Scientists redesign bacteria to tackle the antibiotic resistance crisis
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
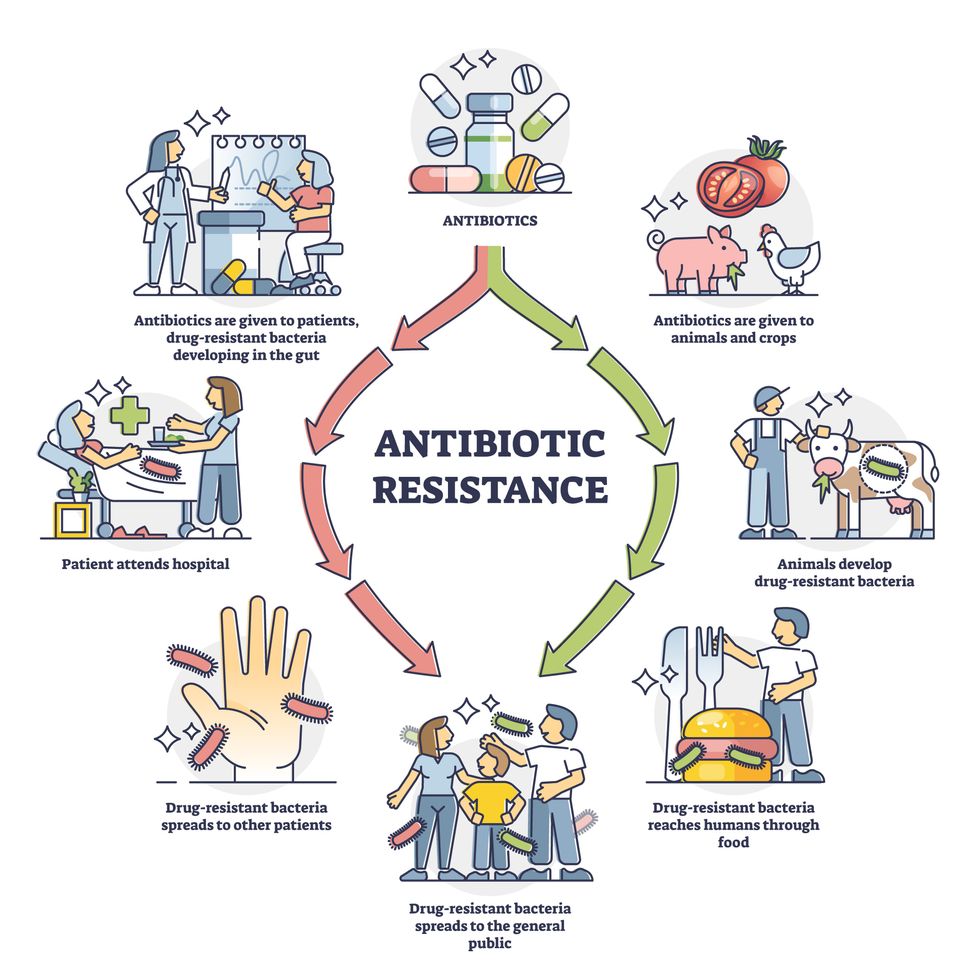
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
Scientists find enzymes in nature that could replace toxic chemicals
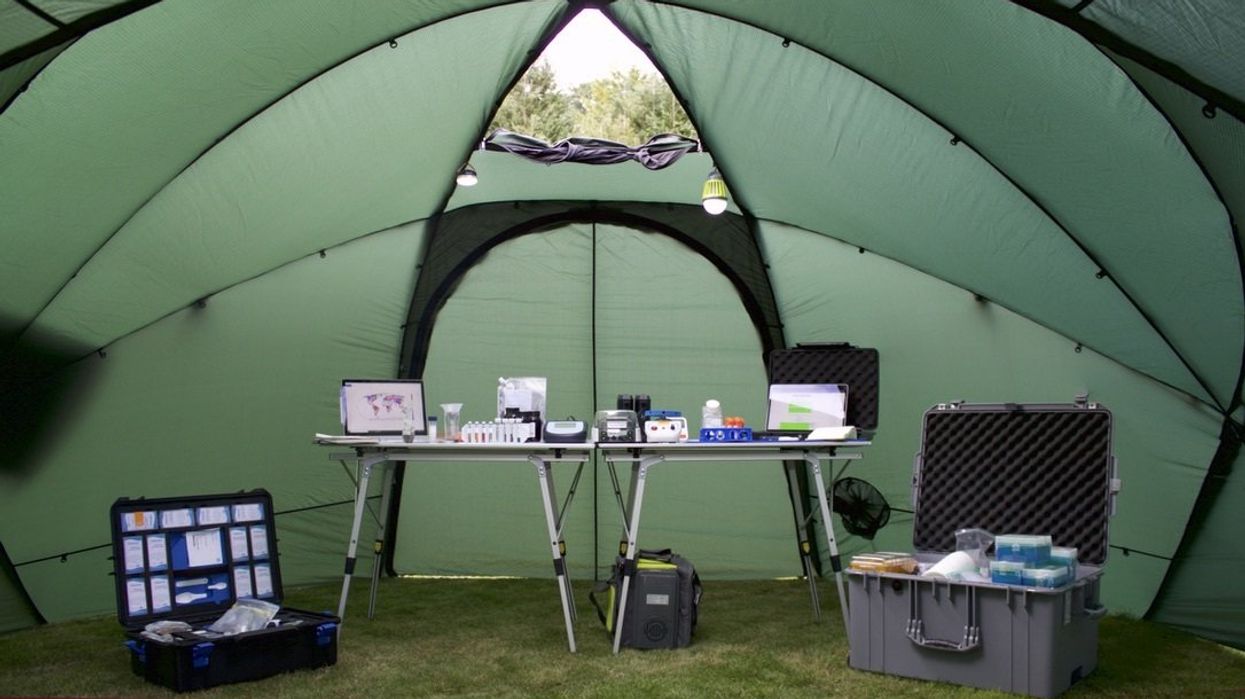
Basecamp Research is using portable labs like this one to gather samples from ecosystems around the world.
Some 900 miles off the coast of Portugal, nine major islands rise from the mid-Atlantic. Verdant and volcanic, the Azores archipelago hosts a wealth of biodiversity that keeps field research scientist, Marlon Clark, returning for more. “You’ve got this really interesting biogeography out there,” says Clark. “There’s real separation between the continents, but there’s this inter-island dispersal of plants and seeds and animals.”
It’s a visual paradise by any standard, but on a microscopic level, there’s even more to see. The Azores’ nutrient-rich volcanic rock — and its network of lagoons, cave systems, and thermal springs — is home to a vast array of microorganisms found in a variety of microclimates with different elevations and temperatures.
Clark works for Basecamp Research, a biotech company headquartered in London, and his job is to collect samples from ecosystems around the world. By extracting DNA from soil, water, plants, microbes and other organisms, Basecamp is building an extensive database of the Earth’s proteins. While DNA itself isn’t a protein, the information stored in DNA is used to create proteins, so extracting, sequencing, and annotating DNA allows for the discovery of unique protein sequences.
