The Brave New World of Using DNA to Store Data

A panoramic view of DNA
Netscape co-founder-turned-venture capitalist billionaire investor Marc Andreessen once posited that software was eating the world. He was right, and the takeover of software resulted in many things. One of them is data. Lots and lots and lots of data. In the previous two years, humanity created more data than it did during its entire existence combined, and the amount will only increase. Think about it: The hundreds of 50KB emails you write a day, the dozens of 10MB photos, the minute-long, 350MB 4K video you shoot on your iPhone X add up to vast quantities of information. All that information needs to be stored. And that's becoming an issue as data volume outpaces storage space.
The race is on to find another medium capable of storing massive amounts of information in as small a space as possible.
"There won't be enough silicon to store all the data we need. It's unlikely that we can make flash memory smaller. We have reached the physical limits," Victor Zhirnov, chief scientist at the Semiconductor Research Corporation, says. "We are facing a crisis that's comparable to the oil crisis in the 1970s. By 2050, we're going to need to store 10 to the 30 bits, compared to 10 to the 23 bits in 2016." That amount of storage space is equivalent to each of the world's seven billion people owning almost six trillion -- that's 10 to the 12th power -- iPhone Xs with 256GB storage space.
The race is on to find another medium capable of storing massive amounts of information in as small a space as possible. Zhirnov and other scientists are looking at the human body, looking to DNA. "Nature has nailed it," Luis Ceze, a professor in the Department of Computer Science and Engineering at the University of Washington, says. "DNA is a molecular storage medium that is remarkable. It's incredibly dense, many, many thousands of times denser than the densest technology that we have today. And DNA is remarkably general. Any information you can map in bits you can store in DNA." It's so dense -- able to store a theoretical maximum of 215 petabytes (215 million gigabytes) in a single gram -- that all the data ever produced could be stored in the back of a tractor trailer truck.
Writing DNA can be an energy-efficient process, too. Consider how the human body is constantly writing and rewriting DNA, and does so on a couple thousand calories a day. And all it needs for storage is a cool, dark place, a significant energy savings when compared to server farms that require huge amounts of energy to run and even more energy to cool.
Picture it: tiny specks of inert DNA made from silicon or another material, stored in cool, dark, dry areas, preserved for all time.
Researchers first succeeded in encoding data onto DNA in 2012, when Harvard University geneticists George Church and Sri Kosuri wrote a 52,000-word book on A, C, G, and T base pairs. Their method only produced 1.28 petabytes per gram of DNA, however, a volume exceeded the next year when a group encoded all 154 Shakespeare sonnets and a 26-second clip of Martin Luther King's "I Have A Dream" speech. In 2017, Columbia University researchers Yaniv Erlich and Dina Zielinski made the process 60 percent more efficient.
The limiting factor today is cost. Erlich said the work his team did cost $7,000 to encode and decode two megabytes of data. To become useful in a widespread way, the price per megabyte needs to plummet. Even advocates concede this point. "Of course it is expensive," Zhirnov says. "But look how much magnetic storage cost in the 1980s. What you store today in your iPhone for virtually nothing would cost many millions of dollars in 1982." There's reason to think the price will continue to fall. Genome readers are improving, getting cheaper, faster, and smaller, and genome sequencing becomes cheaper every year, too. Picture it: tiny specks of inert DNA made from silicon or another material, stored in cool, dark, dry areas, preserved for all time.
"It just takes a few minutes to double a sample. A few more minutes, you double it again. Very quickly, you have thousands or millions of new copies."
Plus, DNA has another advantage over more traditional forms of storage: It's very easy to reproduce. "If you want a second copy of a hard disk drive, you need components for a disk drive, hook both drives up to a computer, and copy. That's a pain," Nick Goldman, a researcher at the European Bioinformatics Institute, says. "DNA, once you have that first sample, it's a process that is absolutely routine in thousands of laboratories around the world to multiply that using polymerase chain reaction [which uses temperature changes or other processes]. It just takes a few minutes to double a sample. A few more minutes, you double it again. Very quickly, you have thousands or millions of new copies."
This ability to duplicate quickly and easily is a positive trait. But, of course, there's also the potential for danger. Does encoding on DNA, the very basis for life, present ethical issues? Could it get out of control and fundamentally alter life as we know it?
