Life is Emerging: Review of Siddhartha Mukherjee’s Song of the Cell
A new book by Pulitzer-winning physician-scientist Siddhartha Mukherjee will be released from Simon & Schuster on October 25, 2022.
The DNA double helix is often the image spiraling at the center of 21st century advances in biomedicine and the growing bioeconomy. And yet, DNA is molecularly inert. DNA, the code for genes, is not alive and is not strictly necessary for life. Ought life be at the center of our communication of living systems? Is not the Cell a superior symbol of life and our manipulation of living systems?
A code for life isn’t a code without the life that instantiates it. A code for life must be translated. The cell is the basic unit of that translation. The cell is the minimal viable package of life as we know it. Therefore, cell biology is at the center of biomedicine’s greatest transformations, suggests Pulitzer-winning physician-scientist Siddhartha Mukherjee in his latest book, The Song of the Cell: The Exploration of Medicine and the New Human.
The Song of the Cell begins with the discovery of cells and of germ theory, featuring characters such as Louis Pasteur and Robert Koch, who brought the cell “into intimate contact with pathology and medicine.” This intercourse would transform biomedicine, leading to the insight that we can treat disease by thinking at the cellular level. The slightest rearrangement of sick cells might be the path toward alleviating suffering for the organism: eroding the cell walls of a bacterium while sparing our human cells; inventing a medium that coaxes sperm and egg to dance into cellular union for in vitro fertilization (IVF); designing molecular missiles that home to the receptors decorating the exterior of cancer cells; teaching adult skin cells to remember their embryonic state for regenerative medicines.
Mukherjee uses the bulk of the book to elucidate key cell types in the human body, along with their “connective relationships” that enable key organs and organ systems to function. This includes the immune system, the heart, the brain, and so on. Mukherjee’s distinctive style features compelling anecdotes and human stories that animate the scientific (and unscientific) processes that have led to our current state of understanding. In his chapter on neurons and the brain, for example, he integrates Santiago Ramon y Cajal’s meticulous black ink sketches of neurons into Mukherjee’s own personal encounter with clinical depression. In one lucid section, he interviews Dr. Helen Mayberg, a pioneering neurologist who takes seriously the descriptive power of her patients’ metaphors, as they suffer from “caves,” “holes,” “voids,” and “force fields” that render their lives gray. Dr. Mayberg aims to stimulate patients’ neuronal cells in a manner that brings back the color.

Beyond exposing the insight and inventiveness that has arisen out of cell-based thinking, it seems that Mukherjee’s bigger project is an epistemological one. The early chapters of The Song of the Cell continually hint at the potential for redefining the basic unit of biology as the cell rather than the gene. The choice to center biomedicine around cells is, above all, a conspicuous choice not to center it around genes (the subject of Mukherjee’s previous book, The Gene), because genes dominate popular science communication.
This choice of cells over genes is most welcome. Cells are alive. Genes are not. Letters—such as the As, Cs, Gs, and Ts that represent the nucleotides of DNA, which make up our genes—must be synthesized into a word or poem or song that offers a glimpse into deeper truths. A key idea embedded in this thinking is that of emergence. Whether in ancient myth or modern art, creation tends to be an emergent process, not a linearly coded script. The cell is our current best guess for the basic unit of life’s emergence, turning a finite set of chemical building blocks—nucleic acids, proteins, sugars, fats—into a replicative, evolving system for fighting stasis and entropy. The cell’s song is one for our times, for it is the song of biology’s emergence out of chemistry and physics, into the “frenetically active process” of homeostasis.
Re-centering our view of biology has practical consequences, too, for how we think about diagnosing and treating disease, and for inventing new medicines. Centering cells presents a challenge: which type of cell to place at the center? Rather than default to the apparent simplicity of DNA as a symbol because it represents the one master code for life, the tension in defining the diversity of cells—a mapping process still far from complete in cutting-edge biology laboratories—can help to create a more thoughtful library of cellular metaphors to shape both the practice and communication of biology.
