Can You Trust Your Gut for Food Advice?
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.

Which foods are actually healthy for your individual gut microbiome? Several companies are offering personalized dietary guidance based on your test results, but their answers in one experiment turned up with some conflicting advice.
I recently got on the scale to weigh myself, thinking I've got to eat better. With so many trendy diets today claiming to improve health, from Keto to Paleo to Whole30, it can be confusing to figure out what we should and shouldn't eat for optimal nutrition.
A number of companies are now selling the concept of "personalized" nutrition based on the genetic makeup of your individual gut bugs.
My next thought was: I've got to lose a few pounds.
Consider a weird factoid: In addition to my fat, skin, bone and muscle, I'm carrying around two or three pounds of straight-up bacteria. Like you, I am the host to trillions of micro-organisms that live in my gut and are collectively known as my microbiome. An explosion of research has occurred in the last decade to try to understand exactly how these microbial populations, which are unique to each of us, may influence our overall health and potentially even our brains and behavior.
Lots of mysteries still remain, but it is established that these "bugs" are crucial to keeping our body running smoothly, performing functions like stimulating the immune system, synthesizing important vitamins, and aiding digestion. The field of microbiome science is evolving rapidly, and a number of companies are now selling the concept of "personalized" nutrition based on the genetic makeup of your individual gut bugs. The two leading players are Viome and DayTwo, but the landscape includes the newly launched startup Onegevity Health and others like Thryve, which offers customized probiotic supplements in addition to dietary recommendations.
The idea has immediate appeal – if science could tell you exactly what to make for lunch and what to avoid, you could forget about the fad diets and go with your own bespoke food pyramid. Wondering if the promise might be too good to be true, I decided to perform my own experiment.
Last fall, I sent the identical fecal sample to both Viome (I paid $425, but the price has since dropped to $299) and DayTwo ($349). A couple of months later, both reports finally arrived, and I eagerly opened each app to compare their recommendations.
First, I examined my results from Viome, which was founded in 2016 in Cupertino, Calif., and declares without irony on its website that "conflicting food advice is now obsolete."
I learned I have "average" metabolic fitness and "average" inflammatory activity in my gut, which are scores that the company defines based on a proprietary algorithm. But I have "low" microbial richness, with only 62 active species of bacteria identified in my sample, compared with the mean of 157 in their test population. I also received a list of the specific species in my gut, with names like Lactococcus and Romboutsia.
But none of it meant anything to me without actionable food advice, so I clicked through to the Recommendations page and found a list of My Superfoods (cranberry, garlic, kale, salmon, turmeric, watermelon, and bone broth) and My Foods to Avoid (chickpeas, kombucha, lentils, and rice noodles). There was also a searchable database of many foods that had been categorized for me, like "bell pepper; minimize" and "beef; enjoy."
"I just don't think sufficient data is yet available to make reliable personalized dietary recommendations based on one's microbiome."
Next, I looked at my results from DayTwo, which was founded in 2015 from research out of the Weizmann Institute of Science in Israel, and whose pitch to consumers is, "Blood sugar made easy. The algorithm diet personalized to you."
This app had some notable differences. There was no result about my metabolic fitness, microbial richness, or list of the species in my sample. There was also no list of superfoods or foods to avoid. Instead, the app encouraged me to build a meal by searching for foods in their database and combining them in beneficial ways for my blood sugar. Two slices of whole wheat bread received a score of 2.7 out of 10 ("Avoid"), but if combined with one cup of large curd cottage cheese, the score improved to 6.8 ("Limit"), and if I added two hard-boiled eggs, the score went up to 7.5 ("Good").
Perusing my list of foods with "Excellent" scores, I noticed some troubling conflicts with the other app. Lentils, which had been a no-no according to Viome, received high marks from DayTwo. Ditto for Kombucha. My purported superfood of cranberry received low marks. Almonds got an almost perfect score (9.7) while Viome told me to minimize them. I found similarly contradictory advice for foods I regularly eat, including navel oranges, peanuts, pork, and beets.
