Scientists want the salamander's secret: how they regenerate tissue
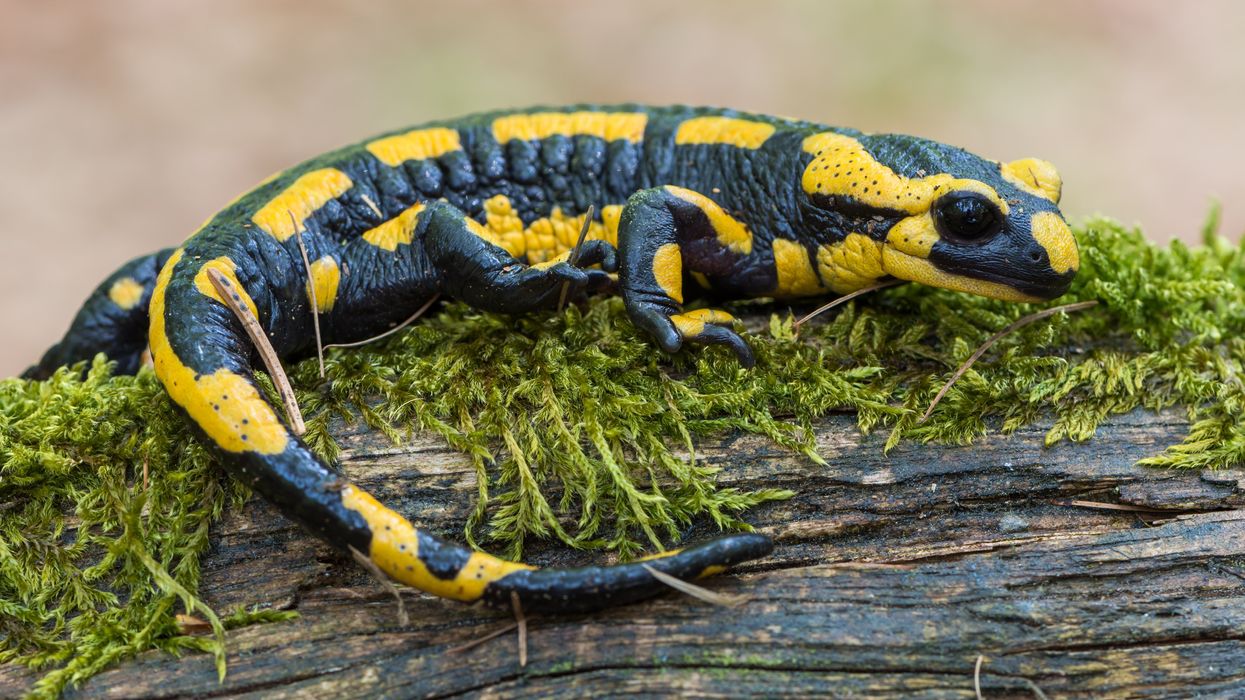

All organisms can repair damaged tissue, but none do it better than salamanders and newts. A surprising area of science could tell us how they manage this feat - and perhaps even help us develop a similar ability.
All organisms have the capacity to repair or regenerate tissue damage. None can do it better than salamanders or newts, which can regenerate an entire severed limb.
That feat has amazed and delighted man from the dawn of time and led to endless attempts to understand how it happens – and whether we can control it for our own purposes. An exciting new clue toward that understanding has come from a surprising source: research on the decline of cells, called cellular senescence.
Senescence is the last stage in the life of a cell. Whereas some cells simply break up or wither and die off, others transition into a zombie-like state where they can no longer divide. In this liminal phase, the cell still pumps out many different molecules that can affect its neighbors and cause low grade inflammation. Senescence is associated with many of the declining biological functions that characterize aging, such as inflammation and genomic instability.