Further, effective problem solving is often about operating at the right level, or the right scale. The cell feels like appropriate level at which to interrogate many of the diseases that ail us, because the senses that guide our own perceptions of sickness and health—the smoldering pain of inflammation, the tunnel vision of a migraine, the dizziness of a fluttering heart—are emergent.
This, unfortunately, is sort of where Mukherjee leaves the reader, under-exploring the consequences of a biology of emergence. Many practical and profound questions have to do with the ways that each scale of life feeds back on the others. In a tome on Cells and “the future human” I wished that Mukherjee had created more space for seeking the ways that cells will shape and be shaped by the future, of humanity and otherwise.
We are entering a phase of real-world bioengineering that features the modularization of cellular parts within cells, of cells within organs, of organs within bodies, and of bodies within ecosystems. In this reality, we would be unwise to assume that any whole is the mere sum of its parts.
For example, when discussing the regenerative power of pluripotent stem cells, Mukherjee raises the philosophical thought experiment of the Delphic boat, also known as the Ship of Theseus. The boat is made of many pieces of wood, each of which is replaced for repairs over the years, with the boat’s structure unchanged. Eventually none of the boat’s original wood remains: Is it the same boat?
Mukherjee raises the Delphic boat in one paragraph at the end of the chapter on stem cells, as a metaphor related to the possibility of stem cell-enabled regeneration in perpetuity. He does not follow any of the threads of potential answers. Given the current state of cellular engineering, about which Mukherjee is a world expert from his work as a physician-scientist, this book could have used an entire section dedicated to probing this question and, importantly, the ways this thought experiment falls apart.
We are entering a phase of real-world bioengineering that features the modularization of cellular parts within cells, of cells within organs, of organs within bodies, and of bodies within ecosystems. In this reality, we would be unwise to assume that any whole is the mere sum of its parts. Wholeness at any one of these scales of life—organelle, cell, organ, body, ecosystem—is what is at stake if we allow biological reductionism to assume away the relation between those scales.
In other words, Mukherjee succeeds in providing a masterful and compelling narrative of the lives of many of the cells that emerge to enliven us. Like his previous books, it is a worthwhile read for anyone curious about the role of cells in disease and in health. And yet, he fails to offer the broader context of The Song of the Cell.
As leading agronomist and essayist Wes Jackson has written, “The sequence of amino acids that is at home in the human cell, when produced inside the bacterial cell, does not fold quite right. Something about the E. coli internal environment affects the tertiary structure of the protein and makes it inactive. The whole in this case, the E. coli cell, affects the part—the newly made protein. Where is the priority of part now?” [1]
Beyond the ways that different kingdoms of life translate the same genetic code, the practical situation for humanity today relates to the ways that the different disciplines of modern life use values and culture to influence our genes, cells, bodies, and environment. It may be that humans will soon become a bit like the Delphic boat, infused with the buzz of fresh cells to repopulate different niches within our bodies, for healthier, longer lives. But in biology, as in writing, a mixed metaphor can cause something of a cacophony. For we are not boats with parts to be replaced piecemeal. And nor are whales, nor alpine forests, nor topsoil. Life isn’t a sum of parts, and neither is a song that rings true.
[1] Wes Jackson, "Visions and Assumptions," in Nature as Measure (p. 52-53).
Massive benefits of AI come with environmental and human costs. Can AI itself be part of the solution?
Generative AI has a large carbon footprint and other drawbacks. But AI can help mitigate its own harms—by plowing through mountains of data on extreme weather and human displacement.
The recent explosion of generative artificial intelligence tools like ChatGPT and Dall-E enabled anyone with internet access to harness AI’s power for enhanced productivity, creativity, and problem-solving. With their ever-improving capabilities and expanding user base, these tools proved useful across disciplines, from the creative to the scientific.