The chance is there, but it's remote. The first reason is that storage could be done with only two base pairs, which would serve as replacements for the 0 and 1 digits that make up all digital data. While doing so would decrease the possible density of the storage, it would virtually eliminate the risk that the sequences would be compatible with life.
But even if scientists and researchers choose to use four base pairs, other safeguards are in place that will prevent trouble. According to Ceze, the computer science professor, the snippets of DNA that they write are very short, around 150 nucleotides. This includes the title, the information that's being encoded, and tags to help organize where the snippet should fall in the larger sequence. Furthermore, they generally avoid repeated letters, which dramatically reduces the chance that a protein could be synthesized from the snippet.
"In the future, we'll know enough about someone from a sample of their DNA that we could make a specific poison. That's the danger, not those of us who want to encode DNA for storage."
Inevitably, some DNA will get spilt. "But it's so unlikely that anything that gets created for storage would have a biological interpretation that could interfere with the mechanisms going on in a living organism that it doesn't worry me in the slightest," Goldman says. "We're not of concern for the people who are worried about the ethical issues of synthetic DNA. They are much more concerned about people deliberately engineering anthrax. In the future, we'll know enough about someone from a sample of their DNA that we could make a specific poison. That's the danger, not those of us who want to encode DNA for storage."
In the end, the reality of and risks surrounding encoding on DNA are the same as any scientific advancement: It's another system that is vulnerable to people with bad intentions but not one that is inherently unethical.
"Every human action has some ethical implications," Zhirnov says. "I can use a hammer to build a house or I can use it to harm another person. I don't see why DNA is in any way more or less ethical."
If that house can store all the knowledge in human history, it's worth learning how to build it.
Editor's Note: In response to readers' comments that silicon is one of the earth's most abundant materials, we reached back out to our source, Dr. Victor Zhirnov. He stands by his statement about a coming shortage of silicon, citing this research. The silicon oxide found in beach sand is unsuitable for semiconductors, he says, because the cost of purifying it would be prohibitive. For use in circuit-making, silicon must be refined to a purity of 99.9999999 percent. So the process begins by mining for pure quartz, which can only be found in relatively few places around the world.
Indigenous wisdom plus honeypot ants could provide new antibiotics
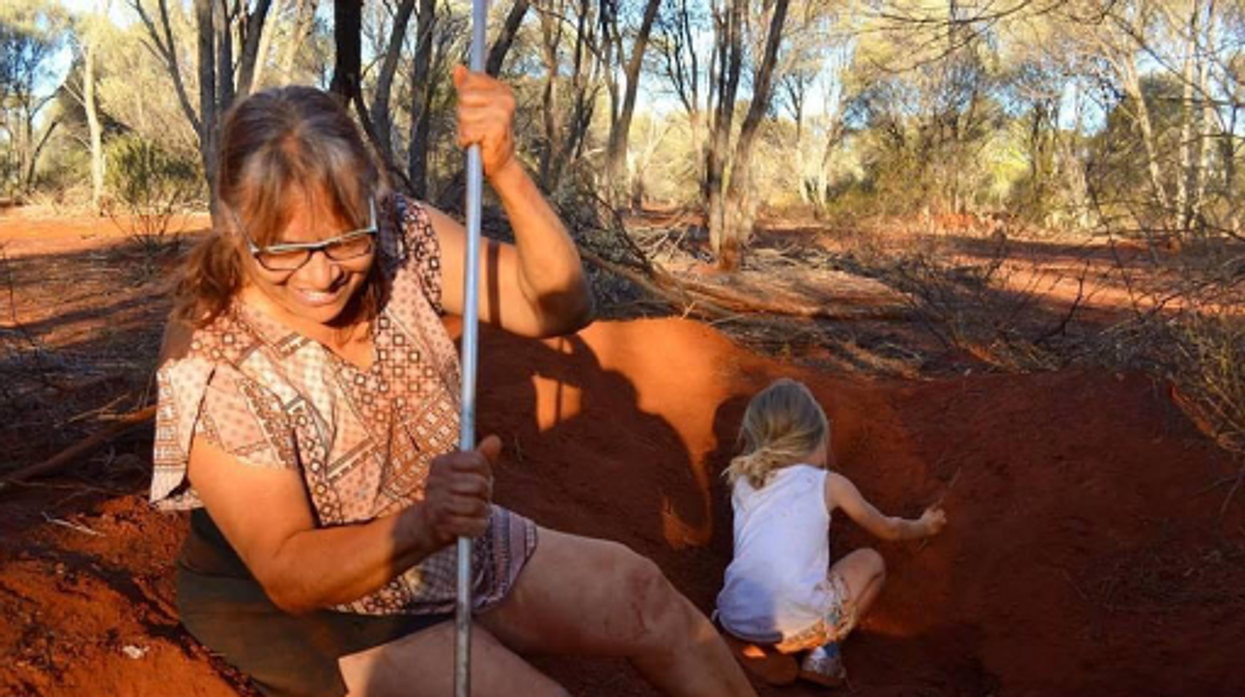
Indigenous people in Australia dig pits next to a honeypot colony. Scientists think the honey can be used to make new antimicrobial drugs.
