Scientists redesign bacteria to tackle the antibiotic resistance crisis
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
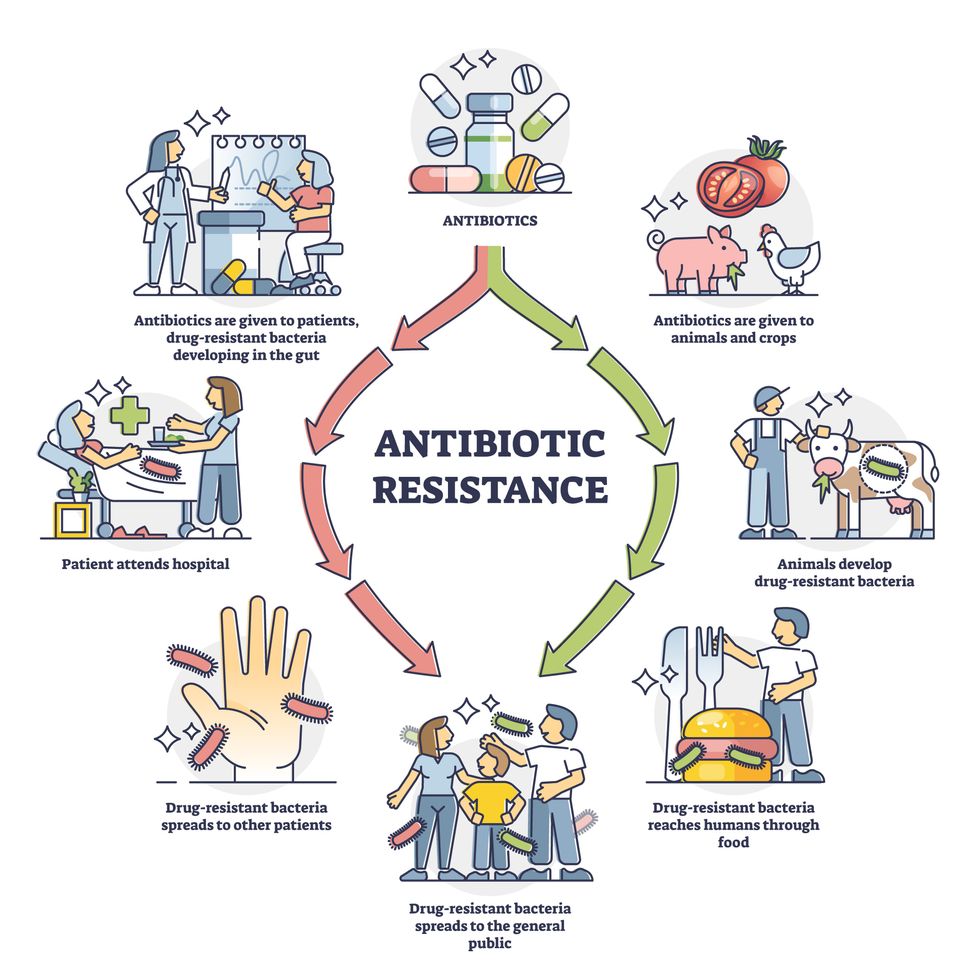
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
In this week's Friday Five: The eyes are the windows to the soul - and biological aging?
Plus, what bean genes mean for health and the planet, a breathing practice that could lower levels of tau proteins in the brain, AI beats humans at assessing heart health, and the benefits of "nature prescriptions"
The Friday Five covers five stories in research that you may have missed this week. There are plenty of controversies and troubling ethical issues in science – and we get into many of them in our online magazine – but this news roundup focuses on new scientific theories and progress to give you a therapeutic dose of inspiration headed into the weekend.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
Here are the stories covered this week:
- The eyes are the windows to the soul - and biological aging?
- What bean genes mean for health and the planet
- This breathing practice could lower levels of tau proteins
- AI beats humans at assessing heart health
- Should you get a nature prescription?