But beneath the technological wonders of human-like conversation and creative expression lies a dirty secret—an alarming environmental and human cost. AI has an immense carbon footprint. Systems like ChatGPT take months to train in high-powered data centers, which demand huge amounts of electricity, much of which is still generated with fossil fuels, as well as water for cooling. “One of the reasons why Open AI needs investments [to the tune of] $10 billion from Microsoft is because they need to pay for all of that computation,” says Kentaro Toyama, a computer scientist at the University of Michigan. There’s also an ecological toll from mining rare minerals required for hardware and infrastructure. This environmental exploitation pollutes land, triggers natural disasters and causes large-scale human displacement. Finally, for data labeling needed to train and correct AI algorithms, the Big Data industry employs cheap and exploitative labor, often from the Global South.
Generative AI tools are based on large language models (LLMs), with most well-known being various versions of GPT. LLMs can perform natural language processing, including translating, summarizing and answering questions. They use artificial neural networks, called deep learning or machine learning. Inspired by the human brain, neural networks are made of millions of artificial neurons. “The basic principles of neural networks were known even in the 1950s and 1960s,” Toyama says, “but it’s only now, with the tremendous amount of compute power that we have, as well as huge amounts of data, that it’s become possible to train generative AI models.”
Though there aren’t any official figures about the power consumption or emissions from data centers, experts estimate that they use one percent of global electricity—more than entire countries.
In recent months, much attention has gone to the transformative benefits of these technologies. But it’s important to consider that these remarkable advances may come at a price.
AI’s carbon footprint
In their latest annual report, 2023 Landscape: Confronting Tech Power, the AI Now Institute, an independent policy research entity focusing on the concentration of power in the tech industry, says: “The constant push for scale in artificial intelligence has led Big Tech firms to develop hugely energy-intensive computational models that optimize for ‘accuracy’—through increasingly large datasets and computationally intensive model training—over more efficient and sustainable alternatives.”
Though there aren’t any official figures about the power consumption or emissions from data centers, experts estimate that they use one percent of global electricity—more than entire countries. In 2019, Emma Strubell, then a graduate researcher at the University of Massachusetts Amherst, estimated that training a single LLM resulted in over 280,000 kg in CO2 emissions—an equivalent of driving almost 1.2 million km in a gas-powered car. A couple of years later, David Patterson, a computer scientist from the University of California Berkeley, and colleagues, estimated GPT-3’s carbon footprint at over 550,000 kg of CO2 In 2022, the tech company Hugging Face, estimated the carbon footprint of its own language model, BLOOM, as 25,000 kg in CO2 emissions. (BLOOM’s footprint is lower because Hugging Face uses renewable energy, but it doubled when other life-cycle processes like hardware manufacturing and use were added.)
Luckily, despite the growing size and numbers of data centers, their increasing energy demands and emissions have not kept pace proportionately—thanks to renewable energy sources and energy-efficient hardware.
But emissions don’t tell the full story.
AI’s hidden human cost
“If historical colonialism annexed territories, their resources, and the bodies that worked on them, data colonialism’s power grab is both simpler and deeper: the capture and control of human life itself through appropriating the data that can be extracted from it for profit.” So write Nick Couldry and Ulises Mejias, authors of the book The Costs of Connection.
The energy requirements, hardware manufacture and the cheap human labor behind AI systems disproportionately affect marginalized communities.
Technologies we use daily inexorably gather our data. “Human experience, potentially every layer and aspect of it, is becoming the target of profitable extraction,” Couldry and Meijas say. This feeds data capitalism, the economic model built on the extraction and commodification of data. While we are being dispossessed of our data, Big Tech commodifies it for their own benefit. This results in consolidation of power structures that reinforce existing race, gender, class and other inequalities.
“The political economy around tech and tech companies, and the development in advances in AI contribute to massive displacement and pollution, and significantly changes the built environment,” says technologist and activist Yeshi Milner, who founded Data For Black Lives (D4BL) to create measurable change in Black people’s lives using data. The energy requirements, hardware manufacture and the cheap human labor behind AI systems disproportionately affect marginalized communities.