Oddly enough, newts are one of the few species that do not accumulate senescent cells as they age, according to research over several years by Maximina Yun. A research group leader at the Center for Regenerative Therapies Dresden and the Max Planck Institute of Molecular and Cell Biology and Genetics, in Dresden, Germany, Yun discovered that senescent cells were induced at some stages of regeneration of the salamander limb, “and then, as the regeneration progresses, they disappeared, they were eliminated by the immune system,” she says. “They were present at particular times and then they disappeared.”
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states.
Previous research on senescence in aging had suggested, logically enough, that applying those cells to the stump of a newly severed salamander limb would slow or even stop its regeneration. But Yun stood that idea on its head. She theorized that senescent cells might also play a role in newt limb regeneration, and she tested it by both adding and removing senescent cells from her animals. It turned out she was right, as the newt limbs grew back faster than normal when more senescent cells were included.
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states, which could then be turned into progenitors, a cell type in between stem cells and specialized cells, needed to regrow the muscle tissue of the missing limb. “We think that this ability to dedifferentiate is intrinsically a big part of why salamanders can regenerate all these very complex structures, which other organisms cannot,” she explains.
Yun sees regeneration as a two part problem. First, the cells must be able to sense that their neighbors from the lost limb are not there anymore. Second, they need to be able to produce the intermediary progenitors for regeneration, , to form what is missing. “Molecularly, that must be encoded like a 3D map,” she says, otherwise the new tissue might grow back as a blob, or liver, or fin instead of a limb.
Wound healing
Another recent study, this time at the Mayo Clinic, provides evidence supporting the role of senescent cells in regeneration. Looking closely at molecules that send information between cells in the wound of a mouse, the researchers found that senescent cells appeared near the start of the healing process and then disappeared as healing progressed. In contrast, persistent senescent cells were the hallmark of a chronic wound that did not heal properly. The function and significance of senescence cells depended on both the timing and the context of their environment.
The paper suggests that senescent cells are not all the same. That has become clearer as researchers have been able to identify protein markers on the surface of some senescent cells. The patterns of these proteins differ for some senescent cells compared to others. In biology, such physical differences suggest functional differences, so it is becoming increasingly likely there are subsets of senescent cells with differing functions that have not yet been identified.
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes.
Scientists initially thought that senescent cells couldn’t play a role in regeneration because they could no longer reproduce, says Anthony Atala, a practicing surgeon and bioengineer who leads the Wake Forest Institute for Regenerative Medicine in North Carolina. But Yun’s study points in the other direction. “What this paper shows clearly is that these cells have the potential to be involved in tissue regeneration [in newts]. The question becomes, will these cells be able to do the same in humans.”
As our knowledge of senescent cells increases, Atala thinks we need to embrace a new analogy to help understand them: humans in retirement. They “have acquired a lot of wisdom throughout their whole life and they can help younger people and mentor them to grow to their full potential. We're seeing the same thing with these cells,” he says. They are no longer putting energy into their own reproduction, but the signaling molecules they secrete “can help other cells around them to regenerate.”
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes. If so, it seems that our genes are unable to express this ability, perhaps as part of a tradeoff in acquiring other traits. It is a fertile area of research.
Dedifferentiation is likely to become an important process in the field of regenerative medicine. One extreme example: a lab has been able to turn back the clock and reprogram adult male skin cells into female eggs, a potential milestone in reproductive health. It will be more difficult to control just how far back one wishes to go in the cell's dedifferentiation – part way or all the way back into a stem cell – and then direct it down a different developmental pathway. Yun is optimistic we can learn these tricks from newts.
Senolytics
A growing field of research is using drugs called senolytics to remove senescent cells and slow or even reverse disease of aging.
“Senolytics are great, but senolytics target different types of senescence,” Yun says. “If senescent cells have positive effects in the context of regeneration, of wound healing, then maybe at the beginning of the regeneration process, you may not want to take them out for a little while.”
