological civilization.”
Around 60 percent of our clothing is made from synthetic fibers like nylon, polyester, and acrylic. These plastics are useful, but often bad for the environment. They shed into our waterways and sometimes damage wildlife. The production of these fibers is a source of greenhouse gas emissions. Now, we have a “sustainable, eco-friendly high-strength and ultra-tough alternative.” We can have silkworms creating silk six times as tough as Kevlar and ten times as strong as nylon.
We shouldn’t get carried away. This isn’t going to transform the textiles industry overnight. Gene-edited silkworms are still only going to produce a comparatively small amount of silk – even if farmed in the millions. But, as Mi himself concedes, this is only the beginning. If Mi’s localization and structure-model techniques are as remarkable as they seem, then this opens up the door to a great many supermaterials.
Nature continues to inspire. We had the bird, the gecko, and the shark. Now we have the spider-silkworm. What new secrets will we unravel in the future? And in what exciting ways will it change the world?
This article originally appeared on Freethink, home of the brightest minds and biggest ideas of all time.