For generations, the Indigenous Tjupan people of Australia enjoyed the sweet treat of honey made by honeypot ants. As a favorite pastime, entire families would go searching for the underground colonies, first spotting a worker ant and then tracing it to its home. The ants, which belong to the species called Camponotus inflatus, usually build their subterranean homes near the mulga trees, Acacia aneura. Having traced an ant to its tree, it would be the women who carefully dug a pit next to a colony, cautious not to destroy the entire structure. Once the ant chambers were exposed, the women would harvest a small amount to avoid devastating the colony’s stocks—and the family would share the treat.
The Tjupan people also knew that the honey had antimicrobial properties. “You could use it for a sore throat,” says Danny Ulrich, a member of the Tjupan nation. “You could also use it topically, on cuts and things like that.”
These hunts have become rarer, as many of the Tjupan people have moved away and, up until now, the exact antimicrobial properties of the ant honey remained unknown. But recently, scientists Andrew Dong and Kenya Fernandes from the University of Sydney, joined Ulrich, who runs the Honeypot Ants tours in Kalgoorlie, a city in Western Australia, on a honey-gathering expedition. Afterwards, they ran a series of experiments analyzing the honey’s antimicrobial activity—and confirmed that the Indigenous wisdom was true. The honey was effective against Staphylococcus aureus, a common pathogen responsible for sore throats, skin infections like boils and sores, and also sepsis, which can result in death. Moreover, the honey also worked against two species of fungi, Cryptococcus and Aspergillus, which can be pathogenic to humans, especially those with suppressed immune systems.
In the era of growing antibiotic resistance and the rising threat of pathogenic fungi, these findings may help scientists identify and make new antimicrobial compounds. “Natural products have been honed over thousands and millions of years by nature and evolution,” says Fernandes. “And some of them have complex and intricate properties that make them really important as potential new antibiotics. “