AI’s recent explosive growth spiked the demand for manual, behind-the-scenes tasks, creating an industry described by Mary Gray and Siddharth Suri as “ghost work” in their book. This invisible human workforce that lies behind the “magic” of AI, is overworked and underpaid, and very often based in the Global South. For example, workers in Kenya who made less than $2 an hour, were the behind the mechanism that trained ChatGPT to properly talk about violence, hate speech and sexual abuse. And, according to an article in Analytics India Magazine, in some cases these workers may not have been paid at all, a case for wage theft. An exposé by the Washington Post describes “digital sweatshops” in the Philippines, where thousands of workers experience low wages, delays in payment, and wage theft by Remotasks, a platform owned by Scale AI, a $7 billion dollar American startup. Rights groups and labor researchers have flagged Scale AI as one company that flouts basic labor standards for workers abroad.
It is possible to draw a parallel with chattel slavery—the most significant economic event that continues to shape the modern world—to see the business structures that allow for the massive exploitation of people, Milner says. Back then, people got chocolate, sugar, cotton; today, they get generative AI tools. “What’s invisible through distance—because [tech companies] also control what we see—is the massive exploitation,” Milner says.
“At Data for Black Lives, we are less concerned with whether AI will become human…[W]e’re more concerned with the growing power of AI to decide who’s human and who’s not,” Milner says. As a decision-making force, AI becomes a “justifying factor for policies, practices, rules that not just reinforce, but are currently turning the clock back generations years on people’s civil and human rights.”
Ironically, AI plays an important role in mitigating its own harms—by plowing through mountains of data about weather changes, extreme weather events and human displacement.
Nuria Oliver, a computer scientist, and co-founder and vice-president of the European Laboratory of Learning and Intelligent Systems (ELLIS), says that instead of focusing on the hypothetical existential risks of today’s AI, we should talk about its real, tangible risks.
“Because AI is a transverse discipline that you can apply to any field [from education, journalism, medicine, to transportation and energy], it has a transformative power…and an exponential impact,” she says.
AI's accountability
“At the core of what we were arguing about data capitalism [is] a call to action to abolish Big Data,” says Milner. “Not to abolish data itself, but the power structures that concentrate [its] power in the hands of very few actors.”
A comprehensive AI Act currently negotiated in the European Parliament aims to rein Big Tech in. It plans to introduce a rating of AI tools based on the harms caused to humans, while being as technology-neutral as possible. That sets standards for safe, transparent, traceable, non-discriminatory, and environmentally friendly AI systems, overseen by people, not automation. The regulations also ask for transparency in the content used to train generative AIs, particularly with copyrighted data, and also disclosing that the content is AI-generated. “This European regulation is setting the example for other regions and countries in the world,” Oliver says. But, she adds, such transparencies are hard to achieve.
Google, for example, recently updated its privacy policy to say that anything on the public internet will be used as training data. “Obviously, technology companies have to respond to their economic interests, so their decisions are not necessarily going to be the best for society and for the environment,” Oliver says. “And that’s why we need strong research institutions and civil society institutions to push for actions.” ELLIS also advocates for data centers to be built in locations where the energy can be produced sustainably.
Ironically, AI plays an important role in mitigating its own harms—by plowing through mountains of data about weather changes, extreme weather events and human displacement. “The only way to make sense of this data is using machine learning methods,” Oliver says.
Milner believes that the best way to expose AI-caused systemic inequalities is through people's stories. “In these last five years, so much of our work [at D4BL] has been creating new datasets, new data tools, bringing the data to life. To show the harms but also to continue to reclaim it as a tool for social change and for political change.” This change, she adds, will depend on whose hands it is in.
DNA gathered from animal poop helps protect wildlife
Alida de Flamingh and her team are collecting elephant dung. It holds a trove of information about animal health, diet and genetic diversity.