Using what they’re finding in the middle of the Atlantic and beyond, Basecamp’s detailed database is constantly growing. The outputs could be essential for cleaning up the damage done by toxic chemicals and finding alternatives to these chemicals.
Catalysts for change
Proteins provide structure and function in all living organisms. Some of these functional proteins are enzymes, which quite literally make things happen.
“Industrial chemistry is heavily polluting, especially the chemistry done in pharmaceutical drug development. Biocatalysis is providing advantages, both to make more complex drugs and to be more sustainable, reducing the pollution and toxicity of conventional chemistry," says Ahir Pushpanath, who heads partnerships for Basecamp.
“Enzymes are perfectly evolved catalysts,” says Ahir Pushpanath, a partnerships lead at Basecamp. ”Enzymes are essentially just a polymer, and polymers are made up of amino acids, which are nature’s building blocks.” He suggests thinking about it like Legos — if you have a bunch of Lego pieces and use them to build a structure that performs a function, “that’s basically how an enzyme works. In nature, these monuments have evolved to do life’s chemistry. If we didn’t have enzymes, we wouldn’t be alive.”
In our own bodies, enzymes catalyze everything from vision to digesting food to regrowing muscles, and these same types of enzymes are necessary in the pharmaceutical, agrochemical and fine chemical industries. But industrial conditions differ from those inside our bodies. So, when scientists need certain chemical reactions to create a particular product or substance, they make their own catalysts in their labs — generally through the use of petroleum and heavy metals.
