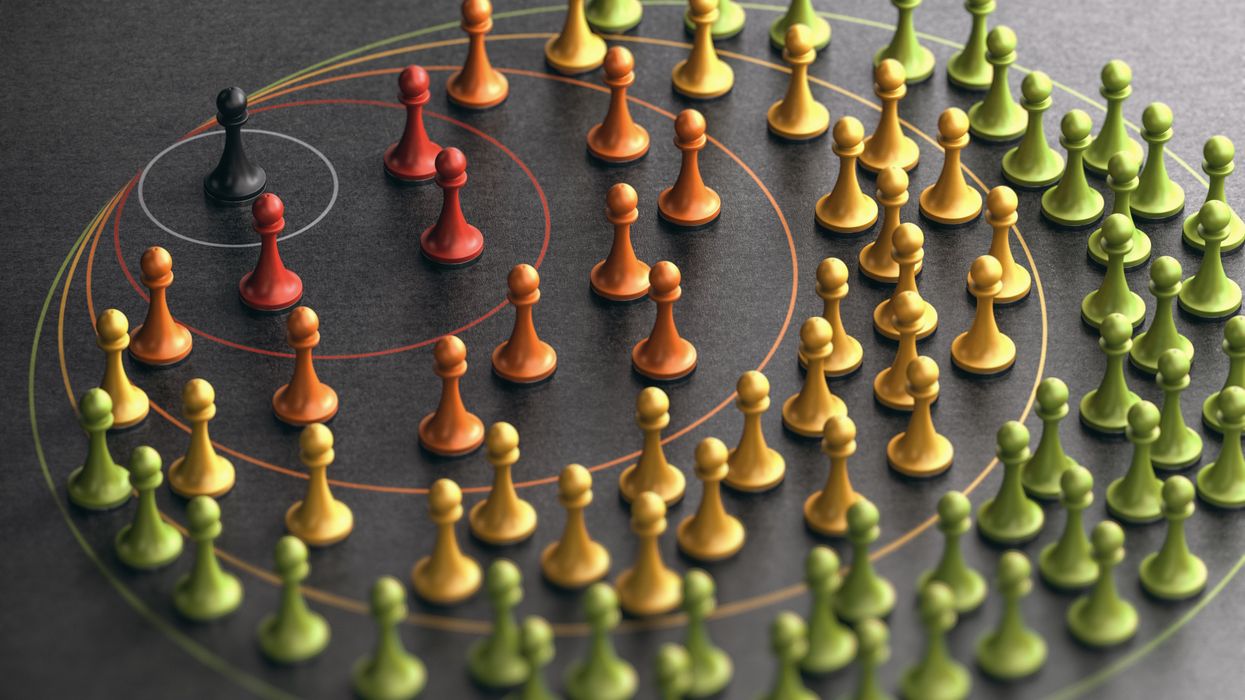
COVID Variants Are Like “a Thief Changing Clothes” – and Our Camera System Barely Exists

Being able to track variants of concern in real time is crucial to our ability to stay ahead of the virus.
Whether it's "natural selection" as Darwin called it, or it's "mutating" as the X-Men called it, living organisms change over time, developing thumbs or more efficient protein spikes, depending on the organism and the demands of its environment. The coronavirus that causes COVID-19, SARS-CoV-2, is not an exception, and now, after the virus has infected millions of people around the globe for more than a year, scientists are beginning to see those changes.
The notorious variants that have popped up include B.1.1.7, sometimes called the UK variant, as well as P.1 and B.1.351, which seem to have emerged in Brazil and South Africa respectively. As vaccinations are picking up pace, officials are warning that now
is not the time to become complacent or relax restrictions because the variants aren't well understood.
Some appear to be more transmissible, and deadlier, while others can evade the immune system's defenses better than earlier versions of the virus, potentially undermining the effectiveness of vaccines to some degree. Genomic surveillance, the process of sequencing the genetic code of the virus widely to observe changes and patterns, is a critical way that scientists can keep track of its evolution and work to understand how the variants might affect humans.
"It's like a thief changing clothes"
It's important to note that viruses mutate all the time. If there were funding and personnel to sequence the genome of every sample of the virus, scientists would see thousands of mutations. Not every variant deserves our attention. The vast majority of mutations are not important at all, but recognizing those that are is a crucial tool in getting and staying ahead of the virus. The work of sequencing, analyzing, observing patterns, and using public health tools as necessary is complicated and confusing to those without years of specialized training.
