COVID Variants Are Like “a Thief Changing Clothes” – and Our Camera System Barely Exists

Being able to track variants of concern in real time is crucial to our ability to stay ahead of the virus.
Whether it's "natural selection" as Darwin called it, or it's "mutating" as the X-Men called it, living organisms change over time, developing thumbs or more efficient protein spikes, depending on the organism and the demands of its environment. The coronavirus that causes COVID-19, SARS-CoV-2, is not an exception, and now, after the virus has infected millions of people around the globe for more than a year, scientists are beginning to see those changes.
The notorious variants that have popped up include B.1.1.7, sometimes called the UK variant, as well as P.1 and B.1.351, which seem to have emerged in Brazil and South Africa respectively. As vaccinations are picking up pace, officials are warning that now
is not the time to become complacent or relax restrictions because the variants aren't well understood.
Some appear to be more transmissible, and deadlier, while others can evade the immune system's defenses better than earlier versions of the virus, potentially undermining the effectiveness of vaccines to some degree. Genomic surveillance, the process of sequencing the genetic code of the virus widely to observe changes and patterns, is a critical way that scientists can keep track of its evolution and work to understand how the variants might affect humans.
"It's like a thief changing clothes"
It's important to note that viruses mutate all the time. If there were funding and personnel to sequence the genome of every sample of the virus, scientists would see thousands of mutations. Not every variant deserves our attention. The vast majority of mutations are not important at all, but recognizing those that are is a crucial tool in getting and staying ahead of the virus. The work of sequencing, analyzing, observing patterns, and using public health tools as necessary is complicated and confusing to those without years of specialized training.
Jeremy Kamil, associate professor of microbiology and immunology at LSU Health Shreveport, in Louisiana, says that the variants developing are like a thief changing clothes. The thief goes in your house, steals your stuff, then leaves and puts on a different shirt and a wig, in the hopes you won't recognize them. Genomic surveillance catches the "thief" even in those different clothes.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context.
Understanding variants, both the uninteresting ones and the potentially concerning ones, gives public health officials and researchers at different levels a useful set of tools. Locally, knowing which variants are circulating in the community helps leaders know whether mask mandates and similar measures should be implemented or discontinued, or whether businesses and schools can open relatively safely.
There's more to it than observing new variants
Analysis is complex, particularly when it comes to understanding which variants are of concern. "So the question is always if a mutation becomes common, is that a random occurrence?" says Phoebe Lostroh, associate professor of molecular biology at Colorado College. "Or is the variant the result of some kind of selection because the mutation changes some property about the virus that makes it reproduce more quickly than variants of the virus that don't have that mutation? For a virus, [mutations can affect outcomes like] how much it replicates inside a person's body, how much somebody breathes it out, whether the particles that somebody might breathe in get smaller and can lead to greater transmission."
Along with all of those factors, accurate and useful genomic surveillance requires an understanding of where variants are occurring, how they are related, and an examination of why they might be prevalent.
For example, if a potentially worrisome variant appears in a community and begins to spread very quickly, it's not time to raise a public health alarm until several important questions have been answered, such as whether the variant is spreading due to specific events, or if it's happening because the mutation has allowed the virus to infect people more efficiently. Kamil offered a hypothetical scenario to explain: Imagine that a member of a community became infected and the virus mutated. That person went to church and three more people were infected, but one of them went to a karaoke bar and while singing infected 100 other people. Examining the conditions under which the virus has spread is, therefore, an essential part of untangling whether a mutation itself made the virus more transmissible or if an infected person's behaviors contributed to a local outbreak.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context. Genomic sequencing can help with that, but only when it's coordinated. When the same mutation occurs frequently, but is localized to one region, it's a concern, but when the same mutation happens in different places at the same time, it's much more likely that the "virus is learning that's a good mutation," explains Kamil.