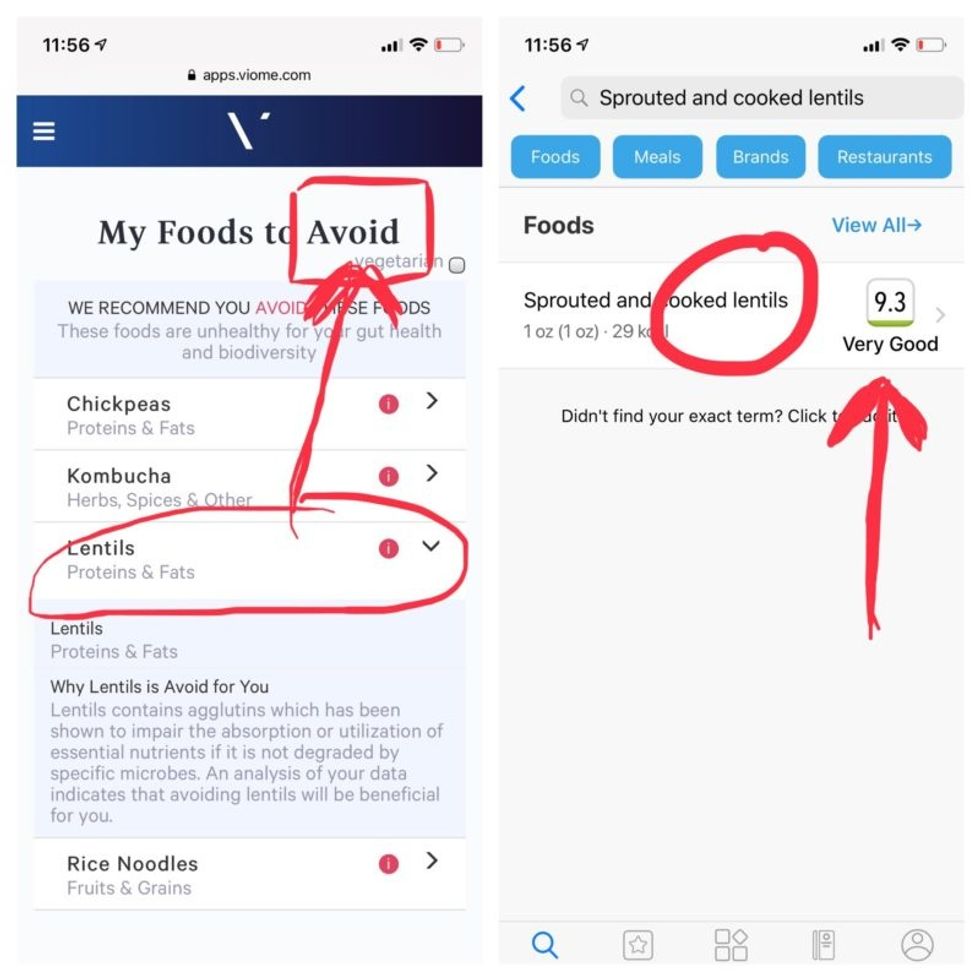
Contradictory dietary guidance that Kira Peikoff received from Viome (left) and DayTwo from an identical sample.
To be sure, there was some overlap. Both apps agreed on rice noodles (bad), chickpeas (bad), honey (bad), carrots (good), and avocado (good), among other foods.
But still, I was left scratching my head. Which set of recommendations should I trust, if either? And what did my results mean for the accuracy of this nascent field?
I called a couple of experts to find out.
"I have worked on the microbiome and nutrition for the last 20 years and I would be absolutely incapable of finding you evidence in the scientific literature that lentils have a detrimental effect based on the microbiome," said Dr. Jens Walter, an Associate Professor and chair for Nutrition, Microbes, and Gastrointestinal Health at the University of Alberta. "I just don't think sufficient data is yet available to make reliable personalized dietary recommendations based on one's microbiome. And even if they would have proprietary algorithms, at least one of them is not doing it right."
There is definite potential for personalized nutrition based on the microbiome, he said, but first, predictive models must be built and standardized, then linked to clinical endpoints, and tested in a large sample of healthy volunteers in order to enable extrapolations for the general population.
"It is mindboggling what you would need to do to make this work," he observed. "There are probably hundreds of relevant dietary compounds, then the microbiome has at least a hundred relevant species with a hundred or more relevant genes each, then you'd have to put all this together with relevant clinical outcomes. And there's a hundred-fold variation in that information between individuals."
However, Walter did acknowledge that the companies might be basing their algorithms on proprietary data that could potentially connect all the dots. I reached out to them to find out.
Amir Golan, the Chief Commercial Officer of DayTwo, told me, "It's important to emphasize this is a prediction, as the microbiome field is in a very early stage of research." But he added, "I believe we are the only company that has very solid science published in top journals and we can bring very actionable evidence and benefit to our uses."
He was referring to pioneering work out of the Weizmann Institute that was published in 2015 in the journal Cell, which logged the glycemic responses of 800 people in response to nearly 50,000 meals; adding information about the subjects' microbiomes enabled more accurate glycemic response predictions. Since then, Golan said, additional trials have been conducted, most recently with the Mayo Clinic, to duplicate the results, and other studies are ongoing whose results have not yet been published.
He also pointed out that the microbiome was merely one component that goes into building a client's profile, in addition to medical records, including blood glucose levels. (I provided my HbA1c levels, a measure of average blood sugar over the previous several months.)
"We are not saying we want to improve your gut microbiome. We provide a dynamic tool to help guide what you should eat to control your blood sugar and think about combinations," he said. "If you eat one thing, or with another, it will affect you in a different way."