Jeremy Kamil, associate professor of microbiology and immunology at LSU Health Shreveport, in Louisiana, says that the variants developing are like a thief changing clothes. The thief goes in your house, steals your stuff, then leaves and puts on a different shirt and a wig, in the hopes you won't recognize them. Genomic surveillance catches the "thief" even in those different clothes.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context.
Understanding variants, both the uninteresting ones and the potentially concerning ones, gives public health officials and researchers at different levels a useful set of tools. Locally, knowing which variants are circulating in the community helps leaders know whether mask mandates and similar measures should be implemented or discontinued, or whether businesses and schools can open relatively safely.
There's more to it than observing new variants
Analysis is complex, particularly when it comes to understanding which variants are of concern. "So the question is always if a mutation becomes common, is that a random occurrence?" says Phoebe Lostroh, associate professor of molecular biology at Colorado College. "Or is the variant the result of some kind of selection because the mutation changes some property about the virus that makes it reproduce more quickly than variants of the virus that don't have that mutation? For a virus, [mutations can affect outcomes like] how much it replicates inside a person's body, how much somebody breathes it out, whether the particles that somebody might breathe in get smaller and can lead to greater transmission."
Along with all of those factors, accurate and useful genomic surveillance requires an understanding of where variants are occurring, how they are related, and an examination of why they might be prevalent.
For example, if a potentially worrisome variant appears in a community and begins to spread very quickly, it's not time to raise a public health alarm until several important questions have been answered, such as whether the variant is spreading due to specific events, or if it's happening because the mutation has allowed the virus to infect people more efficiently. Kamil offered a hypothetical scenario to explain: Imagine that a member of a community became infected and the virus mutated. That person went to church and three more people were infected, but one of them went to a karaoke bar and while singing infected 100 other people. Examining the conditions under which the virus has spread is, therefore, an essential part of untangling whether a mutation itself made the virus more transmissible or if an infected person's behaviors contributed to a local outbreak.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context. Genomic sequencing can help with that, but only when it's coordinated. When the same mutation occurs frequently, but is localized to one region, it's a concern, but when the same mutation happens in different places at the same time, it's much more likely that the "virus is learning that's a good mutation," explains Kamil.
The process is called convergent evolution, and it was a fascinating topic long before COVID. Just as your heritage can be traced through DNA, so can that of viruses, and when separate lineages develop similar traits it's almost like scientists can see evolution happening in real time. A mutation to SARS-CoV-2 that happens in more than one place at once is a mutation that makes it easier in some way for the virus to survive and that is when it may become alarming. The widespread, documented variants P.1 and B.1.351 are examples of convergence because they share some of the same virulent mutations despite having developed thousands of miles apart.
However, even variants that are emerging in different places at the same time don't present the kind of threat SARS-CoV-2 did in 2019. "This is nature," says Kamil. "It just means that this virus will not easily be driven to extinction or complete elimination by vaccines." Although a person who has already had COVID-19 can be reinfected with a variant, "it is almost always much milder disease" than the original infection, Kamil adds. Rather than causing full-fledged disease, variants have the potiental to "penetrate herd immunity, spreading relatively quietly among people who have developed natural immunity or been vaccinated, until the virus finds someone who has no immunity yet, and that person would be at risk of hospitalization-grade severe disease or death."
Surveillance and predictions
According to Lostroh, genomic surveillance can help scientists predict what's going to happen. "With the British strain, for instance, that's more transmissible, you can measure how fast it's doubling in the population and you can sort of tell whether we should take more measures against this mutation. Should we shut things down a little longer because that mutation is present in the population? That could be really useful if you did enough sampling in the population that you knew where it was," says Lostroh. If, for example, the more transmissible strain was present in 50 percent of cases, but in another county or state it was barely present, it would allow for rolling lockdowns instead of sweeping measures.
Variants are also extremely important when it comes to the development, manufacture, and distribution of vaccines. "You're also looking at medical countermeasures, such as whether your vaccine is still effective, or if your antiviral needs to be updated," says Lane Warmbrod, a senior analyst and research associate at Johns Hopkins Center for Health Security.