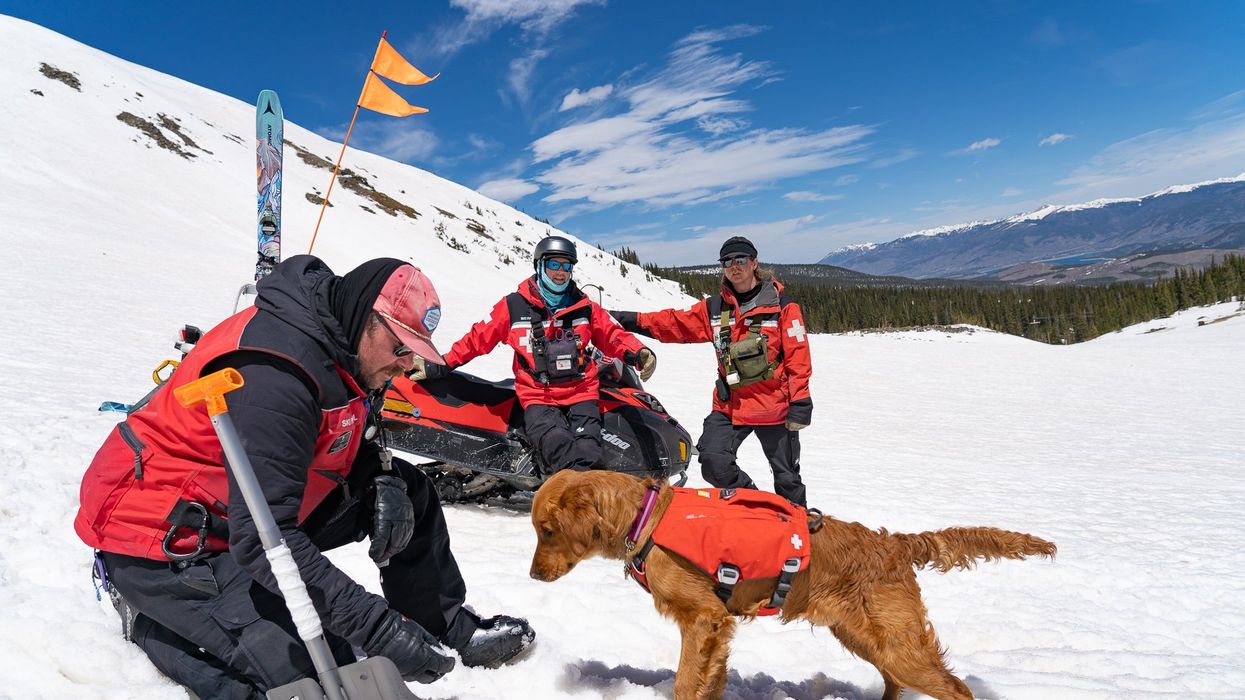
Avalanche rescue dogs train to find and dig out people buried in snow slides
Two-and-a-half year-old Huckleberry, a blue merle Australian shepherd, pulls hard at her leash; her yelps can be heard by skiers and boarders high above on the chairlift that carries them over the ski patrol hut to the top of the mountain. Huckleberry is an avalanche rescue dog — or avy dog, for short. She lives and works with her owner and handler, a ski patroller at Breckenridge Ski Resort in Colorado. As she watches the trainer play a game of hide-and-seek with six-month-old Lume, a golden retriever and avy dog-in-training, Huckleberry continues to strain on her leash; she loves the game. Hide-and-seek is one of the key training methods for teaching avy dogs the rescue skills they need to find someone caught in an avalanche — skier, snowmobiler, hiker, climber.
Lume’s owner waves a T-shirt in front of the puppy. While another patroller holds him back, Lume’s owner runs away and hides. About a minute later — after a lot of barking — Lume is released and commanded to “search.” He springs free, running around the hut to find his owner who reacts with a great amount of excitement and fanfare. Lume’s scent training will continue for the rest of the ski season (Breckenridge plans operating through May or as long as weather permits) and through the off-season. “We make this game progressively harder by not allowing the dog watch the victim run away,” explains Dave Leffler, Breckenridge's ski patroller and head of the avy dog program, who has owned, trained and raised many of them. Eventually, the trainers “dig an open hole in the snow to duck out of sight and gradually turn the hole into a cave where the dog has to dig to get the victim,” explains Leffler.