The process is called convergent evolution, and it was a fascinating topic long before COVID. Just as your heritage can be traced through DNA, so can that of viruses, and when separate lineages develop similar traits it's almost like scientists can see evolution happening in real time. A mutation to SARS-CoV-2 that happens in more than one place at once is a mutation that makes it easier in some way for the virus to survive and that is when it may become alarming. The widespread, documented variants P.1 and B.1.351 are examples of convergence because they share some of the same virulent mutations despite having developed thousands of miles apart.
However, even variants that are emerging in different places at the same time don't present the kind of threat SARS-CoV-2 did in 2019. "This is nature," says Kamil. "It just means that this virus will not easily be driven to extinction or complete elimination by vaccines." Although a person who has already had COVID-19 can be reinfected with a variant, "it is almost always much milder disease" than the original infection, Kamil adds. Rather than causing full-fledged disease, variants have the potiental to "penetrate herd immunity, spreading relatively quietly among people who have developed natural immunity or been vaccinated, until the virus finds someone who has no immunity yet, and that person would be at risk of hospitalization-grade severe disease or death."
Surveillance and predictions
According to Lostroh, genomic surveillance can help scientists predict what's going to happen. "With the British strain, for instance, that's more transmissible, you can measure how fast it's doubling in the population and you can sort of tell whether we should take more measures against this mutation. Should we shut things down a little longer because that mutation is present in the population? That could be really useful if you did enough sampling in the population that you knew where it was," says Lostroh. If, for example, the more transmissible strain was present in 50 percent of cases, but in another county or state it was barely present, it would allow for rolling lockdowns instead of sweeping measures.
Variants are also extremely important when it comes to the development, manufacture, and distribution of vaccines. "You're also looking at medical countermeasures, such as whether your vaccine is still effective, or if your antiviral needs to be updated," says Lane Warmbrod, a senior analyst and research associate at Johns Hopkins Center for Health Security.
Properly funded and extensive genomic surveillance could eventually help control endemic diseases, too, like the seasonal flu, or other common respiratory infections. Kamil says he envisions a future in which genomic surveillance allows for prediction of sickness just as the weather is predicted today. "It's a 51 for infection today at the San Francisco Airport. There's been detection of some respiratory viruses," he says, offering an example. He says that if you're a vulnerable person, if you're immune-suppressed for some reason, you may want to wear a mask based on the sickness report.
The U.S. has the ability, but lacks standards
The benefits of widespread genomic surveillance are clear, and the United States certainly has the necessary technology, equipment, and personnel to carry it out. But, it's not happening at the speed and extent it needs to for the country to gain the benefits.
"The numbers are improving," said Kamil. "We're probably still at less than half a percent of all the samples that have been taken have been sequenced since the beginning of the pandemic."
Although there's no consensus on how many sequences is ideal for a robust surveillance program, modeling performed by the company Illumina suggests about 5 percent of positive tests should be sequenced. The reasons the U.S. has lagged in implementing a sequencing program are complex and varied, but solvable.
Perhaps the most important element that is currently missing is leadership. In order to conduct an effective genomic surveillance program, there need to be standards. The Johns Hopkins Center for Health Security recently published a paper with recommendations as to what kinds of elements need to be standardized in order to make the best use of sequencing technology and analysis.
"Along with which bioinformatic pipelines you're going to use to do the analyses, which sequencing strategy protocol are you going to use, what's your sampling strategy going to be, how is the data is going to be reported, what data gets reported," says Warmbrod. Currently, there's no guidance from the CDC on any of those things. So, while scientists can collect and report information, they may be collecting and reporting different information that isn't comparable, making it less useful for public health measures and vaccine updates.
Globally, one of the most important tools in making the information from genomic surveillance useful is GISAID, a platform designed for scientists to share -- and, importantly, to be credited for -- their data regarding genetic sequences of influenza. Originally, it was launched as a database of bird flu sequences, but has evolved to become an essential tool used by the WHO to make flu vaccine virus recommendations each year. Scientists who share their credentials have free access to the database, and anyone who uses information from the database must credit the scientist who uploaded that information.