“If you look at pretty much all biological systems, too little or too much of something can be bad, you have to be in that central zone” and at the proper time, says Atala. “That's true for proteins, sugars, and the drugs that you take. I think the same thing is true for these cells. Why would they be different?”
Our growing understanding that senescence is not a single thing but a variety of things likely means that effective senolytic drugs will not resemble a single sledge hammer but more a carefully manipulated scalpel where some types of senescent cells are removed while others are added. Combinations and timing could be crucial, meaning the difference between regenerating healthy tissue, a scar, or worse.
Scientists are making machines, wearable and implantable, to act as kidneys
Recent advancements in engineering mean that the first preclinical trials for an artificial kidney could happen soon.
Like all those whose kidneys have failed, Scott Burton’s life revolves around dialysis. For nearly two decades, Burton has been hooked up (or, since 2020, has hooked himself up at home) to a dialysis machine that performs the job his kidneys normally would. The process is arduous, time-consuming, and expensive. Except for a brief window before his body rejected a kidney transplant, Burton has depended on machines to take the place of his kidneys since he was 12-years-old. His whole life, the 39-year-old says, revolves around dialysis.
“Whenever I try to plan anything, I also have to plan my dialysis,” says Burton says, who works as a freelance videographer and editor. “It’s a full-time job in itself.”
Many of those on dialysis are in line for a kidney transplant that would allow them to trade thrice-weekly dialysis and strict dietary limits for a lifetime of immunosuppressants. Burton’s previous transplant means that his body will likely reject another donated kidney unless it matches perfectly—something he’s not counting on. It’s why he’s enthusiastic about the development of artificial kidneys, small wearable or implantable devices that would do the job of a healthy kidney while giving users like Burton more flexibility for traveling, working, and more.
Still, the devices aren’t ready for testing in humans—yet. But recent advancements in engineering mean that the first preclinical trials for an artificial kidney could happen soon, according to Jonathan Himmelfarb, a nephrologist at the University of Washington.
“It would liberate people with kidney failure,” Himmelfarb says.
An engineering marvel
Compared to the heart or the brain, the kidney doesn’t get as much respect from the medical profession, but its job is far more complex. “It does hundreds of different things,” says UCLA’s Ira Kurtz.
Kurtz would know. He’s worked as a nephrologist for 37 years, devoting his career to helping those with kidney disease. While his colleagues in cardiology and endocrinology have seen major advances in the development of artificial hearts and insulin pumps, little has changed for patients on hemodialysis. The machines remain bulky and require large volumes of a liquid called dialysate to remove toxins from a patient’s blood, along with gallons of purified water. A kidney transplant is the next best thing to someone’s own, functioning organ, but with over 600,000 Americans on dialysis and only about 100,000 kidney transplants each year, most of those in kidney failure are stuck on dialysis.
Part of the lack of progress in artificial kidney design is the sheer complexity of the kidney’s job. Each of the 45 different cell types in the kidney do something different.
Part of the lack of progress in artificial kidney design is the sheer complexity of the kidney’s job. To build an artificial heart, Kurtz says, you basically need to engineer a pump. An artificial pancreas needs to balance blood sugar levels with insulin secretion. While neither of these tasks is simple, they are fairly straightforward. The kidney, on the other hand, does more than get rid of waste products like urea and other toxins. Each of the 45 different cell types in the kidney do something different, helping to regulate electrolytes like sodium, potassium, and phosphorous; maintaining blood pressure and water balance; guiding the body’s hormonal and inflammatory responses; and aiding in the formation of red blood cells.

There's been little progress for patients during Ira Kurtz's 37 years as a nephrologist. Artificial kidneys would change that.