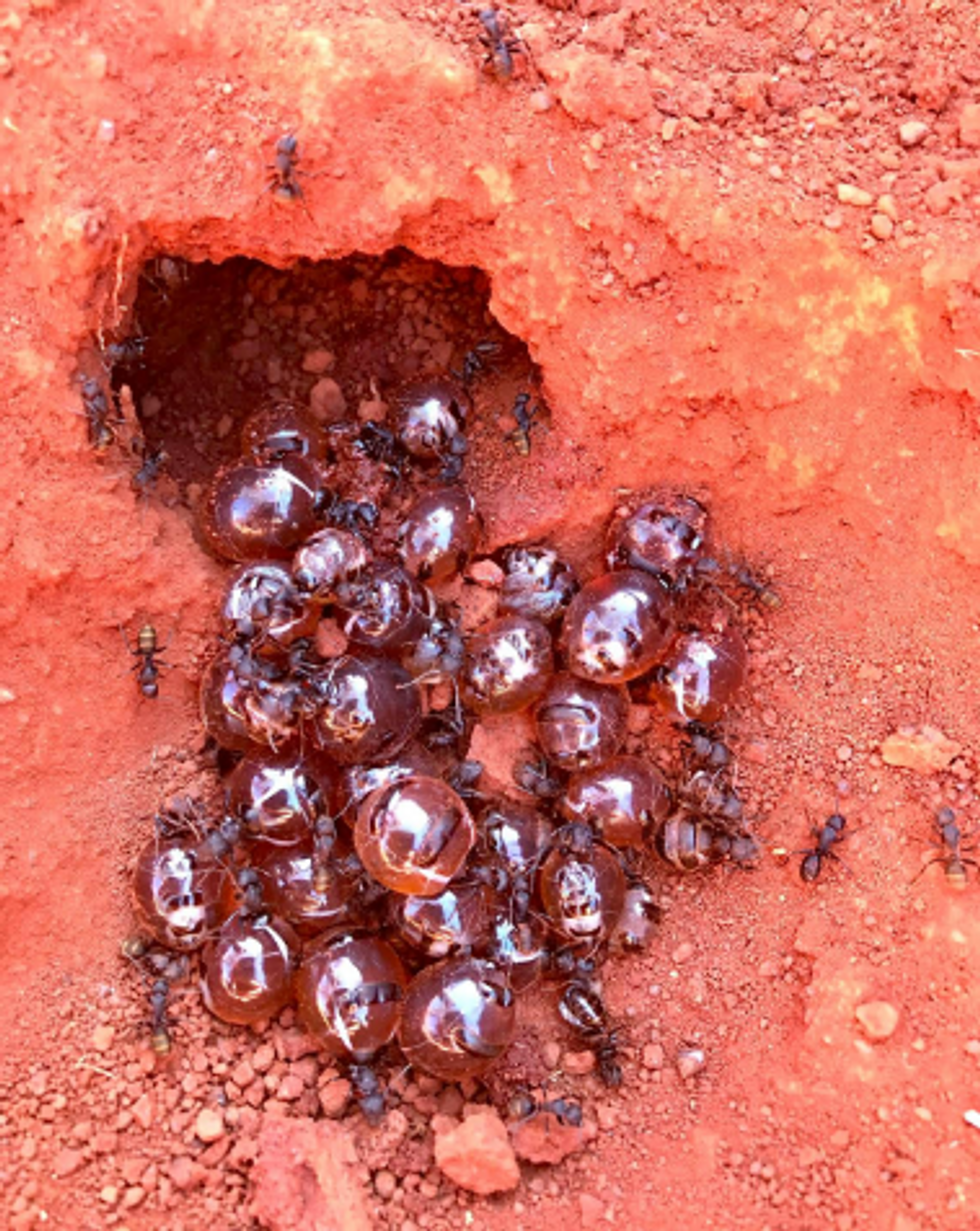

In an era of growing resistance to antibiotics and new threats of fungi infections, the latest findings about honeypot ants are helping scientists identify new antimicrobial drugs.
Danny Ulrich
Bee honey is also known for its antimicrobial properties, but bees produce it very differently than the ants. Bees collect nectar from flowers, which they regurgitate at the hive and pack into the hexagonal honeycombs they build for storage. As they do so, they also add into the mix an enzyme called glucose oxidase produced by their glands. The enzyme converts atmospheric oxygen into hydrogen peroxide, a reactive molecule that destroys bacteria and acts as a natural preservative. After the bees pack the honey into the honeycombs, they fan it with their wings to evaporate the water. Once a honeycomb is full, the bees put a beeswax cover on it, where it stays well-preserved thanks to the enzymatic action, until the bees need it.
Less is known about the chemistry of ants’ honey-making. Similarly to bees, they collect nectar. They also collect the sweet sap of the mulga tree. Additionally, they also “milk” the aphids—small sap-sucking insects that live on the tree. When ants tickle the aphids with their antennae, the latter release a sweet substance, which the former also transfer to their colonies. That’s where the honey management difference becomes really pronounced. The ants don’t build any kind of structures to store their honey. Instead, they store it in themselves.
The workers feed their harvest to their fellow ants called repletes, stuffing them up to the point that their swollen bellies outgrow the ants themselves, looking like amber-colored honeypots—hence the name. Because of their size, repletes don’t move, but hang down from the chamber’s ceiling, acting as living feedstocks. When food becomes scarce, they regurgitate their reserves to their colony’s brethren. It’s not clear whether the repletes die afterwards or can be restuffed again. “That's a good question,” Dong says. “After they've been stretched, they can't really return to exactly the same shape.”