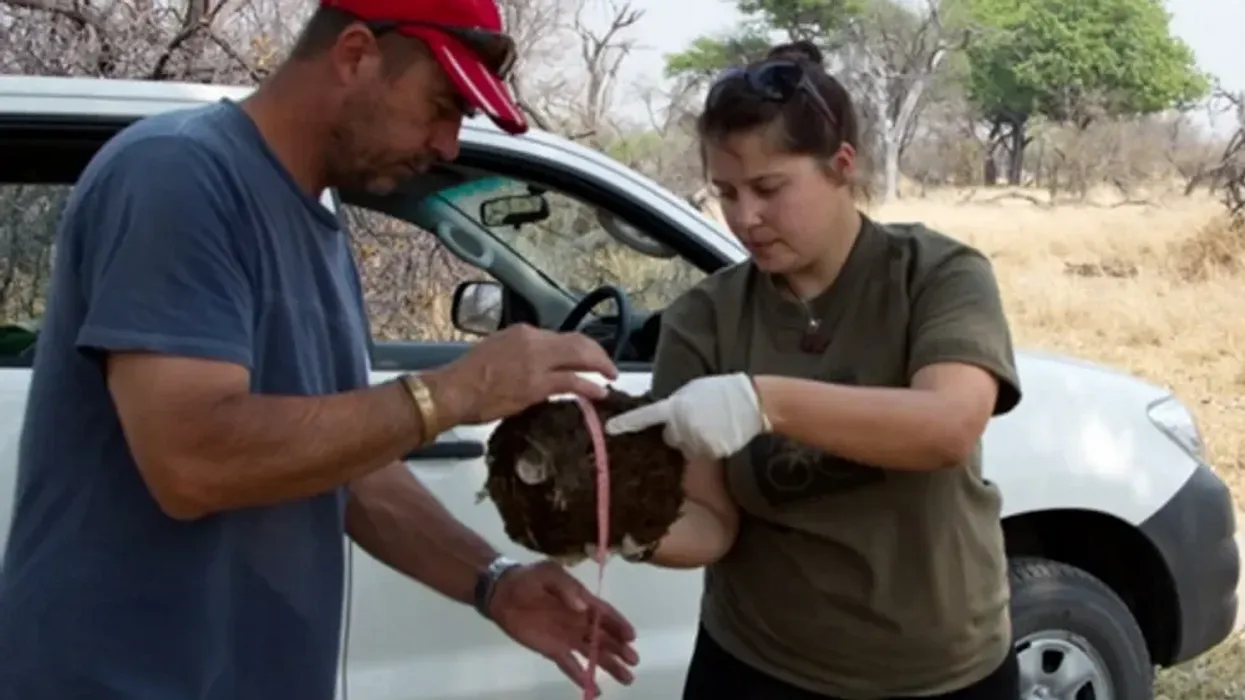
On the savannah near the Botswana-Zimbabwe border, elephants grazed contentedly. Nearby, postdoctoral researcher Alida de Flamingh watched and waited. As the herd moved away, she went into action, collecting samples of elephant dung that she and other wildlife conservationists would study in the months to come. She pulled on gloves, took a swab, and ran it all over the still-warm, round blob of elephant poop.
Sequencing DNA from fecal matter is a safe, non-invasive way to track and ultimately help protect over 42,000 species currently threatened by extinction. Scientists are using this DNA to gain insights into wildlife health, genetic diversity and even the broader environment. Applied to elephants, chimpanzees, toucans and other species, it helps scientists determine the genetic diversity of groups and linkages with other groups. Such analysis can show changes in rates of inbreeding. Populations with greater genetic diversity adapt better to changes and environmental stressors than those with less diversity, thus reducing their risks of extinction, explains de Flamingh, a postdoctoral researcher at the University of Illinois Urbana-Champaign.
Analyzing fecal DNA also reveals information about an animal’s diet and health, and even nearby flora that is eaten. That information gives scientists broader insights into the ecosystem, and the findings are informing conservation initiatives. Examples include restoring or maintaining genetic connections among groups, ensuring access to certain foraging areas or increasing diversity in captive breeding programs.