UCLA
Dialysis primarily filters waste, and does so well enough to keep someone alive, but it isn’t a true artificial kidney because it doesn’t perform the kidney’s other jobs, according to Kurtz, such as sensing levels of toxins, wastes, and electrolytes in the blood. Due to the size and water requirements of existing dialysis machines, the equipment isn’t portable. Physicians write a prescription for a certain duration of dialysis and assess how well it’s working with semi-regular blood tests. The process of dialysis itself, however, is conducted blind. Doctors can’t tell how much dialysis a patient needs based on kidney values at the time of treatment, says Meera Harhay, a nephrologist at Drexel University in Philadelphia.
But it’s the impact of dialysis on their day-to-day lives that creates the most problems for patients. Only one-quarter of those on dialysis are able to remain employed (compared to 85% of similar-aged adults), and many report a low quality of life. Having more flexibility in life would make a major different to her patients, Harhay says.
“Almost half their week is taken up by the burden of their treatment. It really eats away at their freedom and their ability to do things that add value to their life,” she says.
Art imitates life
The challenge for artificial kidney designers was how to compress the kidney’s natural functions into a portable, wearable, or implantable device that wouldn’t need constant access to gallons of purified and sterilized water. The other universal challenge they faced was ensuring that any part of the artificial kidney that would come in contact with blood was kept germ-free to prevent infection.
As part of the 2021 KidneyX Prize, a partnership between the U.S. Department of Health and Human Services and the American Society of Nephrology, inventors were challenged to create prototypes for artificial kidneys. Himmelfarb’s team at the University of Washington’s Center for Dialysis Innovation won the prize by focusing on miniaturizing existing technologies to create a portable dialysis machine. The backpack sized AKTIV device (Ambulatory Kidney to Increase Vitality) will recycle dialysate in a closed loop system that removes urea from blood and uses light-based chemical reactions to convert the urea to nitrogen and carbon dioxide, which allows the dialysate to be recirculated.
Himmelfarb says that the AKTIV can be used when at home, work, or traveling, which will give users more flexibility and freedom. “If you had a 30-pound device that you could put in the overhead bins when traveling, you could go visit your grandkids,” he says.
Kurtz’s team at UCLA partnered with the U.S. Kidney Research Corporation and Arkansas University to develop a dialysate-free desktop device (about the size of a small printer) as the first phase of a progression that will he hopes will lead to something small and implantable. Part of the reason for the artificial kidney’s size, Kurtz says, is the number of functions his team are cramming into it. Not only will it filter urea from blood, but it will also use electricity to help regulate electrolyte levels in a process called electrodeionization. Kurtz emphasizes that these additional functions are what makes his design a true artificial kidney instead of just a small dialysis machine.
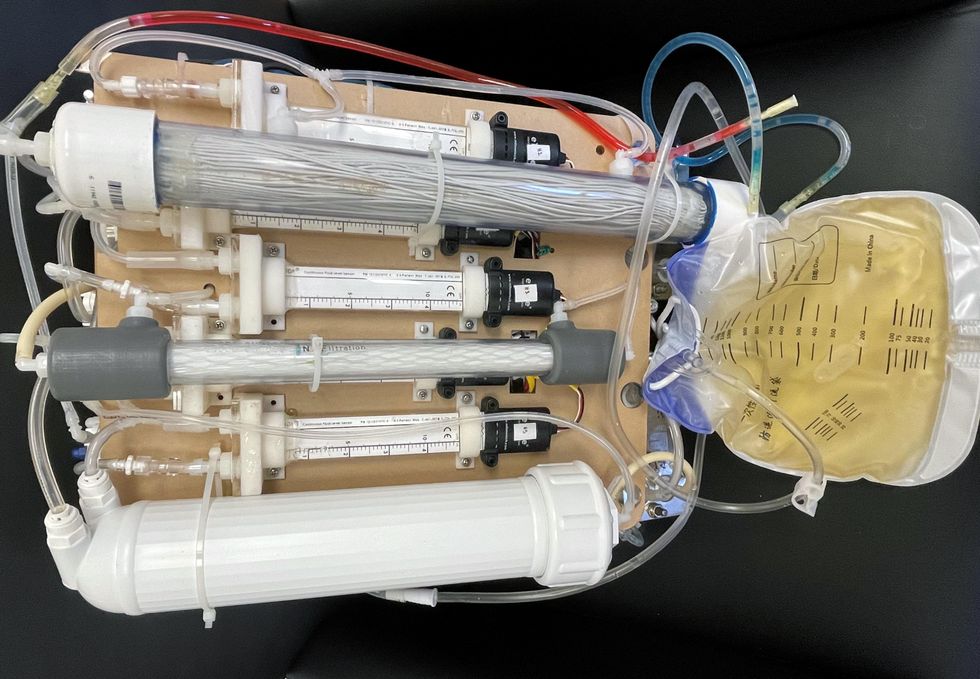

One version of an artificial kidney.