Properly funded and extensive genomic surveillance could eventually help control endemic diseases, too, like the seasonal flu, or other common respiratory infections. Kamil says he envisions a future in which genomic surveillance allows for prediction of sickness just as the weather is predicted today. "It's a 51 for infection today at the San Francisco Airport. There's been detection of some respiratory viruses," he says, offering an example. He says that if you're a vulnerable person, if you're immune-suppressed for some reason, you may want to wear a mask based on the sickness report.
The U.S. has the ability, but lacks standards
The benefits of widespread genomic surveillance are clear, and the United States certainly has the necessary technology, equipment, and personnel to carry it out. But, it's not happening at the speed and extent it needs to for the country to gain the benefits.
"The numbers are improving," said Kamil. "We're probably still at less than half a percent of all the samples that have been taken have been sequenced since the beginning of the pandemic."
Although there's no consensus on how many sequences is ideal for a robust surveillance program, modeling performed by the company Illumina suggests about 5 percent of positive tests should be sequenced. The reasons the U.S. has lagged in implementing a sequencing program are complex and varied, but solvable.
Perhaps the most important element that is currently missing is leadership. In order to conduct an effective genomic surveillance program, there need to be standards. The Johns Hopkins Center for Health Security recently published a paper with recommendations as to what kinds of elements need to be standardized in order to make the best use of sequencing technology and analysis.
"Along with which bioinformatic pipelines you're going to use to do the analyses, which sequencing strategy protocol are you going to use, what's your sampling strategy going to be, how is the data is going to be reported, what data gets reported," says Warmbrod. Currently, there's no guidance from the CDC on any of those things. So, while scientists can collect and report information, they may be collecting and reporting different information that isn't comparable, making it less useful for public health measures and vaccine updates.
Globally, one of the most important tools in making the information from genomic surveillance useful is GISAID, a platform designed for scientists to share -- and, importantly, to be credited for -- their data regarding genetic sequences of influenza. Originally, it was launched as a database of bird flu sequences, but has evolved to become an essential tool used by the WHO to make flu vaccine virus recommendations each year. Scientists who share their credentials have free access to the database, and anyone who uses information from the database must credit the scientist who uploaded that information.
Safety, logistics, and funding matter
Scientists at university labs and other small organizations have been uploading sequences to GISAID almost from the beginning of the pandemic, but their funding is generally limited, and there are no standards regarding information collection or reporting. Private, for-profit labs haven't had motivation to set up sequencing programs, although many of them have the logistical capabilities and funding to do so. Public health departments are understaffed, underfunded, and overwhelmed.
University labs may also be limited by safety concerns. The SARS-CoV-2 virus is dangerous, and there's a question of how samples should be transported to labs for sequencing.
Larger, for-profit organizations often have the tools and distribution capabilities to safely collect and sequence samples, but there hasn't been a profit motive. Genomic sequencing is less expensive now than ever before, but even at $100 per sample, the cost adds up -- not to mention the cost of employing a scientist with the proper credentials to analyze the sequence.
The path forward
The recently passed COVID-19 relief bill does have some funding to address genomic sequencing. Specifically, the American Rescue Plan Act includes $1.75 billion in funding for the Centers for Disease Control and Prevention's Advanced Molecular Detection (AMD) program. In an interview last month, CDC Director Rochelle Walensky said that the additional funding will be "a dial. And we're going to need to dial it up." AMD has already announced a collaboration called the Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance (SPHERES) Initiative that will bring together scientists from public health, academic, clinical, and non-profit laboratories across the country with the goal of accelerating sequencing.
Such a collaboration is a step toward following the recommendations in the paper Warmbrod coauthored. Building capacity now, creating a network of labs, and standardizing procedures will mean improved health in the future. "I want to be optimistic," she says. "The good news is there are a lot of passionate, smart, capable people who are continuing to work with government and work with different stakeholders." She cautions, however, that without a national strategy we won't succeed.
"If we maximize the potential and create that framework now, we can also use it for endemic diseases," she says. "It's a very helpful system for more than COVID if we're smart in how we plan it."
Living with someone changes your microbiome, new research shows
For the first time, research has shown that bacteria of the microbiome are transmitted between many individuals, not just infants and their mothers, in ways that can’t be explained by having the same diet or geography.
Some roommate frustration can be expected, whether it’s a sink piled high with crusty dishes or crumbs where a clean tabletop should be. Now, research suggests a less familiar issue: person-to-person transmission of shared bacterial strains in our gut and oral microbiomes. For the first time, the lab of Nicola Segata, a professor of genetics and computational biology at the University of Trento, located in Italy, has shown that bacteria of the microbiome are transmitted between many individuals, not just infants and their mothers, in ways that can’t be explained by their shared diet or geography.