Viome acknowledged that the two companies are taking very different approaches.
"DayTwo is primarily focused on the glycemic response," Naveen Jain, the CEO, told me. "If you can only eat butter for rest of your life, you will have no glycemic response but will probably die of a heart attack." He laughed. "Whereas we came from very different angle – what is happening inside the gut at a microbial level? When you eat food like spinach, how will that be metabolized in the gut? Will it produce the nutrients you need or cause inflammation?"
He said his team studied 1000 people who were on continuous glucose monitoring and fed them 45,000 meals, then built a proprietary data prediction model, looking at which microbes existed and how they actively broke down the food.
Jain pointed out that DayTwo sequences the DNA of the microbes, while Viome sequences the RNA – the active expression of DNA. That difference, in his opinion, is key to making accurate predictions.
"DNA is extremely stable, so when you eat any food and measure the DNA [in a fecal sample], you get all these false positives--you get DNA from plant food and meat, and you have no idea if those organisms are dead and simply transient, or actually exist. With RNA, you see what is actually alive in the gut."
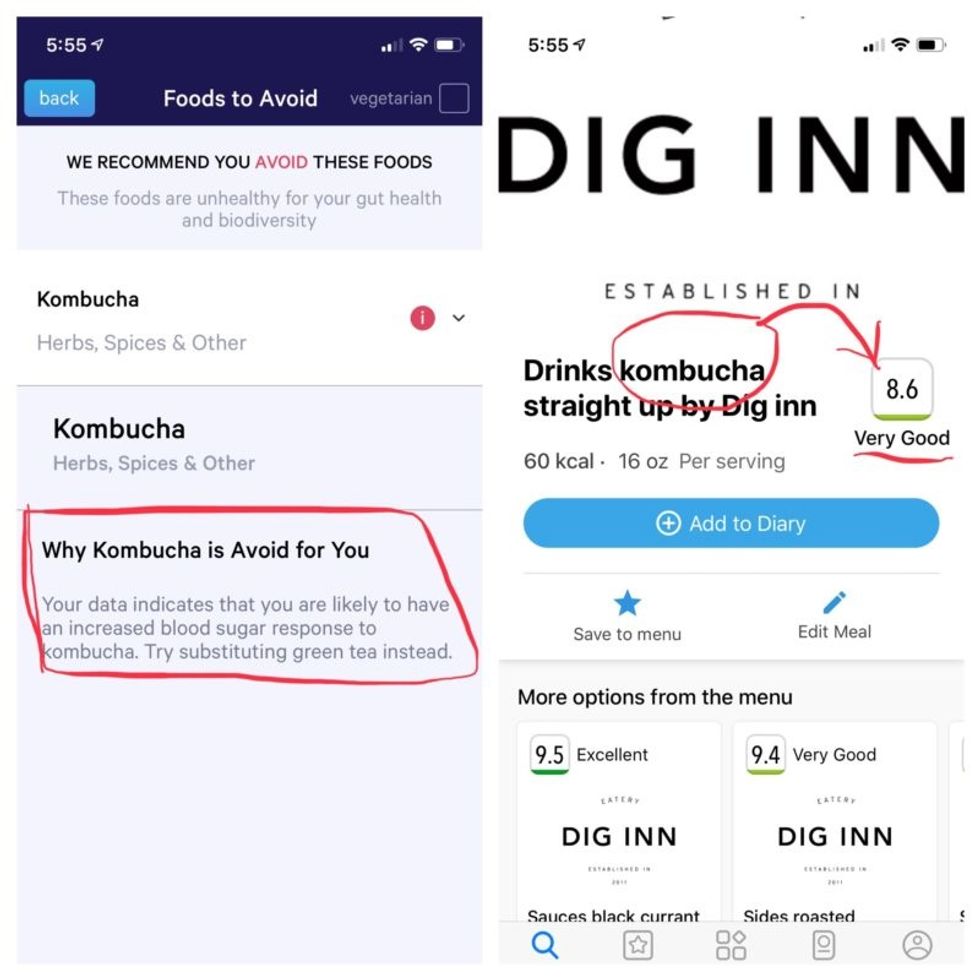
More contradictory food advice from Viome (left) and DayTwo.
Note that controversy exists over how it is possible with a fecal sample to effectively measure RNA, which degrades within minutes, though Jain said that his company has the technology to keep RNA stable for fourteen days.
Viome's approach, Jain maintains, is 90 percent accurate, based on as-yet unpublished data; a patent was filed just last week. DayTwo's approach is 66 percent accurate according to the latest published research.
Natasha Haskey, a registered dietician and doctoral student conducting research in the field of microbiome science and nutrition, is skeptical of both companies. "We can make broad statements, like eat more fruits and vegetables and fiber, but when it comes to specific foods, the science is just not there yet," she said. "I think there is a future, and we will be doing that someday, but not yet. Maybe we will be closer in ten years."