These replete ants are the “treat” the Tjupan women dug for. Once they saw the round-belly ants inside the chambers, they would reach in carefully and get a few scoops of them. “You see a lot of honeypot ants just hanging on the roof of the little openings,” says Ulrich’s mother, Edie Ulrich. The women would share the ants with family members who would eat them one by one. “They're very delicate,” shares Edie Ulrich—you have to take them out carefully, so they don’t accidentally pop and become a wasted resource. “Because you’d lose all this precious honey.”
Dong stumbled upon the honeypot ants phenomenon because he was interested in Indigenous foods and went on Ulrich’s tour. He quickly became fascinated with the insects and their role in the Indigenous culture. “The honeypot ants are culturally revered by the Indigenous people,” he says. Eventually he decided to test out the honey’s medicinal qualities.
The researchers were surprised to see that even the smallest, eight percent concentration of honey was able to arrest the growth of S. aureus.
To do this, the two scientists first diluted the ant honey with water. “We used something called doubling dilutions, which means that we made 32 percent dilutions, and then we halve that to 16 percent and then we half that to eight percent,” explains Fernandes. The goal was to obtain as much results as possible with the meager honey they had. “We had very, very little of the honeypot ant honey so we wanted to maximize the spectrum of results we can get without wasting too much of the sample.”
After that, the researchers grew different microbes inside a nutrient rich broth. They added the broth to the different honey dilutions and incubated the mixes for a day or two at the temperature favorable to the germs’ growth. If the resulting solution turned turbid, it was a sign that the bugs proliferated. If it stayed clear, it meant that the honey destroyed them. The researchers were surprised to see that even the smallest, eight percent concentration of honey was able to arrest the growth of S. aureus. “It was really quite amazing,” Fernandes says. “Eight milliliters of honey in 92 milliliters of water is a really tiny amount of honey compared to the amount of water.”
Similar to bee honey, the ants’ honey exhibited some peroxide antimicrobial activity, researchers found, but given how little peroxide was in the solution, they think the honey also kills germs by a different mechanism. “When we measured, we found that [the solution] did have some hydrogen peroxide, but it didn't have as much of it as we would expect based on how active it was,” Fernandes says. “Whether this hydrogen peroxide also comes from glucose oxidase or whether it's produced by another source, we don't really know,” she adds. The research team does have some hypotheses about the identity of this other germ-killing agent. “We think it is most likely some kind of antimicrobial peptide that is actually coming from the ant itself.”
The honey also has a very strong activity against the two types of fungi, Cryptococcus and Aspergillus. Both fungi are associated with trees and decaying leaves, as well as in the soils where ants live, so the insects likely have evolved some natural defense compounds, which end up inside the honey.
It wouldn’t be the first time when modern medicines take their origin from the natural world or from the indigenous people’s knowledge. The bark of the cinchona tree native to South America contains quinine, a substance that treats malaria. The Indigenous people of the Andes used the bark to quell fever and chills for generations, and when Europeans began to fall ill with malaria in the Amazon rainforest, they learned to use that medicine from the Andean people.
The wonder drug aspirin similarly takes its origin from a bark of a tree—in this case a willow.
Even some anticancer compounds originated from nature. A chemotherapy drug called Paclitaxel, was originally extracted from the Pacific yew trees, Taxus brevifolia. The samples of the Pacific yew bark were first collected in 1962 by researchers from the United States Department of Agriculture who were looking for natural compounds that might have anti-tumor activity. In December 1992, the FDA approved Paclitaxel (brand name Taxol) for the treatment of ovarian cancer and two years later for breast cancer.
In the era when the world is struggling to find new medicines fast enough to subvert a fungal or bacterial pandemic, these discoveries can pave the way to new therapeutics. “I think it's really important to listen to indigenous cultures and to take their knowledge because they have been using these sources for a really, really long time,” Fernandes says. Now we know it works, so science can elucidate the molecular mechanisms behind it, she adds. “And maybe it can even provide a lead for us to develop some kind of new treatments in the future.”
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.
Blood Test Can Detect Lymphoma Cells Before a Tumor Grows Back
David Kurtz making DNA sequencing libraries in his lab.
When David M. Kurtz was doing his clinical fellowship at Stanford University Medical Center in 2009, specializing in lymphoma treatments, he found himself grappling with a question no one could answer. A typical regimen for these blood cancers prescribed six cycles of chemotherapy, but no one knew why. "The number seemed to be drawn out of a hat," Kurtz says. Some patients felt much better after just two doses, but had to endure the toxic effects of the entire course. For some elderly patients, the side effects of chemo are so harsh, they alone can kill. Others appeared to be cancer-free on the CT scans after the requisite six but then succumbed to it months later.