UCLA
“It doesn't have just a static function. It has a bank of sensors that measure chemicals in the blood and feeds that information back to the device,” Kurtz says.
Other startups are getting in on the game. Nephria Bio, a spinout from the South Korean-based EOFlow, is working to develop a wearable dialysis device, akin to an insulin pump, that uses miniature cartridges with nanomaterial filters to clean blood (Harhay is a scientific advisor to Nephria). Ian Welsford, Nephria’s co-founder and CTO, says that the device’s design means that it can also be used to treat acute kidney injuries in resource-limited settings. These potentials have garnered interest and investment in artificial kidneys from the U.S. Department of Defense.
For his part, Burton is most interested in an implantable device, as that would give him the most freedom. Even having a regular outpatient procedure to change batteries or filters would be a minor inconvenience to him.
“Being plugged into a machine, that’s not mimicking life,” he says.
This article was first published by Leaps.org on May 5, 2022.
With this new technology, hospitals and pharmacies could make vaccines and medicines onsite
New research focuses on methods that could change medicine-making worldwide. The scientists propose bursting cells open, removing their DNA and using the cellular gears inside to make therapies.
Most modern biopharmaceutical medicines are produced by workhorse cells—typically bacterial but sometimes mammalian. The cells receive the synthesizing instructions on a snippet of a genetic code, which they incorporate into their DNA. The cellular machinery—ribosomes, RNAs, polymerases, and other compounds—read and use these instructions to build the medicinal molecules, which are harvested and administered to patients.
Although a staple of modern pharma, this process is complex and expensive. One must first insert the DNA instructions into the cells, which they may or may not uptake. One then must grow the cells, keeping them alive and well, so that they produce the required therapeutics, which then must be isolated and purified. To make this at scale requires massive bioreactors and big factories from where the drugs are distributed—and may take a while to arrive where they’re needed. “The pandemic showed us that this method is slow and cumbersome,” says Govind Rao, professor of biochemical engineering who directs the Center for Advanced Sensor Technology at the University of Maryland, Baltimore County (UMBC). “We need better methods that can work faster and can work locally where an outbreak is happening.”
Rao and his team of collaborators, which spans multiple research institutions, believe they have a better approach that may change medicine-making worldwide. They suggest forgoing the concept of using living cells as medicine-producers. Instead, they propose breaking the cells and using the remaining cellular gears for assembling the therapeutic compounds. Instead of inserting the DNA into living cells, the team burst them open, and removed their DNA altogether. Yet, the residual molecular machinery of ribosomes, polymerases and other cogwheels still functioned the way it would in a cell. “Now if you drop your DNA drug-making instructions into that soup, this machinery starts making what you need,” Rao explains. “And because you're no longer worrying about living cells, it becomes much simpler and more efficient.” The collaborators detail their cell-free protein synthesis or CFPS method in their recent paper published in preprint BioAxiv.