Safety, logistics, and funding matter
Scientists at university labs and other small organizations have been uploading sequences to GISAID almost from the beginning of the pandemic, but their funding is generally limited, and there are no standards regarding information collection or reporting. Private, for-profit labs haven't had motivation to set up sequencing programs, although many of them have the logistical capabilities and funding to do so. Public health departments are understaffed, underfunded, and overwhelmed.
University labs may also be limited by safety concerns. The SARS-CoV-2 virus is dangerous, and there's a question of how samples should be transported to labs for sequencing.
Larger, for-profit organizations often have the tools and distribution capabilities to safely collect and sequence samples, but there hasn't been a profit motive. Genomic sequencing is less expensive now than ever before, but even at $100 per sample, the cost adds up -- not to mention the cost of employing a scientist with the proper credentials to analyze the sequence.
The path forward
The recently passed COVID-19 relief bill does have some funding to address genomic sequencing. Specifically, the American Rescue Plan Act includes $1.75 billion in funding for the Centers for Disease Control and Prevention's Advanced Molecular Detection (AMD) program. In an interview last month, CDC Director Rochelle Walensky said that the additional funding will be "a dial. And we're going to need to dial it up." AMD has already announced a collaboration called the Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance (SPHERES) Initiative that will bring together scientists from public health, academic, clinical, and non-profit laboratories across the country with the goal of accelerating sequencing.
Such a collaboration is a step toward following the recommendations in the paper Warmbrod coauthored. Building capacity now, creating a network of labs, and standardizing procedures will mean improved health in the future. "I want to be optimistic," she says. "The good news is there are a lot of passionate, smart, capable people who are continuing to work with government and work with different stakeholders." She cautions, however, that without a national strategy we won't succeed.
"If we maximize the potential and create that framework now, we can also use it for endemic diseases," she says. "It's a very helpful system for more than COVID if we're smart in how we plan it."
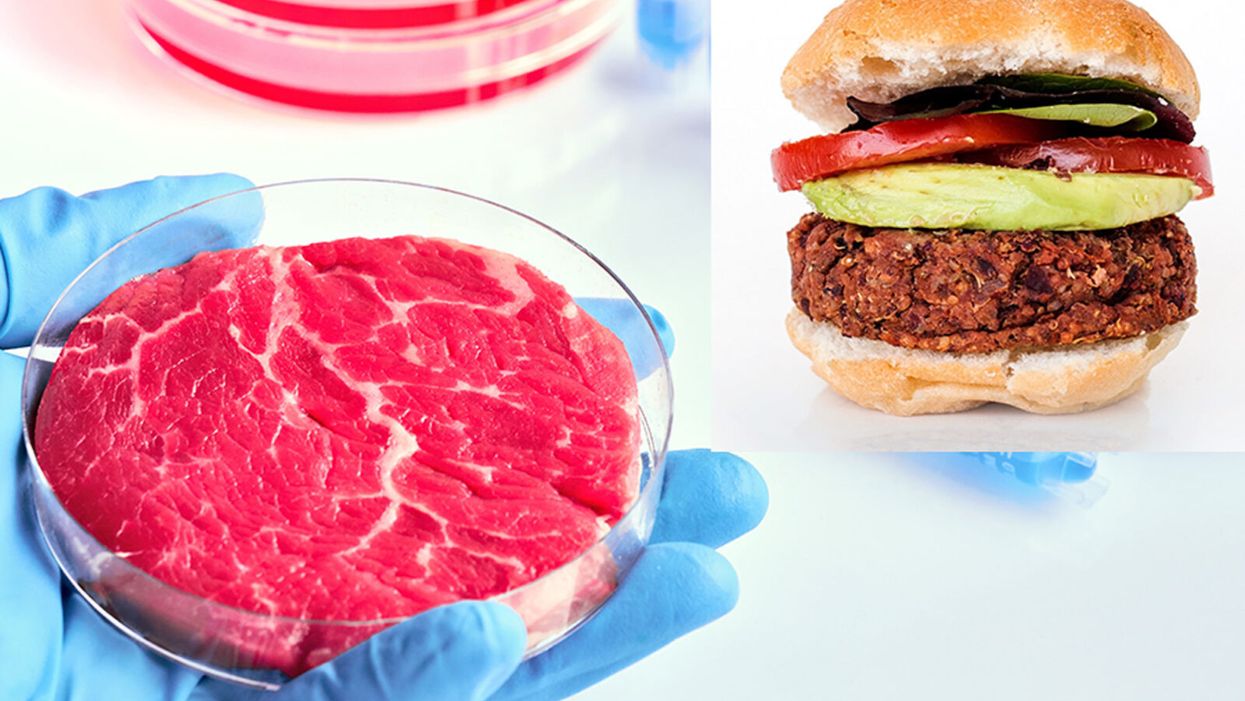
While lab-grown meat is not yet ready to crisis-proof the food supply chain, it offers key benefits that could make it an important solution beyond this pandemic.