It’s a finding with wide-ranging implications, yet frustratingly few predictable outcomes. Our microbiomes are an ever-growing and changing collection of helpful and harmful bacteria that we begin to accumulate the moment we’re born, but experts are still struggling to unravel why and how bacteria from one person’s gut or mouth become established in another person’s microbiome, as opposed to simply passing through.
“If we are looking at the overall species composition of the microbiome, then there is an effect of age of course, and many other factors,” Segata says. “But if we are looking at where our strains are coming from, 99 percent of them are only present in other people’s guts. They need to come from other guts.”
If we could better understand this process, we might be able to control and use it; perhaps hospital patients could avoid infections from other patients when their microbiome is depleted by antibiotics and their immune system is weakened, for example. But scientists are just beginning to link human microbiomes with various ailments. Growing evidence shows that our microbiomes steer our long-term health, impacting conditions like obesity, irritable bowel syndrome, type 2 diabetes, and cancer.
Previous work from Segata’s lab and others illuminated the ways bacteria are passed from mothers to infants during the first few months of life during vaginal birth, breastfeeding and other close contact. And scientists have long known that people in close proximity tend to share bacteria. But the factors related to that overlap, such as genetics and diet, were unclear, especially outside the mother-baby dyad.
“If we look at strain sharing between a mother and an infant at five years of age, for example, we cannot really tell which was due to transmission at birth and which is due to continued transmission because of contact,” Segata says. Experts hypothesized that they could be caused by bacterial similarities in the environment itself, genetics, or bacteria from shared foods that colonized the guts of people in close contact.
Strain sharing was highest in mother-child pairs, with 96 percent of them sharing strains, and only slightly lower in members of shared households, at 95 percent.
In Italy, researchers led by Mireia Valles-Colomer, including Segata, hoped to unravel this mystery. They compared data from 9,715 stool and saliva samples in 31 genomic datasets with existing metadata. Scientists zoomed in on variations in each bacterial strain down to the individual level. They examined not only mother-child pairs, but people living in the same household, adult twins, and people living in the same village in a level of detail that wasn’t possible before, due to its high cost and difficulties in retrieving data about interactions between individuals, Segata explained.
“This paper is, with high granularity, quantifying the percent sharing that you expect between different types of social interactions, controlling for things like genetics and diet,” Gibbons says. Strain sharing was highest in mother-child pairs, with 96 percent of them sharing strains, and only slightly lower in members of shared households, at 95 percent. And at least half of the mother-infant pairs shared 30 percent of their strains; the median was 12 percent among people in shared households. Yet, there was no sharing among eight percent of adult twins who lived separately, and 16 percent of people within villages who resided in different households. The results were published in Nature.
It’s not a regional phenomenon. Although the types of bacterial strains varied depending on whether people lived in western and eastern nations — datasets were drawn from 20 countries on five continents — the patterns of sharing were much the same. To establish these links, scientists focused on individual variations in shared bacterial strains, differences that create unique bacterial “fingerprints” in each person, while controlling for variables like diet, demonstrating that the bacteria had been transmitted between people and were not the result of environmental similarities.
The impact of this bacterial sharing isn’t clear, but shouldn’t be viewed with trepidation, according to Sean Gibbons, a microbiome scientist at the nonprofit Institute for Systems Biology.
“The vast majority of these bugs are actually either benign or beneficial to our health, and the fact that we're swapping and sharing them and that we can take someone else's strain and supplement or better diversify our own little garden is not necessarily a bad thing,” he says.
"There are hundreds of billions of dollars of investment capital moving into these microbiome therapeutic companies; bugs as drugs, so to speak,” says Sean Gibbons, a microbiome scientist at the Institute for Systems Biology.
Everyday habits like exercising and eating vegetables promote a healthy, balanced gut microbiome, which is linked to better metabolic and immune function, and fewer illnesses. While many people’s microbiomes contain bacteria like C. diff or E. coli, these bacteria don’t cause diseases in most cases because they’re present in low levels. But a microbiome that’s been wiped out by, say, antibiotics, may no longer keep these bacteria in check, allowing them to proliferate and make us sick.