These petrochemicals are effective and cost-efficient, but they’re wasteful and often hazardous. With growing concerns around sustainability and long-term public health, it's essential to find alternative solutions to toxic chemicals. “Industrial chemistry is heavily polluting, especially the chemistry done in pharmaceutical drug development,” Pushpanath says.
Basecamp is trying to replace lab-created catalysts with enzymes found in the wild. This concept is called biocatalysis, and in theory, all scientists have to do is find the right enzymes for their specific need. Yet, historically, researchers have struggled to find enzymes to replace petrochemicals. When they can’t identify a suitable match, they turn to what Pushpanath describes as “long, iterative, resource-intensive, directed evolution” in the laboratory to coax a protein into industrial adaptation. But the latest scientific advances have enabled these discoveries in nature instead.
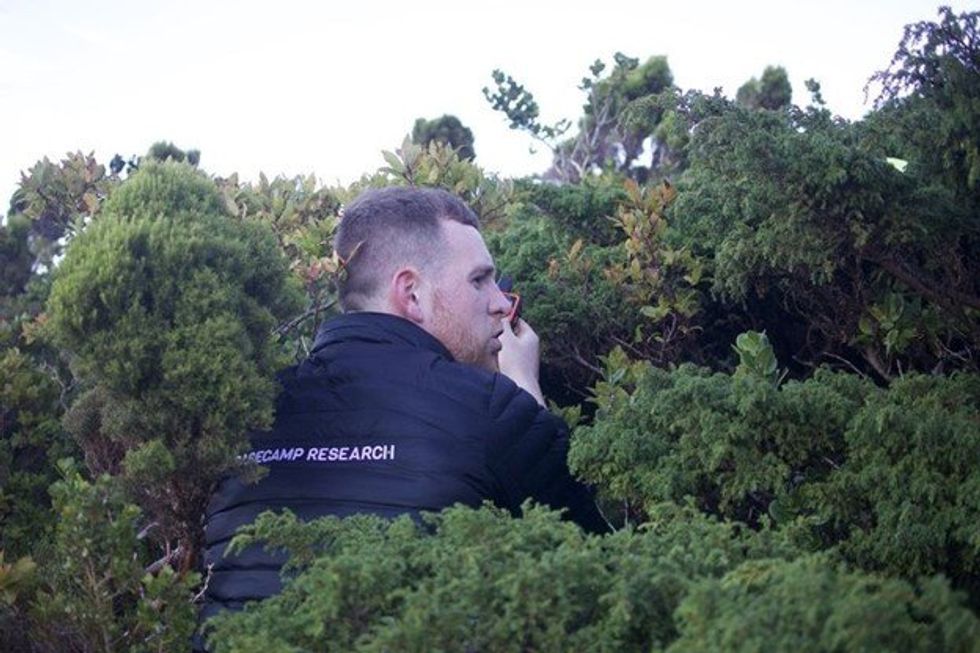

Marlon Clark, a research scientist at Basecamp Research, looks for novel biochemistries in the Azores.
Glen Gowers
Enzyme hunters
Whether it’s Clark and a colleague setting off on an expedition, or a local, on-the-ground partner gathering and processing samples, there’s a lot to be learned from each collection. “Microbial genomes contain complete sets of information that define an organism — much like how letters are a code allowing us to form words, sentences, pages, and books that contain complex but digestible knowledge,” Clark says. He thinks of the environmental samples as biological libraries, filled with thousands of species, strains, and sequence variants. “It’s our job to glean genetic information from these samples.”
“We can actually dream up new proteins using generative AI," Pushpanath says.
Basecamp researchers manage this feat by sequencing the DNA and then assembling the information into a comprehensible structure. “We’re building the ‘stories’ of the biota,” Clark says. The more varied the samples, the more valuable insights his team gains into the characteristics of different organisms and their interactions with the environment. Sequencing allows scientists to examine the order of nucleotides — the organic molecules that form DNA — to identify genetic makeups and find changes within genomes. The process used to be too expensive, but the cost of sequencing has dropped from $10,000 a decade ago to as low as $100. Notably, biocatalysis isn’t a new concept — there have been waves of interest in using natural enzymes in catalysis for over a century, Pushpanath says. “But the technology just wasn’t there to make it cost effective,” he explains. “Sequencing has been the biggest boon.”
AI is probably the second biggest boon.