By the time he is three, Lume, like Huckleberry, will be a fully trained avy pup and will join seven other avy dogs on Breckenridge ski patrol team. Some of the team members, both human and canine, are also certified to work with Colorado Rapid Avalanche Deployment, a coordinated response team that works with the Summit County Sheriff’s office for avalanche emergencies outside of the ski slopes’ boundaries.
There have been 19 avalanche deaths in the U.S. this season, according to avalanche.org, which tracks slides; eight in Colorado. During the entirety of last season there were 17. Avalanche season runs from November through June, but avalanches can occur year-round.
High tech and high stakes
Complementing avy dogs’ ability to smell people buried in a slide, avalanche detection, rescue and recovery is becoming increasingly high tech. There are transceivers, signal locators, ground scanners and drones, which are considered “games changers” by many in avalanche rescue and recovery
For a person buried in an avalanche, the chance of survival plummets after 20 minutes, so every moment counts.
A drone can provide thermal imaging of objects caught in a slide; what looks like a rock from far away might be a human with a heat signature. Transceivers, also known as beacons, send a signal from an avalanche victim to a companion. Signal locators, like RECCO reflectors which are often sewn directly into gear, can echo back a radar signal sent by a detector; most ski resorts have RECCO detector units.
Research suggests that Ground Penetrating Radar (GPR), an electromagnetic tool used by geophysicists to pull images from inside the ground, could be used to locate an avalanche victim. A new study from the Department of Energy’s Sandia National Laboratories suggests that a computer program developed to pinpoint the source of a chemical or biological terrorist attack could also be used to find someone submerged in an avalanche. The search algorithm allows for small robots (described as cockroach-sized) to “swarm” a search area. Researchers say that this distributed optimization algorithm can help find avalanche victims four times faster than current search mechanisms. For a person buried in an avalanche, the chance of survival plummets after 20 minutes, so every moment counts.
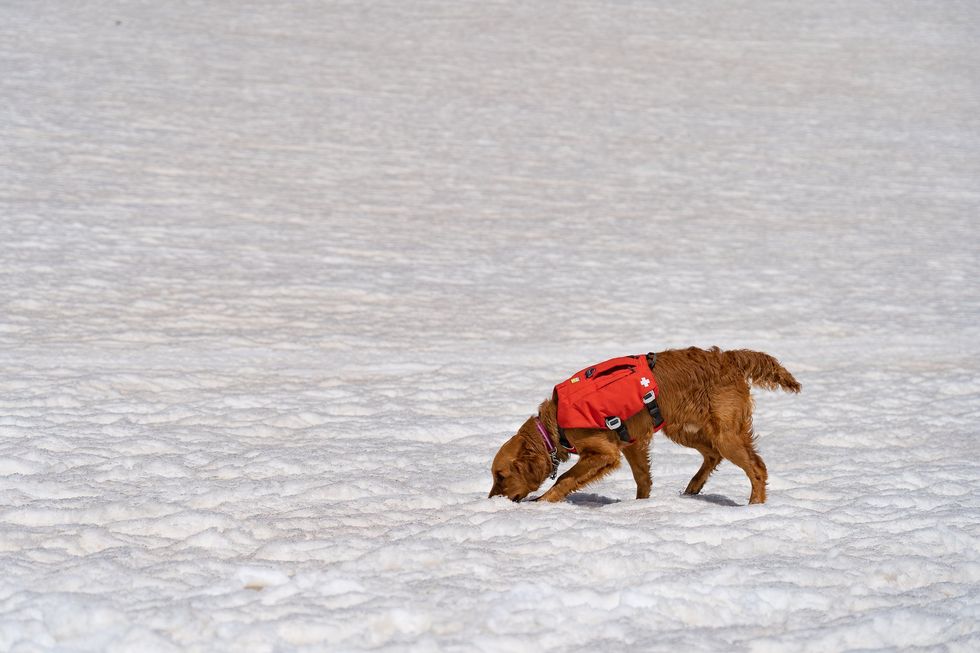

An avy dog in training is picking up scent
Sarah McLear
While rescue gear has been evolving, predicting when a slab will fall remains an emerging science — kind of where weather forecasting science was in the 1980s. Avalanche forecasting still relies on documenting avalanches by going out and looking,” says Ethan Greene, director of the Colorado Avalanche Information Center (CAIC). “So if there's a big snowstorm, and as you might remember, most avalanches happened during snowstorms, we could have 10,000 avalanches that release and we document 50,” says Greene. “Avalanche forecasting is essentially pattern recognition,” he adds--and understanding the layering structure of snow.