Professor Walter wholeheartedly agrees with Haskey, and suggested that if people want to eat a gut-healthy diet, they should focus on beneficial oils, fruits and vegetables, fish, a variety of whole grains, poultry and beans, and limit red meat and cheese, as well as avoid processed meats.
"These services are far over the tips of their science skis," Arthur Caplan, the founding head of New York University's Division of Medical Ethics, said in an email. "We simply don't know enough about the gut microbiome, its fluctuations and variability from person to person to support general [direct-to-consumer] testing. This is simply premature. We need standards for accuracy, specificity, and sensitivity, plus mandatory competent counseling for all such testing. They don't exist. Neither should DTC testing—yet."
Meanwhile, it's time for lunch. I close out my Viome and DayTwo apps and head to the kitchen to prepare a peanut butter sandwich. My gut tells me I'll be just fine.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
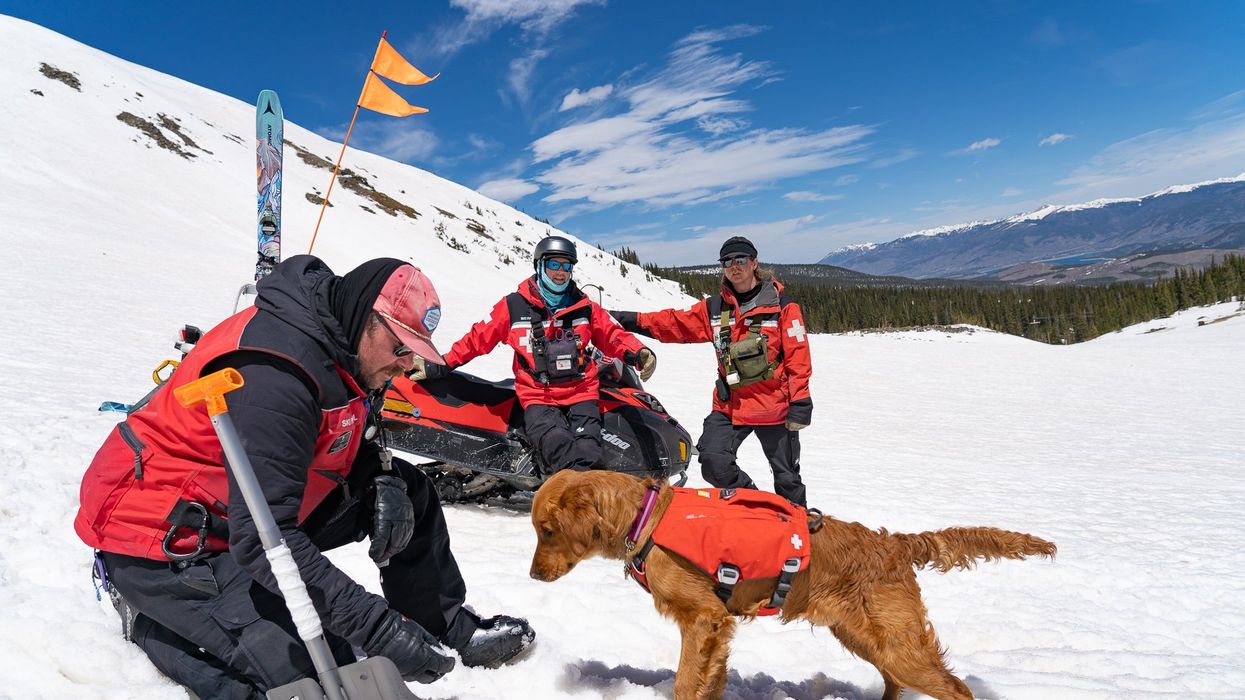
Avalanche rescue dogs train to find and dig out people buried in snow slides
Two-and-a-half year-old Huckleberry, a blue merle Australian shepherd, pulls hard at her leash; her yelps can be heard by skiers and boarders high above on the chairlift that carries them over the ski patrol hut to the top of the mountain. Huckleberry is an avalanche rescue dog — or avy dog, for short. She lives and works with her owner and handler, a ski patroller at Breckenridge Ski Resort in Colorado. As she watches the trainer play a game of hide-and-seek with six-month-old Lume, a golden retriever and avy dog-in-training, Huckleberry continues to strain on her leash; she loves the game. Hide-and-seek is one of the key training methods for teaching avy dogs the rescue skills they need to find someone caught in an avalanche — skier, snowmobiler, hiker, climber.
