Medical Breakthroughs Set to be Fast-Tracked by Innovative New Health Agency
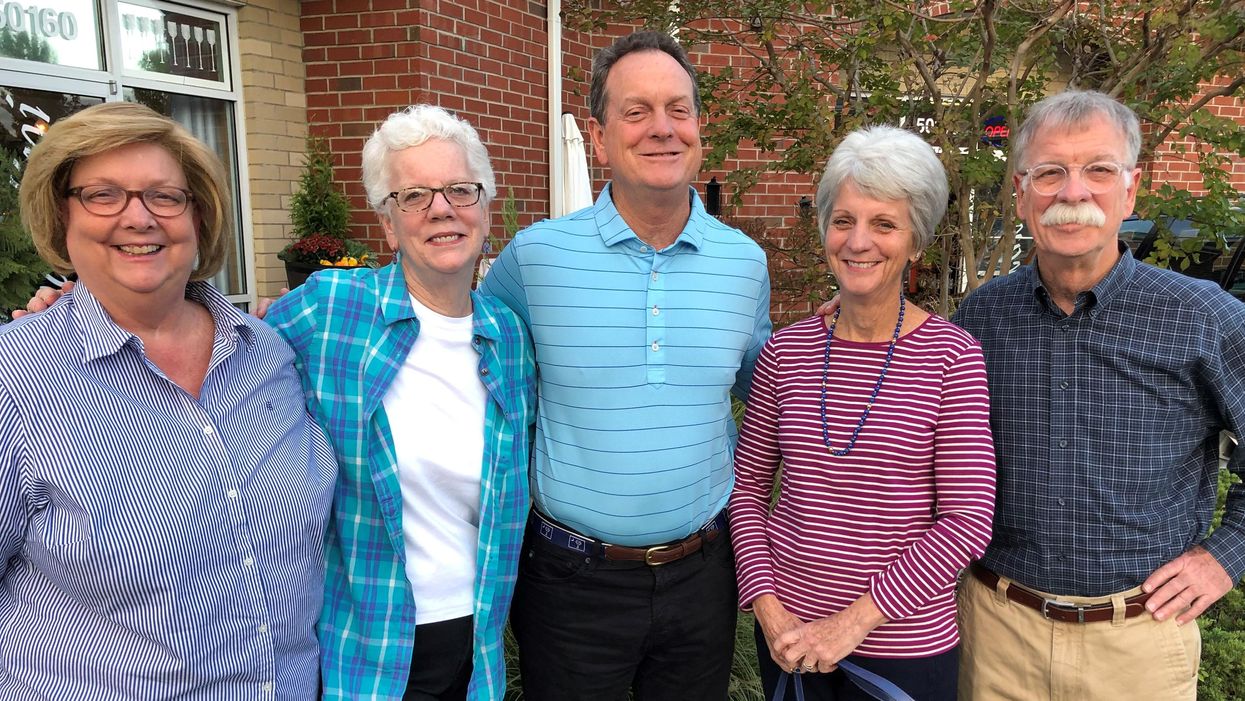

Pippy Rogers, second from left, with her four siblings, who worry that they are at risk for Alzheimer's and are calling for an acceleration of research.
In 2007, Matthew Might's son, Bertrand, was born with a life-threatening disease that was so rare, doctors couldn't diagnose it. Might, a computer scientist and biologist, eventually realized, "Oh my gosh, he's the only patient in the world with this disease right now." To find effective treatments, new methodologies would need to be developed. But there was no process or playbook for doing that.
Might took it upon himself, along with a team of specialists, to try to find a cure. "What Bertrand really taught me was the visceral sense of urgency when there's suffering, and how to act on that," he said.
He calls it "the agency of urgency"—and patients with more common diseases, such as cancer and Alzheimer's, often feel that same need to take matters into their own hands, as they find their hopes for new treatments running up against bureaucratic systems designed to advance in small, steady steps, not leaps and bounds. "We all hope for a cure," said Florence "Pippy" Rogers, a 65-year-old volunteer with Georgia's chapter of the Alzheimer's Association. She lost her mother to the disease and, these days, worries about herself and her four siblings. "We need to keep accelerating research."
We have a fresh example of what can be achieved by fast-tracking discoveries in healthcare: Covid-19 vaccines.