“We can actually dream up new proteins using generative AI,” Pushpanath says, which means that biocataylsis now has real potential to scale.
Glen Gowers, the co-founder of Basecamp, compares the company’s AI approach to that of social networks and streaming services. Consider how these platforms suggest connecting with the friends of your friends, or how watching one comedy film from the 1990s leads to a suggestion of three more.
“They’re thinking about data as networks of relationships as opposed to lists of items,” says Gowers. “By doing the same, we’re able to link the metadata of the proteins — by their relationships to each other, the environments in which they’re found, the way those proteins might look similar in sequence and structure, their surrounding genome context — really, this just comes down to creating a searchable network of proteins.”
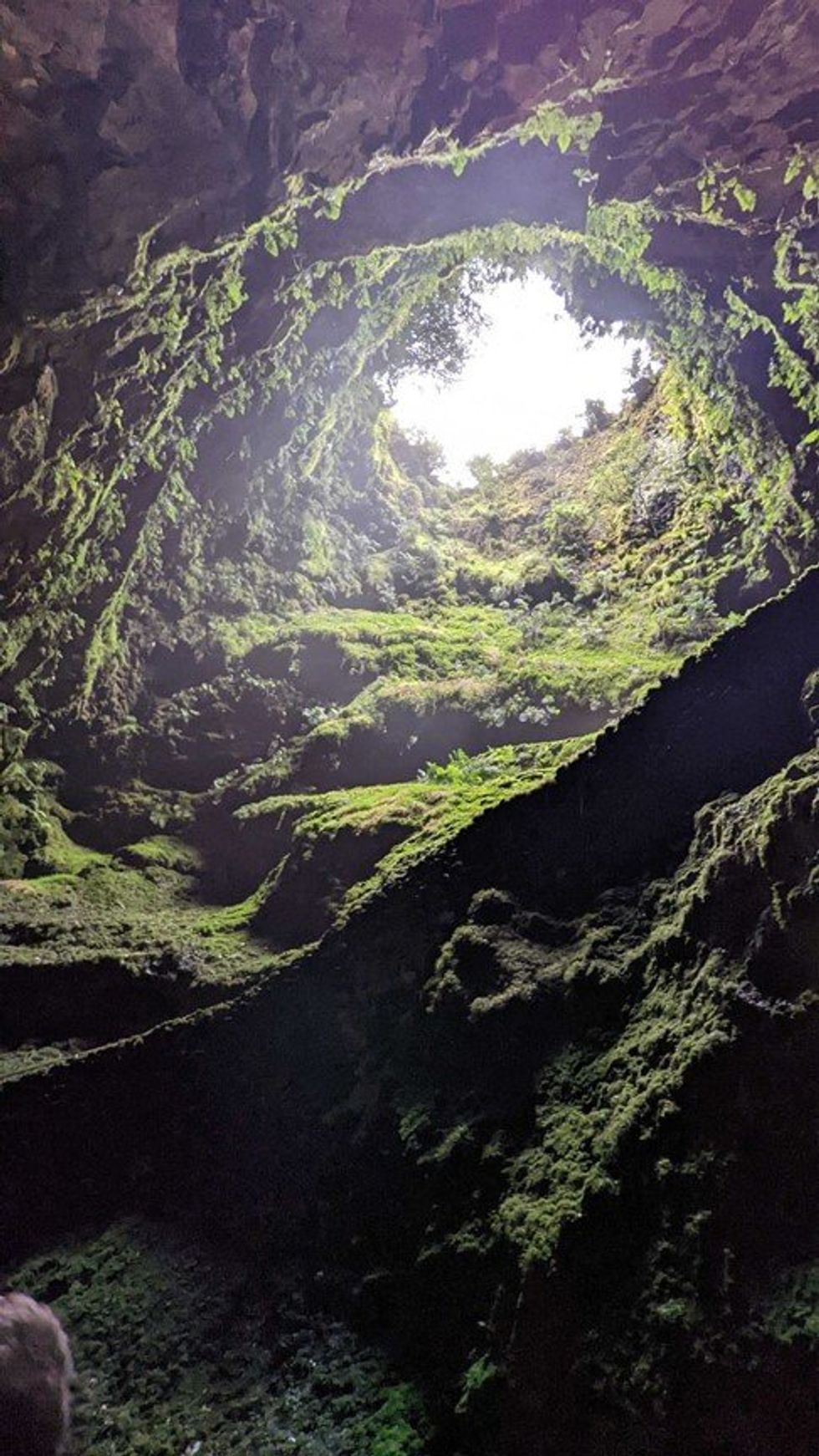

On an Azores island, this volcanic opening may harbor organisms that can help scientists identify enzymes for biocatalysis to replace toxic chemicals.