“A big challenge in the microbiome field is being able to rationally predict whether, if you're exposed to a particular bug, it will stick in the context of your specific microbiome,” Gibbons says.
Gibbons predicts that explorations of microbe-based therapeutics will be “exploding” in the coming decades. “There are hundreds of billions of dollars of investment capital moving into these microbiome therapeutic companies; bugs as drugs, so to speak,” he says. Rather than taking a mass-marketed probiotic, a precise understanding of an individual’s microbiome could help target the introduction of just the right bacteria at just the right time to prevent or treat a particular illness.
Because the current study did not differentiate between different types of contact or relationships among household members sharing bacterial strains or determine the direction of transmission, Segata says his current project is examining children in daycare settings and tracking their microbiomes over time to understand the role genetics and everyday interactions play in the level of transmission that occurs.
This relatively newfound ability to trace bacterial variants to minute levels has unlocked the chance for scientists to untangle when and how bacteria leap from one microbiome to another. As researchers come to better understand the factors that permit a strain to establish itself within a microbiome, they could uncover new strategies to control these microbes, harnessing the makeup of each microbiome to help people to resist life-altering medical conditions.
Are the gains from gain-of-function research worth the risks?
Gain-of-function research can make pathogens more infectious and deadly. It also enables scientists to prepare remedies in advance.
Scientists have long argued that gain-of-function research, which can make viruses and other infectious agents more contagious or more deadly, was necessary to develop therapies and vaccines to counter the pathogens in case they were used for biological warfare. As the SARS-CoV-2 origins are being investigated, one prominent theory suggests it had leaked from a biolab that conducted gain-of-function research, causing a global pandemic that claimed nearly 6.9 million lives. Now some question the wisdom of engaging in this type of research, stating that the risks may far outweigh the benefits.
“Gain-of-function research means genetically changing a genome in a way that might enhance the biological function of its genes, such as its transmissibility or the range of hosts it can infect,” says George Church, professor of genetics at Harvard Medical School. This can occur through direct genetic manipulation as well as by encouraging mutations while growing successive generations of micro-organism in culture. “Some of these changes may impact pathogenesis in a way that is hard to anticipate in advance,” Church says.
In the wake of the global pandemic, the pros and cons of gain-of-function research are being fiercely debated. Some scientists say this type of research is vital for preventing future pandemics or for preparing for bioweapon attacks. Others consider it another disaster waiting to happen. The Government Accounting Office issued a report charging that a framework developed by the U.S. Department of Health & Human Services (HHS) provided inadequate oversight of this potentially deadly research. There’s a movement to stop it altogether. In January, the Viral Gain-of-Function Research Moratorium Act (S. 81) was introduced into the Senate to cease awarding federal research funding to institutions doing gain-of-function studies.
While testifying before the House COVID Origins Select Committee on March 8th, Robert Redfield, former director of the U.S. Centers for Disease Control and Prevention, said that COVID-19 may have resulted from an accidental lab leak involving gain-of-function research. Redfield said his conclusion is based upon the “rapid and high infectivity for human-to-human transmission, which then predicts the rapid evolution of new variants.”
“It is a very, very, very small subset of life science research that could potentially generate a potential pandemic pathogen,” said Gerald Parker, associate dean for Global One Health at Texas A&M University.
“In my opinion,” Redfield continues, “the COVID-19 pandemic presents a case study on the potential dangers of such research. While many believe that gain-of-function research is critical to get ahead of viruses by developing vaccines, in this case, I believe that was the exact opposite.” Consequently, Redfield called for a moratorium on gain-of-function research until there is consensus about the value of such risky science.
What constitutes risky?
The Federal Select Agent Program lists 68 specific infectious agents as risky because they are either very contagious or very deadly. In order to work with these 68 agents, scientists must register with the federal government. Meanwhile, research on deadly pathogens that aren’t easily transmitted, or pathogens that are quite contagious but not deadly, can be conducted without such oversight. “If you’re not working with select agents, you’re not required to register the research with the federal government,” says Gerald Parker, associate dean for Global One Health at Texas A&M University. But the 68-item list may not have everything that could possibly become dangerous or be engineered to be dangerous, thus escaping the government’s scrutiny—an issue that new regulations aim to address.