President Biden has pushed for cancer moonshots since the disease took the life of his son, Beau, in 2015. His administration has now requested $6.5 billion to start a new agency in 2022, called the Advanced Research Projects Agency for Health, or ARPA-H, within the National Institutes of Health. It's based on DARPA, the Department of Defense agency known for hatching world-changing technologies such as drones, GPS and ARPANET, which became the internet.
We have a fresh example of what can be achieved by fast-tracking discoveries in healthcare: Covid-19 vaccines. "Operation Warp Speed was using ARPA-like principles," said Might. "It showed that in a moment of crisis, institutions like NIH can think in an ARPA-like way. So now the question is, why don't we do that all the time?"
But applying the DARPA model to health involves several challenging decisions. I asked experts what could be the hardest question facing advocates of ARPA-H: which health problems it should seek to address. "All the wonderful choices lead to the problem of which ones to choose and prioritize," said Sudip Parikh, CEO of the American Association for the Advancement of Science and executive publisher of the Science family of journals. "There is no objectively right answer."
The Agency of Urgency
ARPA-H will borrow at least three critical ingredients from DARPA: goal-oriented project managers, many from industry; aggressive public-private partnerships; and collaboration among fields that don't always interact. The DARPA concept has been applied to other purposes, including energy and homeland security, with promising results. "We're learning that 'ARPA-ism' is a franchisable model," said Might, a former principal investigator on DARPA projects.
The federal government already pours billions of dollars into advancing research on life-threatening diseases, with much of it channeled through the National Institutes of Health. But the purpose of ARPA-H "isn't just the usual suspects that NIH would fund," said David Walt, a Harvard biochemist, an innovator in gene sequencing and former chair of DARPA's Defense Science Research Council. Whereas some NIH-funded studies aim to gradually improve our understanding of diseases, ARPA-H projects will give full focus to real-world applications; they'll use essential findings from NIH research as starting points, drawing from them to rapidly engineer new technologies that could save lives.
And, ultimately, billions in healthcare costs, if ARPA-H lives up to its predecessor's track record; DARPA's breakthroughs have been economic game-changers, while its fail-fast approach—quickly pulling the plug on projects that aren't panning out—helps to avoid sunken costs. ARPA-H could fuel activities similar to the human genome project, which used existing research to map the base pairs that make up DNA, opening new doors for the biotech industry, sparking economic growth and creating hundreds of thousands of new jobs.
Despite a nearly $4 trillion health economy, "we aren't innovating when it comes to technological capabilities for health," said Liz Feld, president of the Suzanne Wright Foundation for pancreatic cancer.
Individual Diseases Ripe for Innovation
Although the need for innovation is clear, which diseases ARPA-H should tackle is less apparent. One important consideration when choosing health priorities could be "how many people suffer from a disease," said Nancy Kass, a professor of bioethics and public health at Johns Hopkins.
That perspective could justify cancer as a top objective. Cancer and heart disease have long been the two major killers in the U.S. Leonidas Platanias, professor of oncology at Northwestern and director of its cancer center, noted that we've already made significant progress on heart disease. "Anti-cholesterol drugs really have a wide impact," he said. "I don't want to compare one disease to another, but I think cancer may be the most challenging. We need even bigger breakthroughs." He wondered whether ARPA-H should be linked to the part of NIH dedicated to cancer, the National Cancer Institute, "to take maximum advantage of what happens" there.
Previous cancer moonshots have laid a foundation for success. And this sort of disease-by-disease approach makes sense in a way. "We know that concentrating on some diseases has led to treatments," said Parikh. "Think of spinal muscular atrophy or cystic fibrosis. Now, imagine if immune therapies were discovered ten years earlier."
But many advocates think ARPA-H should choose projects that don't revolve around any one disease. "It absolutely has to be disease agnostic," said Feld, president of the pancreatic cancer foundation. "We cannot reach ARPA-H's potential if it's subject to the advocacy of individual patient groups who think their disease is worse than the guy's disease next to them. That's not the way the DARPA model works." Platanias agreed that ARPA-H should "pick the highest concepts and developments that have the best chance" of success.
Finding Connections Between Diseases
Kass, the Hopkins bioethicist, believes that ARPA-H should walk a balance, with some projects focusing on specific diseases and others aspiring to solutions with broader applications, spanning multiple diseases. Being impartial, some have noted, might involve looking at the total "life years" saved by a health innovation; the more diseases addressed by a given breakthrough, the more years of healthy living it may confer. The social and economic value should increase as well.