Lume’s owner waves a T-shirt in front of the puppy. While another patroller holds him back, Lume’s owner runs away and hides. About a minute later — after a lot of barking — Lume is released and commanded to “search.” He springs free, running around the hut to find his owner who reacts with a great amount of excitement and fanfare. Lume’s scent training will continue for the rest of the ski season (Breckenridge plans operating through May or as long as weather permits) and through the off-season. “We make this game progressively harder by not allowing the dog watch the victim run away,” explains Dave Leffler, Breckenridge's ski patroller and head of the avy dog program, who has owned, trained and raised many of them. Eventually, the trainers “dig an open hole in the snow to duck out of sight and gradually turn the hole into a cave where the dog has to dig to get the victim,” explains Leffler.
By the time he is three, Lume, like Huckleberry, will be a fully trained avy pup and will join seven other avy dogs on Breckenridge ski patrol team. Some of the team members, both human and canine, are also certified to work with Colorado Rapid Avalanche Deployment, a coordinated response team that works with the Summit County Sheriff’s office for avalanche emergencies outside of the ski slopes’ boundaries.
There have been 19 avalanche deaths in the U.S. this season, according to avalanche.org, which tracks slides; eight in Colorado. During the entirety of last season there were 17. Avalanche season runs from November through June, but avalanches can occur year-round.
High tech and high stakes
Complementing avy dogs’ ability to smell people buried in a slide, avalanche detection, rescue and recovery is becoming increasingly high tech. There are transceivers, signal locators, ground scanners and drones, which are considered “games changers” by many in avalanche rescue and recovery
For a person buried in an avalanche, the chance of survival plummets after 20 minutes, so every moment counts.
A drone can provide thermal imaging of objects caught in a slide; what looks like a rock from far away might be a human with a heat signature. Transceivers, also known as beacons, send a signal from an avalanche victim to a companion. Signal locators, like RECCO reflectors which are often sewn directly into gear, can echo back a radar signal sent by a detector; most ski resorts have RECCO detector units.
Research suggests that Ground Penetrating Radar (GPR), an electromagnetic tool used by geophysicists to pull images from inside the ground, could be used to locate an avalanche victim. A new study from the Department of Energy’s Sandia National Laboratories suggests that a computer program developed to pinpoint the source of a chemical or biological terrorist attack could also be used to find someone submerged in an avalanche. The search algorithm allows for small robots (described as cockroach-sized) to “swarm” a search area. Researchers say that this distributed optimization algorithm can help find avalanche victims four times faster than current search mechanisms. For a person buried in an avalanche, the chance of survival plummets after 20 minutes, so every moment counts.
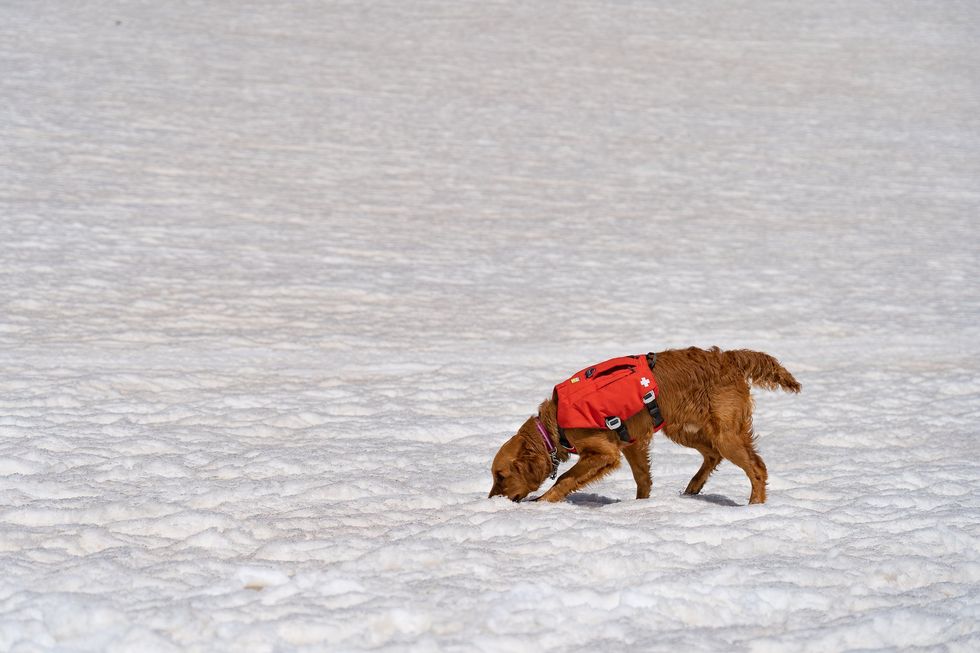

An avy dog in training is picking up scent
Sarah McLear
While rescue gear has been evolving, predicting when a slab will fall remains an emerging science — kind of where weather forecasting science was in the 1980s. Avalanche forecasting still relies on documenting avalanches by going out and looking,” says Ethan Greene, director of the Colorado Avalanche Information Center (CAIC). “So if there's a big snowstorm, and as you might remember, most avalanches happened during snowstorms, we could have 10,000 avalanches that release and we document 50,” says Greene. “Avalanche forecasting is essentially pattern recognition,” he adds--and understanding the layering structure of snow.