"Anecdotally, one patient decided to stop therapy after one dose because he felt it was so toxic that he opted for hospice instead," says Kurtz, now an oncologist at the center. "Five years down the road, he was alive and well. For him, just one dose was enough." Others would return for their one-year check up and find that their tumors grew back. Kurtz felt that while CT scans and MRIs were powerful tools, they weren't perfect ones. They couldn't tell him if there were any cancer cells left, stealthily waiting to germinate again. The scans only showed the tumor once it was back.
Blood cancers claim about 68,000 people a year, with a new diagnosis made about every three minutes, according to the Leukemia Research Foundation. For patients with B-cell lymphoma, which Kurtz focuses on, the survival chances are better than for some others. About 60 percent are cured, but the remaining 40 percent will relapse—possibly because they will have a negative CT scan, but still harbor malignant cells. "You can't see this on imaging," says Michael Green, who also treats blood cancers at University of Texas MD Anderson Medical Center.
The new blood test is sensitive enough to spot one cancerous perpetrator amongst one million other DNA molecules.
Kurtz wanted a better diagnostic tool, so he started working on a blood test that could capture the circulating tumor DNA or ctDNA. For that, he needed to identify the specific mutations typical for B-cell lymphomas. Working together with another fellow PhD student Jake Chabon, Kurtz finally zeroed-in on the tumor's genetic "appearance" in 2017—a pair of specific mutations sitting in close proximity to each other—a rare and telling sign. The human genome contains about 3 billion base pairs of nucleotides—molecules that compose genes—and in case of the B-cell lymphoma cells these two mutations were only a few base pairs apart. "That was the moment when the light bulb went on," Kurtz says.
The duo formed a company named Foresight Diagnostics, focusing on taking the blood test to the clinic. But knowing the tumor's mutational signature was only half the process. The other was fishing the tumor's DNA out of patients' bloodstream that contains millions of other DNA molecules, explains Chabon, now Foresight's CEO. It would be like looking for an escaped criminal in a large crowd. Kurtz and Chabon solved the problem by taking the tumor's "mug shot" first. Doctors would take the biopsy pre-treatment and sequence the tumor, as if taking the criminal's photo. After treatments, they would match the "mug shot" to all DNA molecules derived from the patient's blood sample to see if any molecular criminals managed to escape the chemo.
Foresight isn't the only company working on blood-based tumor detection tests, which are dubbed liquid biopsies—other companies such as Natera or ArcherDx developed their own. But in a recent study, the Foresight team showed that their method is significantly more sensitive in "fishing out" the cancer molecules than existing tests. Chabon says that this test can detect circulating tumor DNA in concentrations that are nearly 100 times lower than other methods. Put another way, it's sensitive enough to spot one cancerous perpetrator amongst one million other DNA molecules.
They also aim to extend their test to detect other malignancies such as lung, breast or colorectal cancers.
"It increases the sensitivity of detection and really catches most patients who are going to progress," says Green, the University of Texas oncologist who wasn't involved in the study, but is familiar with the method. It would also allow monitoring patients during treatment and making better-informed decisions about which therapy regimens would be most effective. "It's a minimally invasive test," Green says, and "it gives you a very high confidence about what's going on."
Having shown that the test works well, Kurtz and Chabon are planning a new trial in which oncologists would rely on their method to decide when to stop or continue chemo. They also aim to extend their test to detect other malignancies such as lung, breast or colorectal cancers. The latest genome sequencing technologies have sequenced and catalogued over 2,500 different tumor specimens and the Foresight team is analyzing this data, says Chabon, which gives the team the opportunity to create more molecular "mug shots."
The team hopes that that their blood cancer test will become available to patients within about five years, making doctors' job easier, and not only at the biological level. "When I tell patients, "good news, your cancer is in remission', they ask me, 'does it mean I'm cured?'" Kurtz says. "Right now I can't answer this question because I don't know—but I would like to." His company's test, he hopes, will enable him to reply with certainty. He'd very much like to have the power of that foresight.
This article is republished from our archives to coincide with Blood Cancer Awareness Month, which highlights progress in cancer diagnostics and treatment.
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.