For multiple payoffs, ARPA-H could concentrate on rare diseases, which can yield important insights for many other diseases, said Might. Every case of cancer and Alzheimer's is, in a way, its own rare disease. Cancer is a genetic disease, like his son Bertrand's rare disorder, and mutations vary widely across cancer patients. "It's safe to say that no two people have ever actually had the same cancer," said Might. In theory, solutions for rare diseases could help us understand how to individualize treatments for more common diseases.
Many experts I talked with support another priority for ARPA-H with implications for multiple diseases: therapies that slow down the aging process. "Aging is the greatest risk factor for every major disease that NIH is studying," said Matt Kaeberlein, a bio-gerontologist at the University of Washington. Yet, "half of one percent of the NIH budget goes to researching the biology of aging. An ARPA-H sized budget would push the field forward at a pace that's hard to imagine."
Might agreed. "It could take ARPA-H to get past the weird stigmas around aging-related research. It could have a tremendous impact on the field."
For example, ARPA-H could try to use mRNA technology to express proteins that affect biological aging, said Kaeberlein. It's an engineering project well-suited to the DARPA model. So is harnessing machine learning to identify biomarkers that assess how fast people are aging. Biological aging clocks, if validated, could quickly reveal whether proposed therapies for aging are working or not. "I think there's huge value in that," said Kaeberlein.
By delivering breakthroughs in computation, ARPA-H could improve diagnostics for many different diseases. That could include improving biowearables for continuously monitoring blood pressure—a hypothetical mentioned in the White House's concept paper on ARPA-H—and advanced imaging technologies. "The high cost of medical imaging is a leading reason why our healthcare costs are the highest in the world," said Feld. "There's no detection test for ALS. No brain detection for Alzheimer's. Innovations in detection technology would save on cost and human suffering."
Some biotech companies may be skeptical about the financial rewards of accelerating such technologies. But ARPA-H could fund public-private partnerships to "de-risk" biotech's involvement—an incentive that harkens back to the advance purchase contracts that companies got during Covid. (Some groups have suggested that ARPA-H could provide advance purchase agreements.)
Parikh is less bullish on creating diagnostics through ARPA-H. Like DARPA, Biden's health agency will enjoy some independence from federal oversight; it may even be located hundreds of miles from DC. That freedom affords some breathing room for innovation, but it could also make it tougher to ensure that algorithms fully consider diverse populations. "That part I really would like the government more involved in," Parikh said.
Might thinks ARPA-H should also explore innovations in clinical trials, which many patients and medical communities view as grindingly slow and requiring too many participants. "We can approve drugs for very tiny patient populations, even at the level of the individual," he said, while emphasizing the need for safety. But Platanias thinks the FDA has become much more flexible in recent years. In the cancer field, at least, "You now see faster approvals for more drugs. Having [more] shortcuts on clinical trial approvals is not necessarily a good idea."
With so many options on the table, ARPA-H needs to show the public a clear framework for measuring the value of potential projects. Kass warned that well-resourced advocates could skew the agency's priorities. They've affected health outcomes before, she noted; fundraising may partly explain larger increases in life expectancy for cystic fibrosis than sickle cell anemia. Engaging diverse communities is a must for ARPA-H. So are partnerships to get the agency's outputs to people who need them. "Research is half the equation," said Kass. "If we don't ensure implementation and access, who cares." The White House concept paper on ARPA-H made a similar point.
As Congress works on authorizing ARPA-H this year, Might is doing what he can to ensure better access to innovation on a patient-by-patient basis. Last year, his son, Bertrand, passed away suddenly from his disorder. He was 12. But Might's sense of urgency has persisted, as he directs the Precision Medicine Institute at the University of Alabama-Birmingham. That urgency "can be carried into an agency like ARPA-H," he said. "It guides what I do as I apply for funding, because I'm trying to build the infrastructure that other parents need. So they don't have to build it from scratch like I did."