Approximately 27 percent of mammals and 28 percent of all assessed species are close to dying out. The IUCN Red List of threatened species, simply called the Red List, is the world’s most comprehensive record of animals’ risk of extinction status. The more information scientists gather, the better their chances of reducing those risks. In Africa, populations of vertebrates declined 69 percent between 1970 and 2022, according to the World Wildlife Fund (WWF).
“We put on sterile gloves and use a sterile swab to collect wet mucus and materials from the outside of the dung ball,” says Alida de Flamingh, a postdoctoral researcher at the University of Illinois Urbana-Champaign.
“When people talk about species, they often talk about ecosystems, but they often overlook genetic diversity,” says Christina Hvilsom, senior geneticist at the Copenhagen Zoo. “It’s easy to count (individuals) to assess whether the population size is increasing or decreasing, but diversity isn’t something we can see with our bare eyes. Yet, it’s actually the foundation for the species and populations.” DNA analysis can provide this critical information.
Assessing elephants’ health
“Africa’s elephant populations are facing unprecedented threats,” says de Flamingh, the postdoc, who has studied them since 2009. Challenges include ivory poaching, habitat destruction and smaller, more fragmented habitats that result in smaller mating pools with less genetic diversity. Additionally, de Flamingh studies the microbial communities living on and in elephants – their microbiomes – looking for parasites or dangerous microbes.
Approximately 415,000 elephants inhabit Africa today, but de Flamingh says the number would be four times higher without these challenges. The IUCN Red List reports African savannah elephants are endangered and African forest elephants are critically endangered. Elephants support ecosystem biodiversity by clearing paths that help other species travel. Their very footprints create small puddles that can host smaller organisms such as tadpoles. Elephants are often described as ecosystems’ engineers, so if they disappear, the rest of the ecosystem will suffer too.
There’s a process to collecting elephant feces. “We put on sterile gloves (which we change for each sample) and use a sterile swab to collect wet mucus and materials from the outside of the dung ball,” says de Flamingh. They rub a sample about the size of a U.S. quarter onto a paper card embedded with DNA preservation technology. Each card is air dried and stored in a packet of desiccant to prevent mold growth. This way, samples can be stored at room temperature indefinitely without the DNA degrading.
Earlier methods required collecting dung in bags, which needed either refrigeration or the addition of preservatives, or the riskier alternative of tranquilizing the animals before approaching them to draw blood samples. The ability to collect and sequence the DNA made things much easier and safer.
“Our research provides a way to assess elephant health without having to physically interact with elephants,” de Flamingh emphasizes. “We also keep track of the GPS coordinates of each sample so that we can create a map of the sampling locations,” she adds. That helps researchers correlate elephants’ health with geographic areas and their conditions.
Although de Flamingh works with elephants in the wild, the contributions of zoos in the United States and collaborations in South Africa (notably the late Professor Rudi van Aarde and the Conservation Ecology Research Unit at the University of Pretoria) were key in studying this method to ensure it worked, she points out.
Protecting chimpanzees
Genetic work with chimpanzees began about a decade ago. Hvilsom and her group at the Copenhagen Zoo analyzed DNA from nearly 1,000 fecal samples collected between 2003 and 2018 by a team of international researchers. The goal was to assess the status of the West African subspecies, which is critically endangered after rapid population declines. Of the four subspecies of chimpanzees, the West African subspecies is considered the most at-risk.
In total, the WWF estimates the numbers of chimpanzees inhabiting Africa’s forests and savannah woodlands at between 173,000 and 300,000. Poaching, disease and human-caused changes to their lands are their major risks.
By analyzing genetics obtained from fecal samples, Hvilsom estimated the chimpanzees’ population, ascertained their family relationships and mapped their migration routes.
“One of the threats is mining near the Nimba Mountains in Guinea,” a stronghold for the West African subspecies, Hvilsom says. The Nimba Mountains are a UNESCO World Heritage Site, but they are rich in iron ore, which is used to make the steel that is vital to the Asian construction boom. As she and colleagues wrote in a recent paper, “Many extractive industries are currently developing projects in chimpanzee habitat.”