However, determining where the hazards lie can be tricky. While a dense layer of snow over a softer, weaker layer may be a recipe for an avalanche, there’s so much variability in snowpack that no one formula can predict the trigger. Further, observing and measuring snow at a single point may not be representative of all nearby slopes. Finally, there’s not enough historical data to help avalanche scientists create better prediction models.
That, however, may be changing.
Last year, an international group of researchers created computer simulations of snow cover using 16 years of meteorological data to forecast avalanche hazards, publishing their research in Cold Regions Science and Technology. They believe their models, which categorize different kinds of avalanches, can support forecasting and determine whether the avalanche is natural (caused by temperature changes, wind, additional snowfall) or artificial (triggered by a human or animal).
With smell receptors ranging from 800 million for an average dog, to 4 billion for scent hounds, canines remain key to finding people caught in slides.
With data from two sites in British Columbia and one in Switzerland, researchers built computer simulations of five different avalanche types. “In terms of real time avalanche forecasting, this has potential to fill in a lot of data gaps, where we don't have field observations of what the snow looks like,” says Simon Horton, a postdoctoral fellow with the Simon Fraser University Centre for Natural Hazards Research and a forecaster with Avalanche Canada, who participated in the study. While complex models that simulate snowpack layers have been around for a few decades, they weren’t easy to apply until recently. “It's been difficult to find out how to apply that to actual decision-making and improving safety,” says Horton. If you can derive avalanche problem types from simulated snowpack properties, he says, you’ll learn “a lot about how you want to manage that risk.”
The five categories include “new snow,” which is unstable and slides down the slope, “wet snow,” when rain or heat makes it liquidly, as well as “wind-drifted snow,” “persistent weak layers” and “old snow.” “That's when there's some type of deeply buried weak layer in the snow that releases without any real change in the weather,” Horton explains. “These ones tend to cause the most accidents.” One step by a person on that structurally weak layer of snow will cause a slide. Horton is hopeful that computer simulations of avalanche types can be used by scientists in different snow climates to help predict hazard levels.
Greene is doubtful. “If you have six slopes that are lined up next to each other, and you're going to try to predict which one avalanches and the exact dimensions and what time, that's going to be really hard to do. And I think it's going to be a long time before we're able to do that,” says Greene.
What both researchers do agree on, though, is that what avalanche prediction really needs is better imagery through satellite detection. “Just being able to count the number of avalanches that are out there will have a huge impact on what we do,” Greene says. “[Satellites] will change what we do, dramatically.” In a 2022 paper, scientists at the University of Aberdeen in England used satellites to study two deadly Himalayan avalanches. The imaging helped them determine that sediment from a 2016 ice avalanche plus subsequent snow avalanches contributed to the 2021 avalanche that caused a flash flood, killing over 200 people. The researchers say that understanding the avalanches characteristics through satellite imagery can inform them how one such event increases the magnitude of another in the same area.