The coronavirus pandemic exposed significant weaknesses in the country's food supply chain. Grocery store meat counters were bare. Transportation interruptions influenced supply. Finding beef, poultry, and pork at the store has been, in some places, as challenging as finding toilet paper.
In traditional agriculture models, it takes at least three months to raise chicken, six to nine months for pigs, and 18 months for cattle.
It wasn't a lack of supply -- millions of animals were in the pipeline.
"There's certainly enough food out there, but it can't get anywhere because of the way our system is set up," said Amy Rowat, an associate professor of integrative biology and physiology at UCLA. "Having a more self-contained, self-sufficient way to produce meat could make the supply chain more robust."
Cultured meat could be one way of making the meat supply chain more resilient despite disruptions due to pandemics such as COVID-19. But is the country ready to embrace lab-grown food?
According to a Good Food Institute study, GenZ is almost twice as likely to embrace meat alternatives for reasons related to social and environmental awareness, even prior to the pandemic. That's because this group wants food choices that reflect their values around food justice, equity, and animal welfare.
Largely, the interest in protein alternatives has been plant-based foods. However, factors directly related to COVID-19 may accelerate consumer interest in the scaling up of cell-grown products, according to Liz Specht, the associate director of science and technology at The Good Food Institute. The latter is a nonprofit organization that supports scientists, investors, and entrepreneurs working to develop food alternatives to conventional animal products.
While lab-grown food isn't ready yet to definitively crisis-proof the food supply chain, experts say it offers promise.
Matching Supply and Demand
Companies developing cell-grown meat claim it can take as few as two months to develop a cell into an edible product, according to Anthony Chow, CFA at Agronomics Limited, an investment company focused on meat alternatives. Tissue is taken from an animal and placed in a culture that contains nutrients and proteins the cells need to grow and expand. He cites a Good Food Institute report that claims a 2.5-millimeter sample can grow three and a half tons of meat in 40 days, allowing for exponential growth when needed.
In traditional agriculture models, it takes at least three months to raise chicken, six to nine months for pigs, and 18 months for cattle. To keep enough maturing animals in the pipeline, farms must plan the number of animals to raise months -- even years -- in advance. Lab-grown meat advocates say that because cultured meat supplies can be flexible, it theoretically allows for scaling up or down in significantly less time.
"Supply and demand has drastically changed in some way around the world and cultivated meat processing would be able to adapt much quicker than conventional farming," Chow said.
Scaling Up
Lab-grown meat may provide an eventual solution, but not in the immediate future, said Paul Mozdziak, a professor of physiology at North Carolina State University who researches animal cell culture techniques, transgenic animal production, and muscle biology.
"The challenge is in culture media," he said. "It's going to take some innovation to get the cells to grow at quantities that are going to be similar to what you can get from an animal. These are questions that everybody in the space is working on."
Chow says some of the most advanced cultured meat companies, such as BlueNal, anticipate introducing products to the market midway through next year. However, he thinks COVID-19 has slowed the process. Once introduced, they will be at a premium price, most likely available at restaurants before they hit grocery store shelves.