Analyzing DNA allows researchers to identify individual chimpanzees more accurately than simply observing them, she says. Normally, field researchers would install cameras and manually inspect each picture to determine how many chimpanzees were in an area. But, Hvilsom says, “That’s very tricky. Chimpanzees move a lot and are fast, so it’s difficult to get clear pictures. Often, they find and destroy the cameras. Also, they live in large areas, so you need a lot of cameras.”
By analyzing genetics obtained from fecal samples, Hvilsom estimated the chimpanzees’ population, ascertained their family relationships and mapped their migration routes based upon DNA comparisons with other chimpanzee groups. The mining companies and builders are using this information to locate future roads where they won’t disrupt migration – a more effective solution than trying to build artificial corridors for wildlife.
“The current route cuts off communities of chimpanzees,” Hvilsom elaborates. That effectively prevents young adult chimps from joining other groups when the time comes, eventually reducing the currently-high levels of genetic diversity.
“The mining company helped pay for the genetics work,” Hvilsom says, “as part of its obligation to assess and monitor biodiversity and the effect of the mining in the area.”
Of 50 toucan subspecies, 11 are threatened or near-threatened with extinction because of deforestation and poaching.
Identifying toucan families
Feces aren't the only substance researchers draw DNA samples from. Jeffrey Coleman, a Ph.D. candidate at the University of Texas at Austin relies on blood tests for studying the genetic diversity of toucans---birds species native to Central America and nearby regions. They live in the jungles, where they hop among branches, snip fruit from trees, toss it in the air and catch it with their large beaks. “Toucans are beautiful, charismatic birds that are really important to the ecosystem,” says Coleman.
Of their 50 subspecies, 11 are threatened or near-threatened with extinction because of deforestation and poaching. “When people see these aesthetically pleasing birds, they’re motivated to care about conservation practices,” he points out.
Coleman works with the Dallas World Aquarium and its partner zoos to analyze DNA from blood draws, using it to identify which toucans are related and how closely. His goal is to use science to improve the genetic diversity among toucan offspring.
Specifically, he’s looking at sections of the genome of captive birds in which the nucleotides repeat multiple times, such as AGATAGATAGAT. Called microsatellites, these consecutively-repeating sections can be passed from parents to children, helping scientists identify parent-child and sibling-sibling relationships. “That allows you to make strategic decisions about how to pair (captive) individuals for mating...to avoid inbreeding,” Coleman says.

Jeffrey Coleman is studying the microsatellites inside the toucan genomes.
Courtesy Jeffrey Coleman
The alternative is to use a type of analysis that looks for a single DNA building block – a nucleotide – that differs in a given sequence. Called single nucleotide polymorphisms (SNPs, pronounced “snips”), they are very common and very accurate. Coleman says they are better than microsatellites for some uses. But scientists have already developed a large body of microsatellite data from multiple species, so microsatellites can shed more insights on relations.
Regardless of whether conservation programs use SNPs or microsatellites to guide captive breeding efforts, the goal is to help them build genetically diverse populations that eventually may supplement endangered populations in the wild. “The hope is that the ecosystem will be stable enough and that the populations (once reintroduced into the wild) will be able to survive and thrive,” says Coleman. History knows some good examples of captive breeding success.
The California condor, which had a total population of 27 in 1987, when the last wild birds were captured, is one of them. A captive breeding program boosted their numbers to 561 by the end of 2022. Of those, 347 of those are in the wild, according to the National Park Service.
Conservationists hope that their work on animals’ genetic diversity will help preserve and restore endangered species in captivity and the wild. DNA analysis is crucial to both types of efforts. The ability to apply genome sequencing to wildlife conservation brings a new level of accuracy that helps protect species and gives fresh insights that observation alone can’t provide.
“A lot of species are threatened,” Coleman says. “I hope this research will be a resource people can use to get more information on longer-term genealogies and different populations.”