Emma Bolton
Uwe Bornscheuer, professor at the Institute of Biochemistry at the University of Greifswald, and co-founder of Enzymicals, another biocatalysis company, says that the development of machine learning is a critical component of this work. “It’s a very hot topic, because the challenge in protein engineering is to predict which mutation at which position in the protein will make an enzyme suitable for certain applications,” Bornscheuer explains. These predictions are difficult for humans to make at all, let alone quickly. “It is clear that machine learning is a key technology.”
Benefiting from nature’s bounty
Biodiversity commonly refers to plants and animals, but the term extends to all life, including microbial life, and some regions of the world are more biodiverse than others. Building relationships with global partners is another key element to Basecamp’s success. Doing so in accordance with the access and benefit sharing principles set forth by the Nagoya Protocol — an international agreement that seeks to ensure the benefits of using genetic resources are distributed in a fair and equitable way — is part of the company's ethos. “There's a lot of potential for us, and there’s a lot of potential for our partners to have exactly the same impact in building and discovering commercially relevant proteins and biochemistries from nature,” Clark says.
Bornscheuer points out that Basecamp is not the first company of its kind. A former San Diego company called Diversa went public in 2000 with similar work. “At that time, the Nagoya Protocol was not around, but Diversa also wanted to ensure that if a certain enzyme or microorganism from Costa Rica, for example, were used in an industrial process, then people in Costa Rica would somehow profit from this.”
An eventual merger turned Diversa into Verenium Corporation, which is now a part of the chemical producer BASF, but it laid important groundwork for modern companies like Basecamp to continue to scale with today’s technologies.
“To collect natural diversity is the key to identifying new catalysts for use in new applications,” Bornscheuer says. “Natural diversity is immense, and over the past 20 years we have gained the advantages that sequencing is no longer a cost or time factor.”
This has allowed Basecamp to rapidly grow its database, outperforming Universal Protein Resource or UniProt, which is the public repository of protein sequences most commonly used by researchers. Basecamp’s database is three times larger, totaling about 900 million sequences. (UniProt isn’t compliant with the Nagoya Protocol, because, as a public database, it doesn’t provide traceability of protein sequences. Some scientists, however, argue that Nagoya compliance hinders progress.)
“Eventually, this work will reduce chemical processes. We’ll have cleaner processes, more sustainable processes," says Uwe Bornscheuer, a professor at the University of Greifswald.
With so much information available, Basecamp’s AI has been trained on “the true dictionary of protein sequence life,” Pushpanath says, which makes it possible to design sequences for particular applications. “Through deep learning approaches, we’re able to find protein sequences directly from our database, without the need for further laboratory-directed evolution.”
Recently, a major chemical company was searching for a specific transaminase — an enzyme that catalyzes a transfer of amino groups. “They had already spent a year-and-a-half and nearly two million dollars to evolve a public-database enzyme, and still had not reached their goal,” Pushpanath says. “We used our AI approaches on our novel database to yield 10 candidates within a week, which, when validated by the client, achieved the desired target even better than their best-evolved candidate.”
Basecamp’s other huge potential is in bioremediation, where natural enzymes can help to undo the damage caused by toxic chemicals. “Biocatalysis impacts both sides,” says Gowers. “It reduces the usage of chemicals to make products, and at the same time, where contamination sites do exist from chemical spills, enzymes are also there to kind of mop those up.”
So far, Basecamp's round-the-world sampling has covered 50 percent of the 14 major biomes, or regions of the planet that can be distinguished by their flora, fauna, and climate, as defined by the World Wildlife Fund. The other half remains to be catalogued — a key milestone for understanding our planet’s protein diversity, Pushpanath notes.
There’s still a long road ahead to fully replace petrochemicals with natural enzymes, but biocatalysis is on an upward trajectory. "Eventually, this work will reduce chemical processes,” Bornscheuer says. “We’ll have cleaner processes, more sustainable processes.”