"I think in five years' time it will be in a different place," he said. "I don't think that this will have relevance for this pandemic, but certainly beyond that."
"Plant-based meats may be perceived as 'alternatives' to meat, whereas lab-grown meat is producing the same meat, just in a much more efficient manner, without the environmental implications."
Of course, all the technological solutions in the world won't solve the problem unless people are open-minded about embracing them. At least for now, a lab-grown burger or bluefin tuna might still be too strange for many people, especially in the U.S.
For instance, a 2019 article published by "Frontiers in Sustainable Food Systems" reflects results from a study of 3,030 consumers showing that 29 percent of U.S. customers, 59 percent of Chinese consumers, and 56 percent of Indian consumers were either 'very' or 'extremely likely' to try cultivated meat.
"Lab-grown meat is genuine meat, at the cellular level, and therefore will match conventional meat with regard to its nutritional content and overall sensory experience. It could be argued that plant-based meat will never be able to achieve this," says Laura Turner, who works with Chow at Agronomics Limited. "Plant-based meats may be perceived as 'alternatives' to meat, whereas lab-grown meat is producing the same meat, just in a much more efficient manner, without the environmental implications."
A Solution Beyond This Pandemic
The coronavirus has done more than raise awareness of the fragility of food supply chains. It has also been a wakeup call for consumers and policy makers that it is time to radically rethink our meat, Specht says. Those factors have elevated the profile of lab-grown meat.
"I think the economy is getting a little bit more steam and if I was an investor, I would be getting excited about it," adds Mozdziak.
Beyond crises, Mozdziak explains that as affluence continues to increase globally, meat consumption increases exponentially. Yet farm animals can only grow so quickly and traditional farming won't be able to keep up.
"Even Tyson is saying that by 2050, there's not going to be enough capacity in the animal meat space to meet demand," he notes. "If we don't look at some innovative technologies, how are we going to overcome that?"
A COVID-19 moonshot program challenged chemists around the world to submit ideas for how best to design a drug to target the virus.
By mid-March, Alpha Lee was growing restless. A pioneer of AI-driven drug discovery, Lee leads a team of researchers at the University of Cambridge, but his lab had been closed amidst the government-initiated lockdowns spreading inexorably across Europe.
If the Moonshot proves successful, they hope it could serve as a future benchmark for finding new medicines for chronic diseases.
Having spoken to his collaborators across the globe – many of whom were seeing their own experiments and research projects postponed indefinitely due to the pandemic – he noticed a similar sense of frustration and helplessness in the face of COVID-19.
While there was talk of finding a novel treatment for the virus, Lee was well aware the process was likely to be long and laborious. Traditional methods of drug discovery risked suffering the same fate as the efforts to find a cure for SARS in the early 2000, which took years and were ultimately abandoned long before a drug ever reached the market.
To avoid such an outcome, Lee was convinced that global collaboration was required. Together with a collection of scientists in the UK, US and Israel, he launched the 'COVID Moonshot' – a project which encouraged chemists worldwide to share their ideas for potential drug designs. If the Moonshot proves successful, they hope it could serve as a future benchmark for finding new medicines for chronic diseases.
Solving a Complex Jigsaw
In February, ShanghaiTech University published the first detailed snapshots of the SARS-CoV-2 coronavirus's proteins using a technique called X-ray crystallography. In particular, they revealed a high-resolution profile of the virus's main protease – the part of its structure that enables it to replicate inside a host – and the main drug target. The images were tantalizing.
"We could see all the tiny pieces sitting in the structure like pieces of a jigsaw," said Lee. "All we needed was for someone to come up with the best idea of joining these pieces together with a drug. Then you'd be left with a strong molecule which sits in the protease, and stops it from working, killing the virus in the process."
Normally, ideas for how best to design such a drug would be kept as carefully guarded secrets within individual labs and companies due to their potential value. But as a result, the steady process of trial and error to reach an optimum design can take years to come to fruition.