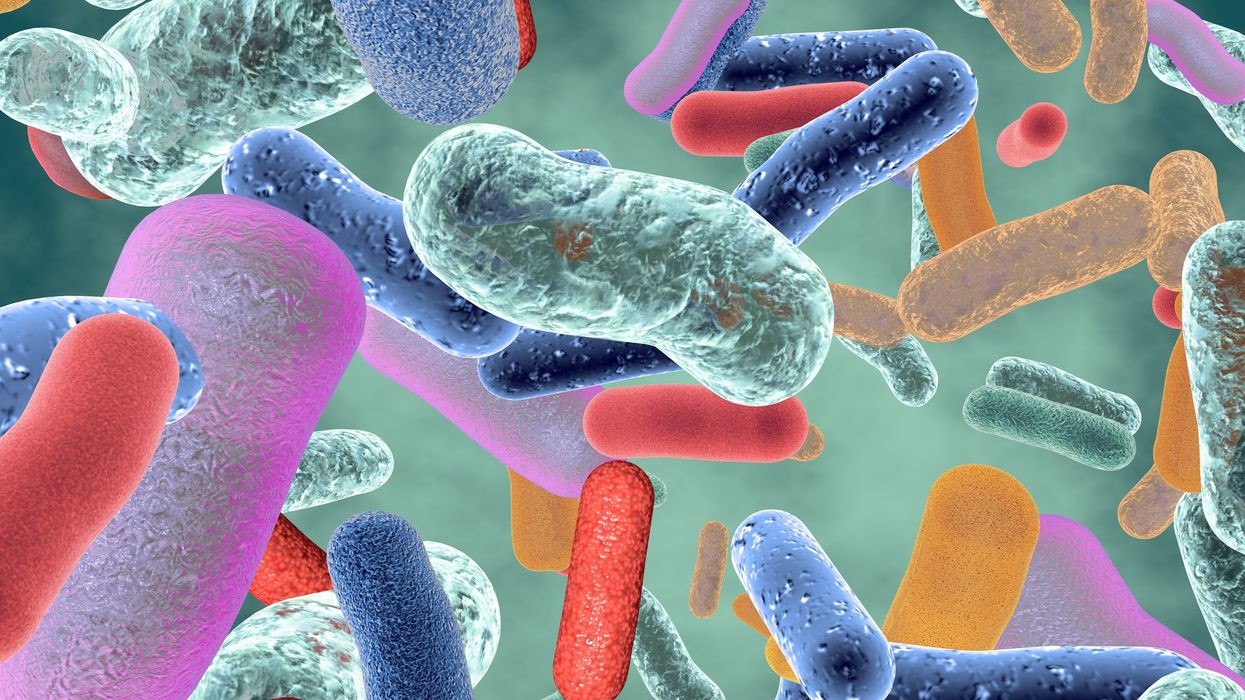
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
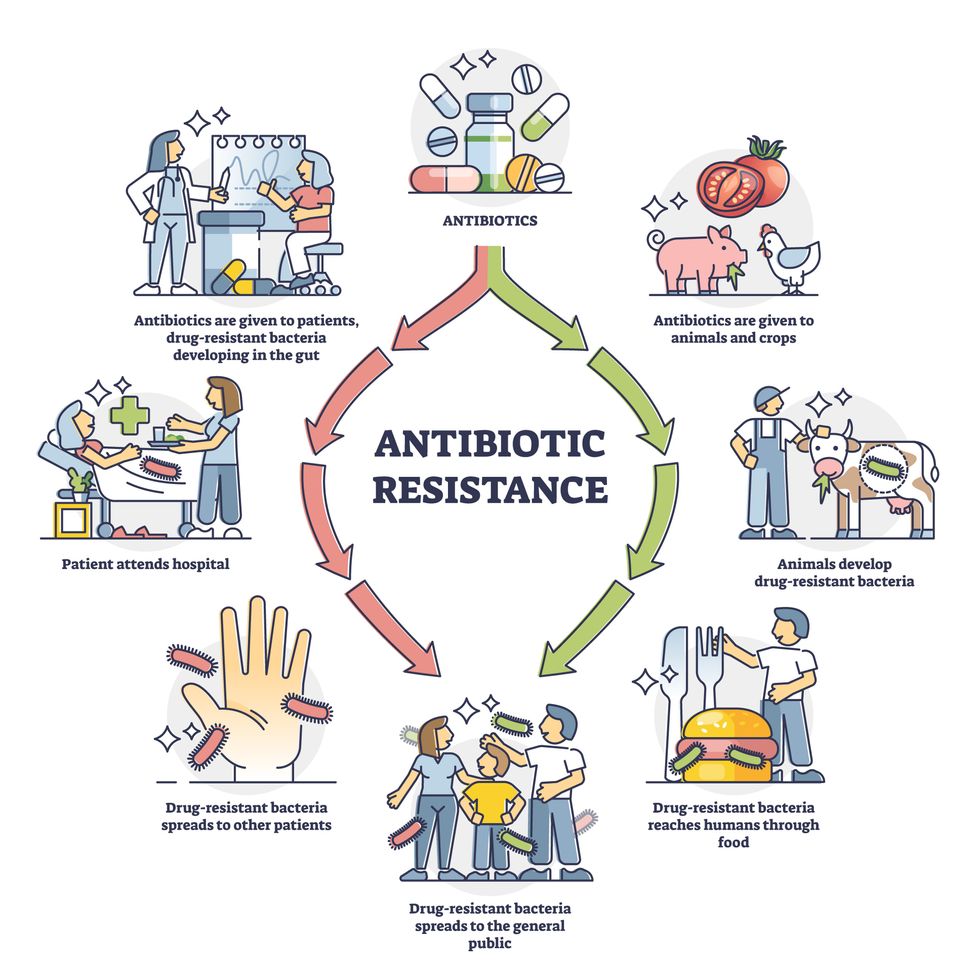
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
Meet Dr. Renee Wegrzyn, the first Director of President Biden's new health agency, ARPA-H
Today's podcast guest, Dr. Renee Wegrzyn, directs ARPA-H, a new agency formed last year to spearhead health innovations. Time will tell if ARPA-H will produce advances on the level of its fellow agency, DARPA.
In today’s podcast episode, I talk with Renee Wegrzyn, appointed by President Biden as the first director of a health agency created last year, the Advanced Research Projects Agency for Health, or ARPA-H. It’s inspired by DARPA, the agency that develops innovations for the Defense department and has been credited with hatching world-changing technologies such as ARPANET, which became the internet.
Time will tell if ARPA-H will lead to similar achievements in the realm of health. That’s what President Biden and Congress expect in return for funding ARPA-H at 2.5 billion dollars over three years.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
How will the agency figure out which projects to take on, especially with so many patient advocates for different diseases demanding moonshot funding for rapid progress?
I talked with Dr. Wegrzyn about the opportunities and challenges, what lessons ARPA-H is borrowing from Operation Warp Speed, how she decided on the first ARPA-H project that was announced recently, why a separate agency was needed instead of reforming HHS and the National Institutes of Health to be better at innovation, and how ARPA-H will make progress on disease prevention in addition to treatments for cancer, Alzheimer’s and diabetes, among many other health priorities.