However, determining where the hazards lie can be tricky. While a dense layer of snow over a softer, weaker layer may be a recipe for an avalanche, there’s so much variability in snowpack that no one formula can predict the trigger. Further, observing and measuring snow at a single point may not be representative of all nearby slopes. Finally, there’s not enough historical data to help avalanche scientists create better prediction models.
That, however, may be changing.
Last year, an international group of researchers created computer simulations of snow cover using 16 years of meteorological data to forecast avalanche hazards, publishing their research in Cold Regions Science and Technology. They believe their models, which categorize different kinds of avalanches, can support forecasting and determine whether the avalanche is natural (caused by temperature changes, wind, additional snowfall) or artificial (triggered by a human or animal).
With smell receptors ranging from 800 million for an average dog, to 4 billion for scent hounds, canines remain key to finding people caught in slides.
With data from two sites in British Columbia and one in Switzerland, researchers built computer simulations of five different avalanche types. “In terms of real time avalanche forecasting, this has potential to fill in a lot of data gaps, where we don't have field observations of what the snow looks like,” says Simon Horton, a postdoctoral fellow with the Simon Fraser University Centre for Natural Hazards Research and a forecaster with Avalanche Canada, who participated in the study. While complex models that simulate snowpack layers have been around for a few decades, they weren’t easy to apply until recently. “It's been difficult to find out how to apply that to actual decision-making and improving safety,” says Horton. If you can derive avalanche problem types from simulated snowpack properties, he says, you’ll learn “a lot about how you want to manage that risk.”
The five categories include “new snow,” which is unstable and slides down the slope, “wet snow,” when rain or heat makes it liquidly, as well as “wind-drifted snow,” “persistent weak layers” and “old snow.” “That's when there's some type of deeply buried weak layer in the snow that releases without any real change in the weather,” Horton explains. “These ones tend to cause the most accidents.” One step by a person on that structurally weak layer of snow will cause a slide. Horton is hopeful that computer simulations of avalanche types can be used by scientists in different snow climates to help predict hazard levels.
Greene is doubtful. “If you have six slopes that are lined up next to each other, and you're going to try to predict which one avalanches and the exact dimensions and what time, that's going to be really hard to do. And I think it's going to be a long time before we're able to do that,” says Greene.
What both researchers do agree on, though, is that what avalanche prediction really needs is better imagery through satellite detection. “Just being able to count the number of avalanches that are out there will have a huge impact on what we do,” Greene says. “[Satellites] will change what we do, dramatically.” In a 2022 paper, scientists at the University of Aberdeen in England used satellites to study two deadly Himalayan avalanches. The imaging helped them determine that sediment from a 2016 ice avalanche plus subsequent snow avalanches contributed to the 2021 avalanche that caused a flash flood, killing over 200 people. The researchers say that understanding the avalanches characteristics through satellite imagery can inform them how one such event increases the magnitude of another in the same area.