Small changes in how a person talks could reveal Alzheimer’s earlier
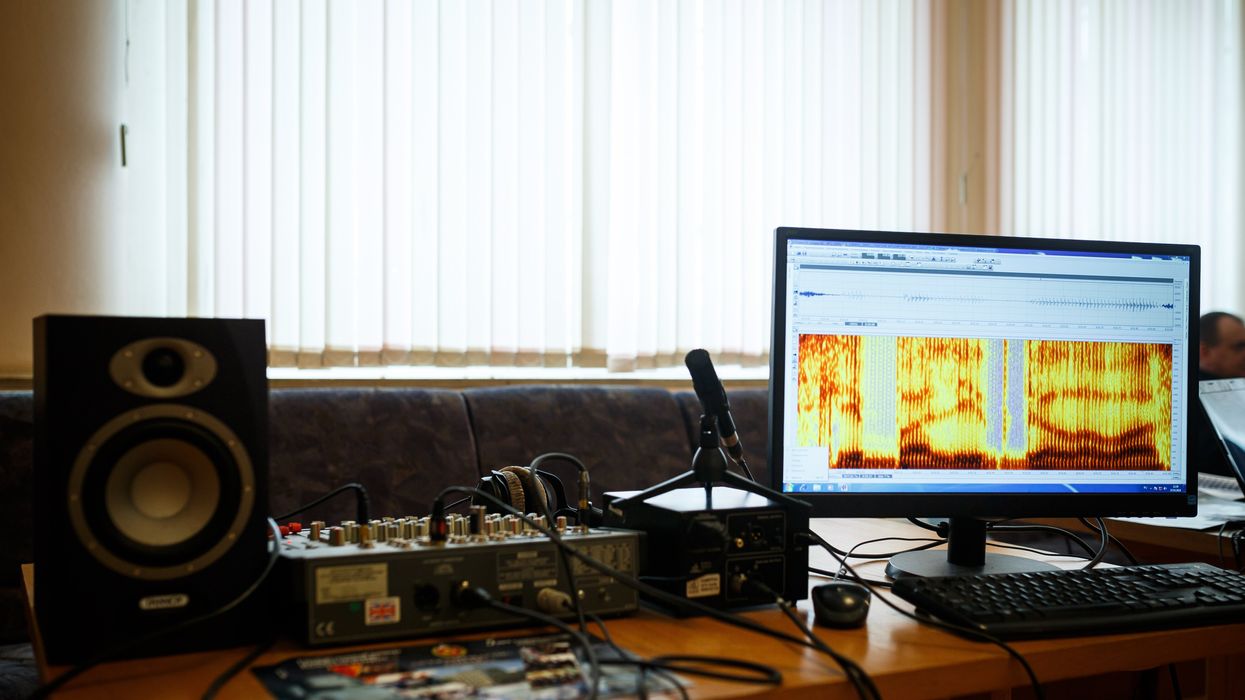
In an initial study, Canary analyzed speech recordings with AI and identified early stage Alzheimer’s with 96 percent accuracy.
Dave Arnold retired in his 60s and began spending time volunteering in local schools. But then he started misplacing items, forgetting appointments and losing his sense of direction. Eventually he was diagnosed with early stage Alzheimer’s.
“Hearing the diagnosis made me very emotional and tearful,” he said. “I immediately thought of all my mom had experienced.” His mother suffered with the condition for years before passing away. Over the last year, Arnold has worked for the Alzheimer’s Association as one of its early stage advisors, sharing his insights to help others in the initial stages of the disease.
Arnold was diagnosed sooner than many others. It's important to find out early, when interventions can make the most difference. One promising avenue is looking at how people talk. Research has shown that Alzheimer’s affects a part of the brain that controls speech, resulting in small changes before people show other signs of the disease.
Now, Canary Speech, a company based in Utah, is using AI to examine elements like the pitch of a person’s voice and their pauses. In an initial study, Canary analyzed speech recordings with AI and identified early stage Alzheimer’s with 96 percent accuracy.
Developing the AI model
Canary Speech’s CEO, Henry O’Connell, met cofounder Jeff Adams about 40 years before they started the company. Back when they first crossed paths, they were both living in Bethesda, Maryland; O’Connell was a research fellow at the National Institutes of Health studying rare neurological diseases, while Adams was working to decode spy messages. Later on, Adams would specialize in building mathematical models to analyze speech and sound as a team leader in developing Amazon's Alexa.