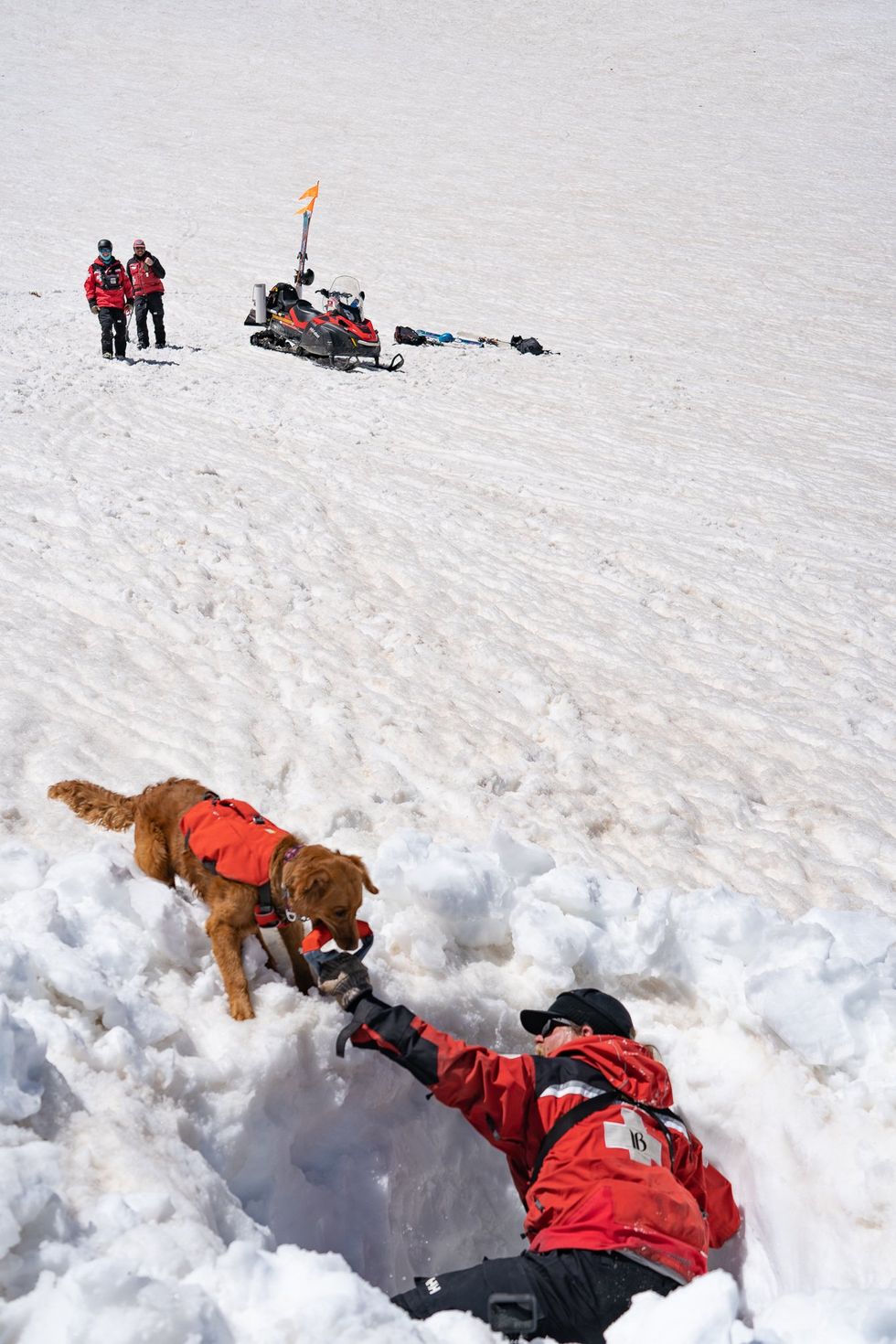

Avy dogs trainers hide in dug-out holes in the snow, teaching the dogs to find buried victims
Sarah McLear
Lifesaving combo: human tech and Mother Nature’s gear
Even as avalanche forecasting evolves, dogs with their built-in rescue mechanisms will remain invaluable. With smell receptors ranging from 800 million for an average dog, to 4 billion for scent hounds, canines remain key to finding people caught in slides. (Humans in comparison, have a meager 12 million.) A new study published in the Journal of Neuroscience revealed that in dogs smell and vision are connected in the brain, which has not been found in other animals. “They can detect the smell of their owner's fingerprints on a glass slide six weeks after they touched it,” says Nicholas Dodman, professor emeritus at Cummings School of Veterinary Medicine at Tufts University. “And they can track from a boat where a box filled with meat was buried in the water, 100 feet below,” says Dodman, who is also co-founder and president of the Center for Canine Behavior Studies.
Another recent study from Queens College in Belfast, United Kingdom, further confirms that dogs can smell when humans are stressed. They can also detect the smell of a person’s breath and the smell of the skin cells of a deceased person.
The emerging avalanche-predicting human-made tech and the incredible nature-made tech of dogs’ olfactory talents is the lifesaving “equipment” that Leffler believes in. Even when human-made technology develops further, it will be most efficient when used together with the millions of dogs’ smell receptors, Leffler believes. “It is a combination of technology and the avalanche dog that will always be effective in finding an avalanche victim.”
Living with someone changes your microbiome, new research shows
For the first time, research has shown that bacteria of the microbiome are transmitted between many individuals, not just infants and their mothers, in ways that can’t be explained by having the same diet or geography.
Some roommate frustration can be expected, whether it’s a sink piled high with crusty dishes or crumbs where a clean tabletop should be. Now, research suggests a less familiar issue: person-to-person transmission of shared bacterial strains in our gut and oral microbiomes. For the first time, the lab of Nicola Segata, a professor of genetics and computational biology at the University of Trento, located in Italy, has shown that bacteria of the microbiome are transmitted between many individuals, not just infants and their mothers, in ways that can’t be explained by their shared diet or geography.
It’s a finding with wide-ranging implications, yet frustratingly few predictable outcomes. Our microbiomes are an ever-growing and changing collection of helpful and harmful bacteria that we begin to accumulate the moment we’re born, but experts are still struggling to unravel why and how bacteria from one person’s gut or mouth become established in another person’s microbiome, as opposed to simply passing through.