However, given the scale of the global emergency, Lee felt that the scientific community would be open to collective brainstorming on a mass scale. "Big Pharma usually wouldn't necessarily do this, but time is of the essence here," he said. "It was a case of, 'Let's just rethink every drug discovery stage to see -- ok, how can we go as fast as we can?'"
On March 13, he launched the COVID moonshot, calling for chemists around the globe to come up with the most creative ideas they could think of, on their laptops at home. No design was too weird or wacky to be considered, and crucially nothing would be patented. The entire project would be done on a not-for-profit basis, meaning that any drug that makes it to market will have been created simply for the good of humanity.
It caught fire: Within just two weeks, more than 2,300 potential drug designs had been submitted. By the middle of July, over 10,000 had been received from scientists around the globe.
The Road Toward Clinical Trials
With so many designs to choose from, the team has been attempting to whittle them down to a shortlist of the most promising. Computational drug discovery experts at Diamond and the Weizmann Institute of Science in Rehovot, Israel, have enabled the Moonshot team to develop algorithms for predicting how quick and easy each design would be to make, and to predict how well each proposed drug might bind to the virus in real life.
The latter is an approach known as computational covalent docking and has previously been used in cancer research. "This was becoming more popular even before COVID-19, with several covalent drugs approved by the FDA in recent years," said Nir London, professor of organic chemistry at the Weizmann Institute, and one of the Moonshot team members. "However, all of these were for oncology. A covalent drug against SARS-CoV-2 will certainly highlight covalent drug-discovery as a viable option."
Through this approach, the team have selected 850 compounds to date, which they have manufactured and tested in various preclinical trials already. Fifty of these compounds - which appear to be especially promising when it comes to killing the virus in a test tube – are now being optimized further.
Lee is hoping that at least one of these potential drugs will be shown to be effective in curing animals of COVID-19 within the next six months, a step that would allow the Moonshot team to reach out to potential pharmaceutical partners to test their compounds in humans.
Future Implications
If the project does succeed, some believe it could open the door to scientific crowdsourcing as a future means of generating novel medicine ideas for other diseases. Frank von Delft, professor of protein science and structural biology at the University of Oxford's Nuffield Department of Medicine, described it as a new form of 'citizen science.'
"There's a vast resource of expertise and imagination that is simply dying to be tapped into," he said.
Others are slightly more skeptical, pointing out that the uniqueness of the current crisis has meant that many scientists were willing to contribute ideas without expecting any future compensation in return. This meant that it was easy to circumvent the traditional hurdles that prevent large-scale global collaborations from happening – namely how to decide who will profit from the final product and who will hold the intellectual property (IP) rights.
"I think it is too early to judge if this is a viable model for future drug discovery," says London. "I am not sure that without the existential threat we would have seen so many contributions, and so many people and institutions willing to waive compensation and future royalties. Many scientists found themselves at home, frustrated that they don't have a way to contribute to the fight against COVID-19, and this project gave them an opportunity. Plus many can get behind the fact that this project has no associated IP and no one will get rich off of this effort. This breaks down a lot of the typical barriers and red-tape for wider collaboration."
"If a drug would sprout from one of these crowdsourced ideas, it would serve as a very powerful argument to consider this mode of drug discovery further in the future."
However the Moonshot team believes that if they can succeed, it will at the very least send a strong statement to policy makers and the scientific community that greater efforts should be made to make such large-scale collaborations more feasible.
"All across the scientific world, we've seen unprecedented adoption of open-science, collaboration and collegiality during this crisis, perhaps recognizing that only a coordinated global effort could address this global challenge," says London. "If a drug would sprout from one of these crowdsourced ideas, it would serve as a very powerful argument to consider this mode of drug discovery further in the future."
[An earlier version of this article was published on June 8th, 2020 as part of a standalone magazine called GOOD10: The Pandemic Issue. Produced as a partnership among LeapsMag, The Aspen Institute, and GOOD, the magazine is available for free online.]