It wasn't until 2015 that they decided to make use of the fit between their backgrounds. ““We established Canary Speech in 2017 to build a product that could be used in multiple languages in clinical environments,” O'Connell says.
The need is growing. About 55 million people worldwide currently live with Alzheimer’s, a number that is expected to double by 2050. Some scientists think the disease results from a buildup of plaque in the brain. It causes mild memory loss at first and, over time, this issue get worse while other symptoms, such as disorientation and hallucinations, can develop. Treatment to manage the disease is more effective in the earlier stages, but detection is difficult since mild symptoms are often attributed to the normal aging process.
O’Connell and Adams specialize in the complex ways that Alzheimer’s effects how people speak. Using AI, their mathematical model analyzes 15 million data points every minute, focusing on certain features of speech such as pitch, pauses and elongation of words. It also pays attention to how the vibrations of vocal cords change in different stages of the disease.
To create their model, the team used a type of machine learning called deep neural nets, which looks at multiple layers of data - in this case, the multiple features of a person’s speech patterns.
“Deep neural nets allow us to look at much, much larger data sets built out of millions of elements,” O’Connell explained. “Through machine learning and AI, we’ve identified features that are very sensitive to an Alzheimer’s patient versus [people without the disease] and also very sensitive to mild cognitive impairment, early stage and moderate Alzheimer's.” Based on their learnings, Canary is able to classify the disease stage very quickly, O’Connell said.
“When we’re listening to sublanguage elements, we’re really analyzing the direct result of changes in the brain in the physical body,” O’Connell said. “The brain controls your vocal cords: how fast they vibrate, the expansion of them, the contraction.” These factors, along with where people put their tongues when talking, function subconsciously and result in subtle changes in the sounds of speech.
Further testing is needed
In an initial trial, Canary analyzed speech recordings from phone calls to a large U.S. health insurer. They looked at the audio recordings of 651 policyholders who had early stage Alzheimer’s and 1018 who did not have the condition, aiming for a representative sample of age, gender and race. They used this data to create their first diagnostic model and found that it was 96 percent accurate in identifying Alzheimer’s.
Christian Herff, an assistant professor of neuroscience at Maastricht University in the Netherlands, praised this approach while adding that further testing is needed to assess its effectiveness.
“I think the general idea of identifying increased risk for cognitive impairment based on speech characteristics is very feasible, particularly when change in a user’s voice is monitored, for example, by recording speech every year,” Herff said. He noted that this can only be a first indication, not a full diagnosis. The accuracy still needs to be validated in studies that follows individuals over a period of time, he said.
Toby Walsh, a professor of artificial intelligence at the University of New South Wales, also thinks Canary’s tool has potential but highlights that Canary could diagnose some people who don’t really have the disease. “This is an interesting and promising application of AI,” he said, “but these tools need to be used carefully. Imagine the anxiety of being misdiagnosed with Alzheimer’s.”
As with many other AI tools, privacy and bias are additional issues to monitor closely, Walsh said.
Other languages
A related issue is that not everyone is fluent in English. Mahnaz Arvaneh, a senior lecturer in automatic control and systems engineering at the University of Sheffield, said this could be a blind spot.
“The system may not be very accurate for those who have English as their second language as their speaking patterns would be different, and any issue might be because of language deficiency rather than cognitive issues,” Arvaneh said.
The team is expanding to multiple languages starting with Japanese and Spanish. The elements of the model that make up the algorithm are very similar, but they need to be validated and retrained in a different language, which will require access to more data.
Recently, Canary analyzed the phone calls of 233 Japanese patients who had mild cognitive impairment and 704 healthy people. Using an English model they were able to identify the Japanese patients who had mild cognitive impairment with 78 percent accuracy. They also developed a model in Japanese that was 45 percent accurate, and they’re continuing to train it with more data.
The future
Canary is using their model to look at other diseases like Huntington’s and Parkinson’s. They’re also collaborating with pharmaceuticals to validate potential therapies for Alzheimer’s. By looking at speech patterns over time, Canary can get an indication of how well these drugs are working.