“If we are looking at the overall species composition of the microbiome, then there is an effect of age of course, and many other factors,” Segata says. “But if we are looking at where our strains are coming from, 99 percent of them are only present in other people’s guts. They need to come from other guts.”
If we could better understand this process, we might be able to control and use it; perhaps hospital patients could avoid infections from other patients when their microbiome is depleted by antibiotics and their immune system is weakened, for example. But scientists are just beginning to link human microbiomes with various ailments. Growing evidence shows that our microbiomes steer our long-term health, impacting conditions like obesity, irritable bowel syndrome, type 2 diabetes, and cancer.
Previous work from Segata’s lab and others illuminated the ways bacteria are passed from mothers to infants during the first few months of life during vaginal birth, breastfeeding and other close contact. And scientists have long known that people in close proximity tend to share bacteria. But the factors related to that overlap, such as genetics and diet, were unclear, especially outside the mother-baby dyad.
“If we look at strain sharing between a mother and an infant at five years of age, for example, we cannot really tell which was due to transmission at birth and which is due to continued transmission because of contact,” Segata says. Experts hypothesized that they could be caused by bacterial similarities in the environment itself, genetics, or bacteria from shared foods that colonized the guts of people in close contact.
Strain sharing was highest in mother-child pairs, with 96 percent of them sharing strains, and only slightly lower in members of shared households, at 95 percent.
In Italy, researchers led by Mireia Valles-Colomer, including Segata, hoped to unravel this mystery. They compared data from 9,715 stool and saliva samples in 31 genomic datasets with existing metadata. Scientists zoomed in on variations in each bacterial strain down to the individual level. They examined not only mother-child pairs, but people living in the same household, adult twins, and people living in the same village in a level of detail that wasn’t possible before, due to its high cost and difficulties in retrieving data about interactions between individuals, Segata explained.
“This paper is, with high granularity, quantifying the percent sharing that you expect between different types of social interactions, controlling for things like genetics and diet,” Gibbons says. Strain sharing was highest in mother-child pairs, with 96 percent of them sharing strains, and only slightly lower in members of shared households, at 95 percent. And at least half of the mother-infant pairs shared 30 percent of their strains; the median was 12 percent among people in shared households. Yet, there was no sharing among eight percent of adult twins who lived separately, and 16 percent of people within villages who resided in different households. The results were published in Nature.
It’s not a regional phenomenon. Although the types of bacterial strains varied depending on whether people lived in western and eastern nations — datasets were drawn from 20 countries on five continents — the patterns of sharing were much the same. To establish these links, scientists focused on individual variations in shared bacterial strains, differences that create unique bacterial “fingerprints” in each person, while controlling for variables like diet, demonstrating that the bacteria had been transmitted between people and were not the result of environmental similarities.
The impact of this bacterial sharing isn’t clear, but shouldn’t be viewed with trepidation, according to Sean Gibbons, a microbiome scientist at the nonprofit Institute for Systems Biology.
“The vast majority of these bugs are actually either benign or beneficial to our health, and the fact that we're swapping and sharing them and that we can take someone else's strain and supplement or better diversify our own little garden is not necessarily a bad thing,” he says.
"There are hundreds of billions of dollars of investment capital moving into these microbiome therapeutic companies; bugs as drugs, so to speak,” says Sean Gibbons, a microbiome scientist at the Institute for Systems Biology.
Everyday habits like exercising and eating vegetables promote a healthy, balanced gut microbiome, which is linked to better metabolic and immune function, and fewer illnesses. While many people’s microbiomes contain bacteria like C. diff or E. coli, these bacteria don’t cause diseases in most cases because they’re present in low levels. But a microbiome that’s been wiped out by, say, antibiotics, may no longer keep these bacteria in check, allowing them to proliferate and make us sick.
“A big challenge in the microbiome field is being able to rationally predict whether, if you're exposed to a particular bug, it will stick in the context of your specific microbiome,” Gibbons says.
Gibbons predicts that explorations of microbe-based therapeutics will be “exploding” in the coming decades. “There are hundreds of billions of dollars of investment capital moving into these microbiome therapeutic companies; bugs as drugs, so to speak,” he says. Rather than taking a mass-marketed probiotic, a precise understanding of an individual’s microbiome could help target the introduction of just the right bacteria at just the right time to prevent or treat a particular illness.
Because the current study did not differentiate between different types of contact or relationships among household members sharing bacterial strains or determine the direction of transmission, Segata says his current project is examining children in daycare settings and tracking their microbiomes over time to understand the role genetics and everyday interactions play in the level of transmission that occurs.
This relatively newfound ability to trace bacterial variants to minute levels has unlocked the chance for scientists to untangle when and how bacteria leap from one microbiome to another. As researchers come to better understand the factors that permit a strain to establish itself within a microbiome, they could uncover new strategies to control these microbes, harnessing the makeup of each microbiome to help people to resist life-altering medical conditions.