Dr. Wegrzyn’s resume leaves no doubt of her suitability for this role. She was a program manager at DARPA where she focused on applying gene editing and synthetic biology to the goal of improving biosecurity. For her work there, she received the Superior Public Service Medal and, in case that wasn’t enough ARPA experience, she also worked at another ARPA that leads advanced projects in intelligence, called I-ARPA. Before that, she ran technical teams in the private sector working on gene therapies and disease diagnostics, among other areas. She has been a vice president of business development at Gingko Bioworks and headed innovation at Concentric by Gingko. Her training and education includes a PhD and undergraduate degree in applied biology from the Georgia Institute of Technology and she did her postdoc as an Alexander von Humboldt Fellow in Heidelberg, Germany.
Dr. Wegrzyn told me that she’s “in the hot seat.” The pressure is on for ARPA-H especially after the need and potential for health innovation was spot lit by the pandemic and the unprecedented speed of vaccine development. We'll soon find out if ARPA-H can produce gamechangers in health that are equivalent to DARPA’s creation of the internet.
Show links:
ARPA-H - https://arpa-h.gov/
Dr. Wegrzyn profile - https://arpa-h.gov/people/renee-wegrzyn/
Dr. Wegrzyn Twitter - https://twitter.com/rwegrzyn?lang=en
President Biden Announces Dr. Wegrzyn's appointment - https://www.whitehouse.gov/briefing-room/statement...
Leaps.org coverage of ARPA-H - https://leaps.org/arpa/
ARPA-H program for joints to heal themselves - https://arpa-h.gov/news/nitro/ -
ARPA-H virtual talent search - https://arpa-h.gov/news/aco-talent-search/

Dr. Renee Wegrzyn was appointed director of ARPA-H last October.