In January 2017, the White House Office of Science and Technology Policy (OSTP) issued additional guidance. It required federal departments and agencies to follow a series of steps when reviewing proposed research that could create, transfer, or use potential pandemic pathogens resulting from the enhancement of a pathogen’s transmissibility or virulence in humans.
In defining risky pathogens, OSTP included viruses that were likely to be highly transmissible and highly virulent, and thus very deadly. The Proposed Biosecurity Oversight Framework for the Future of Science, outlined in 2023, broadened the scope to require federal review of research “that is reasonably anticipated to enhance the transmissibility and/or virulence of any pathogen” likely to pose a threat to public health, health systems or national security. Those types of experiments also include the pathogens’ ability to evade vaccines or therapeutics, or diagnostic detection.
However, Parker says that dangers of generating a pandemic-level germ are tiny. “It is a very, very, very small subset of life science research that could potentially generate a potential pandemic pathogen.” Since gain-of-function guidelines were first issued in 2017, only three such research projects have met those requirements for HHS review. They aimed to study influenza and bird flu. Only two of those projects were funded, according to the NIH Office of Science Policy. For context, NIH funded approximately 11,000 of the 54,000 grant applications it received in 2022.
Guidelines governing gain-of-function research are being strengthened, but Church points out they aren’t ideal yet. “They need to be much clearer about penalties and avoiding positive uses before they would be enforceable.”
What do we gain from gain-of-function research?
The most commonly cited reason to conduct gain-of-function research is for biodefense—the government’s ability to deal with organisms that may pose threats to public health.
In the era of mRNA vaccines, the advance preparedness argument may be even less relevant.
“The need to work with potentially dangerous viruses is central to our preparedness,” Parker says. “It’s essential that we know and understand the basic biology, microbiology, etc. of some of these dangerous pathogens.” That includes increasing our knowledge of the molecular mechanisms by which a virus could become a sustained threat to humans. “Knowing that could help us detect [risks] earlier,” Parker says—and could make it possible to have medical countermeasures, like vaccines and therapeutics, ready.
Most vaccines, however, aren’t affected by this type of research. Essentially, scientists hope they will never need to use it. Moreover, Paul Mango, HSS former deputy chief of staff for policy, and author of the 2022 book Warp Speed, says he believes that in the era of mRNA vaccines, the advance preparedness argument may be even less relevant. “That’s because these vaccines can be developed and produced in less than 12 months, unlike traditional vaccines that require years of development,” he says.
Can better oversight guarantee safety?
Another situation, which Parker calls unnecessarily dangerous, is when regulatory bodies cannot verify that the appropriate biosafety and biosecurity controls are in place.
Gain-of-function studies, Parker points out, are conducted at the basic research level, and they’re performed in high-containment labs. “As long as all the processes, procedures and protocols are followed and there’s appropriate oversight at the institutional and scientific level, it can be conducted safely.”
Globally, there are 69 Biosafety Level 4 (BSL4) labs operating, under construction or being planned, according to recent research from King’s College London and George Mason University for Global BioLabs. Eleven of these 18 high-containment facilities that are planned or under construction are in Asia. Overall, three-quarters of the BSL4 labs are in cities, increasing public health risks if leaks occur.
Researchers say they are confident in the oversight system for BSL4 labs within the U.S. They are less confident in international labs. Global BioLabs’ report concurs. It gives the highest scores for biosafety to industrialized nations, led by France, Australia, Canada, the U.S. and Japan, and the lowest scores to Saudi Arabia, India and some developing African nations. Scores for biosecurity followed similar patterns.
“There are no harmonized international biosafety and biosecurity standards,” Parker notes. That issue has been discussed for at least a decade. Now, in the wake of SARS and the COVID-19 pandemic, scientists and regulators are likely to push for unified oversight standards. “It’s time we got serious about international harmonization of biosafety and biosecurity standards and guidelines,” Parker says. New guidelines are being worked on. The National Science Advisory Board for Biosecurity (NSABB) outlined its proposed recommendations in the document titled Proposed Biosecurity Oversight Framework for the Future of Science.
The debates about whether gain-of-function research is useful or poses unnecessary risks to humanity are likely to rage on for a while. The public too has a voice in this debate and should weigh in by communicating with their representatives in government, or by partaking in educational forums or initiatives offered by universities and other institutions. In the meantime, scientists should focus on improving the research regulations, Parker notes. “We need to continue to look for lessons learned and for gaps in our oversight system,” he says. “That’s what we need to do right now.”