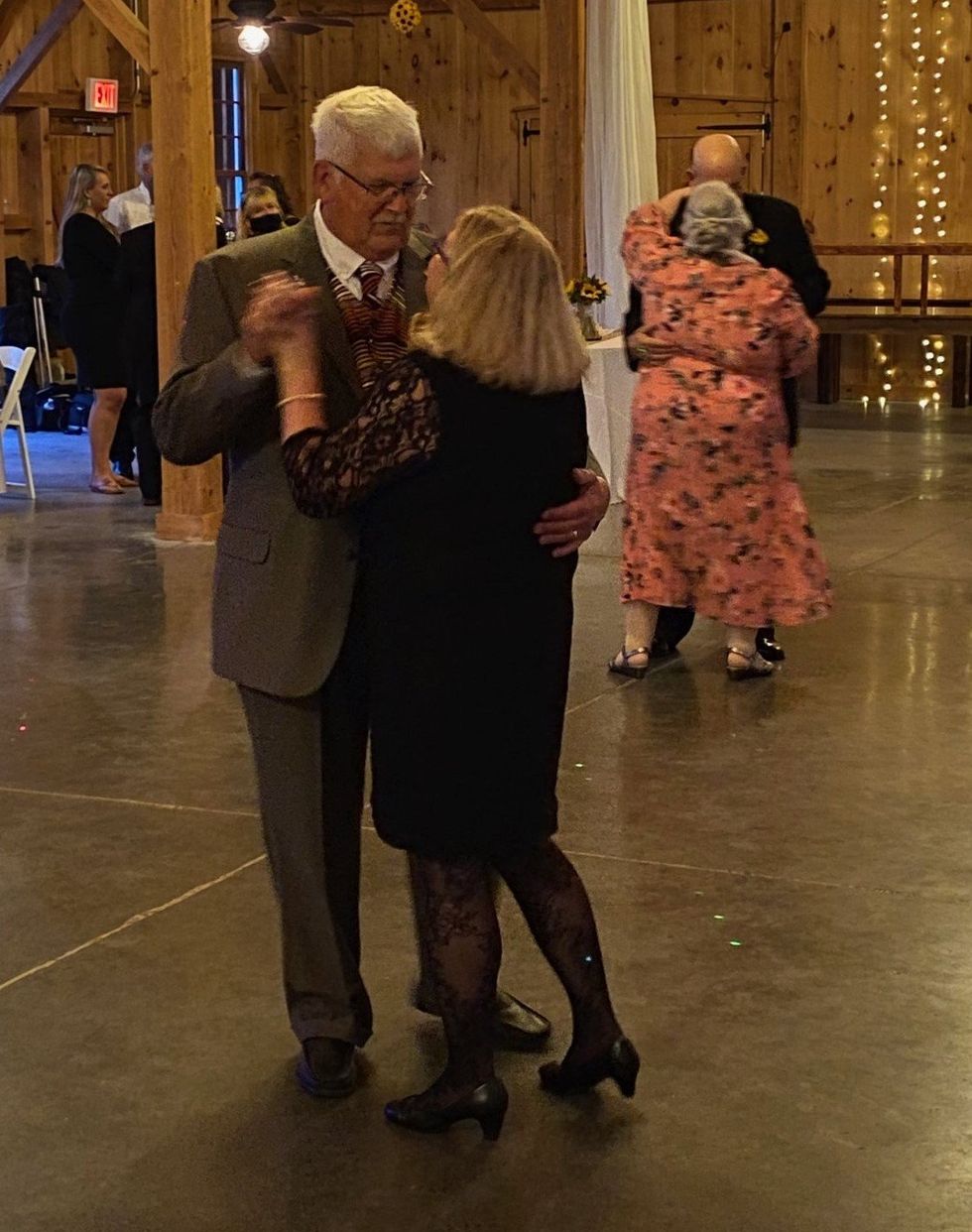

Dave Arnold and his wife dance at his nephew’s wedding in Rochester, New York, ten years ago, before his Alzheimer's diagnosis.
Dave Arnold
Ultimately, they want to integrate their tool into everyday life. “We want it to be used in a smartphone, or a teleconference call so that individuals could be examined in their home,” O’Connell said. “We could follow them over time and work with clinical teams and hospitals to improve the evaluation of patients and contribute towards an accurate diagnosis.”
Arnold, the patient with early stage Alzheimer’s, sees great promise. “The process of getting a diagnosis is already filled with so much anxiety,” he said. “Anything that can be done to make it easier and less stressful would be a good thing, as long as it’s proven accurate.”