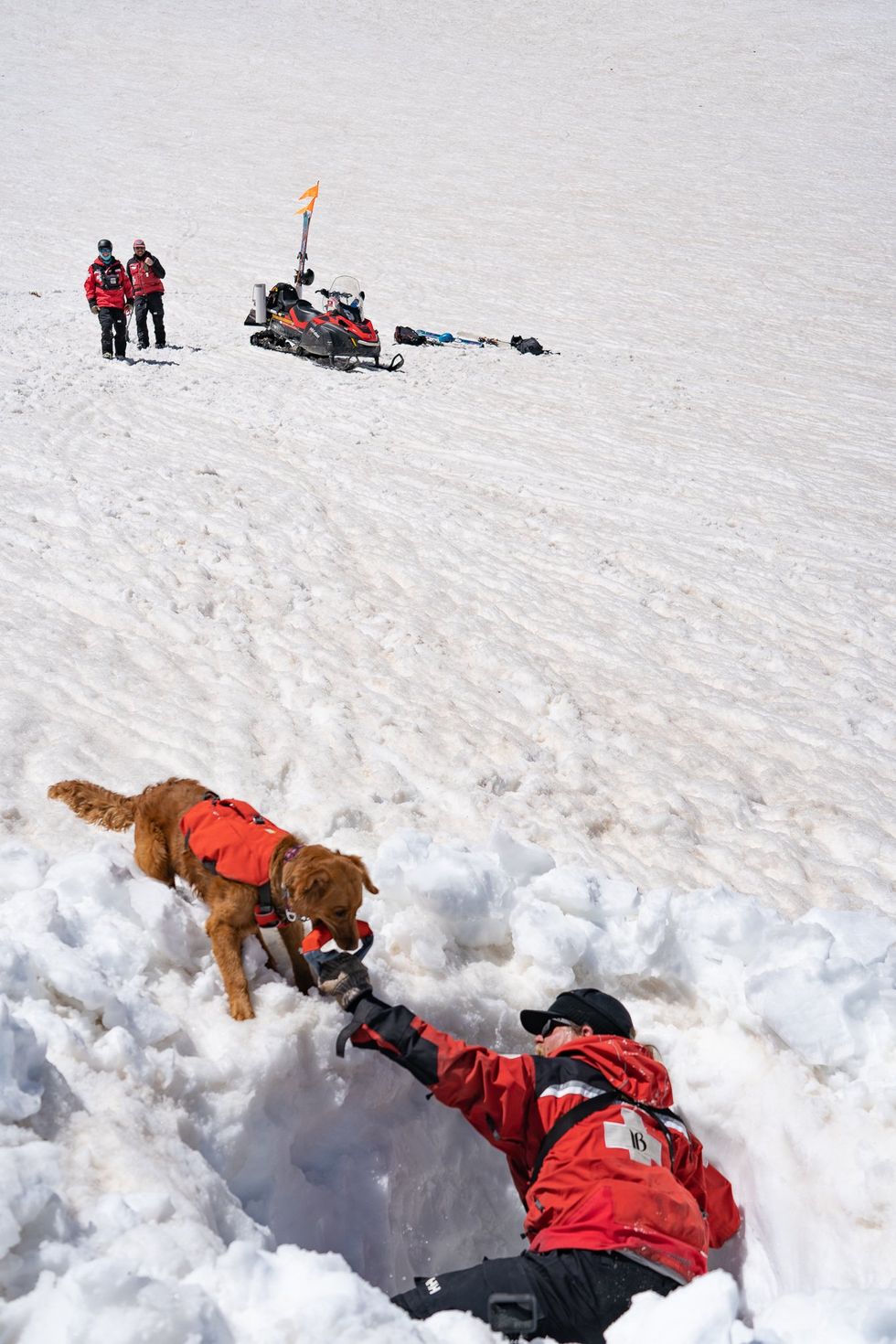

Avy dogs trainers hide in dug-out holes in the snow, teaching the dogs to find buried victims
Sarah McLear
Lifesaving combo: human tech and Mother Nature’s gear
Even as avalanche forecasting evolves, dogs with their built-in rescue mechanisms will remain invaluable. With smell receptors ranging from 800 million for an average dog, to 4 billion for scent hounds, canines remain key to finding people caught in slides. (Humans in comparison, have a meager 12 million.) A new study published in the Journal of Neuroscience revealed that in dogs smell and vision are connected in the brain, which has not been found in other animals. “They can detect the smell of their owner's fingerprints on a glass slide six weeks after they touched it,” says Nicholas Dodman, professor emeritus at Cummings School of Veterinary Medicine at Tufts University. “And they can track from a boat where a box filled with meat was buried in the water, 100 feet below,” says Dodman, who is also co-founder and president of the Center for Canine Behavior Studies.
Another recent study from Queens College in Belfast, United Kingdom, further confirms that dogs can smell when humans are stressed. They can also detect the smell of a person’s breath and the smell of the skin cells of a deceased person.
The emerging avalanche-predicting human-made tech and the incredible nature-made tech of dogs’ olfactory talents is the lifesaving “equipment” that Leffler believes in. Even when human-made technology develops further, it will be most efficient when used together with the millions of dogs’ smell receptors, Leffler believes. “It is a combination of technology and the avalanche dog that will always be effective in finding an avalanche victim.”