While CFPS does not use living cells, it still needs the basic building blocks to assemble proteins from—such as amino acids, nucleotides and certain types of enzymes. These are regularly added into this “soup” to keep the molecular factory chugging. “We just mix everything in as a batch and we let it integrate,” says James Robert Swartz, professor of chemical engineering and bioengineering at Stanford University and co-author of the paper. “And we make sure that we provide enough oxygen.” Rao likens the process to making milk from milk powder.
For a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology.
The idea of a cell-free protein synthesis is older than one might think. Swartz first experimented with it around 1997, when he was a chemical engineer at Genentech. While working on engineering bacteria to make pharmaceuticals, he discovered that there was a limit to what E. coli cells, the workhorse darling of pharma, could do. For example, it couldn’t grow and properly fold some complex proteins. “We tried many genetic engineering approaches, many fermentation, development, and environmental control approaches,” Swartz recalls—to no avail.
“The organism had its own agenda,” he quips. “And because everything was happening within the organism, we just couldn't really change those conditions very easily. Some of them we couldn’t change at all—we didn’t have control.”
It was out of frustration with the defiant bacteria that a new idea took hold. Could the cells be opened instead, so that the protein-forming reactions could be influenced more easily? “Obviously, we’d lose the ability for them to reproduce,” Swartz says. But that also meant that they no longer needed to keep the cells alive and could focus on making the specific reactions happen. “We could take the catalysts, the enzymes, and the more complex catalysts and activate them, make them work together, much as they would in a living cell, but the way we wanted.”
In 1998, Swartz joined Stanford, and began perfecting the biochemistry of the cell-free method, identifying the reactions he wanted to foster and stopping those he didn’t want. He managed to make the idea work, but for a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology. For their BioArxiv paper, the team tested the method by growing a specific antiviral protein called griffithsin.
First identified by Barry O’Keefe at National Cancer Institute over a decade ago, griffithsin is an antiviral known to interfere with many viruses’ ability to enter cells—including HIV, SARS, SARS-CoV-2, MERS and others. Originally isolated from the red algae Griffithsia, it works differently from antibodies and antibody cocktails.
Most antiviral medicines tend to target the specific receptors that viruses use to gain entry to the cells they infect. For example, SARS-CoV-2 uses the infamous spike protein to latch onto the ACE2 receptor of mammalian cells. The antibodies or other antiviral molecules stick to the spike protein, shutting off its ability to cling onto the ACE2 receptors. Unfortunately, the spike proteins mutate very often, so the medicines lose their potency. On the contrary, griffithsin has the ability to cling to the different parts of viral shells called capsids—namely to the molecules of mannose, a type of sugar. That extra stuff, glued all around the capsid like dead weight, makes it impossible for the virus to squeeze into the cell.
“Every time we have a vaccine or an antibody against a specific SARS-CoV-2 strain, that strain then mutates and so you lose efficacy,” Rao explains. “But griffithsin molecules glom onto the viral capsid, so the capsid essentially becomes a sticky mess and can’t enter the cell.” Mannose molecules also don’t mutate as easily as viruses’ receptors, so griffithsin-based antivirals do not have to be constantly updated. And because mannose molecules are found on many viruses’ capsids, it makes griffithsin “a universal neutralizer,” Rao explains.
“When griffithsin was discovered, we recognized that it held a lot of promise as a potential antiviral agent,” O’Keefe says. In 2010, he published a paper about griffithsin efficacy in neutralizing viruses of the corona family—after the first SARS outbreak in the early 2000s, the scientific community was interested in such antivirals. Yet, griffithsin is still not available as an off-the-shelf product. So during the Covid pandemic, the team experimented with synthesizing griffithsin using the cell-free production method. They were able to generate potent griffithsin in less than 24 hours without having to grow living cells.
The antiviral protein isn't the only type of medicine that can be made cell-free. The proteins needed for vaccine production could also be made the same way. “Such portable, on-demand drug manufacturing platforms can produce antiviral proteins within hours, making them ideal for combating future pandemics,” Rao says. “We would be able to stop the pandemic before it spreads.”
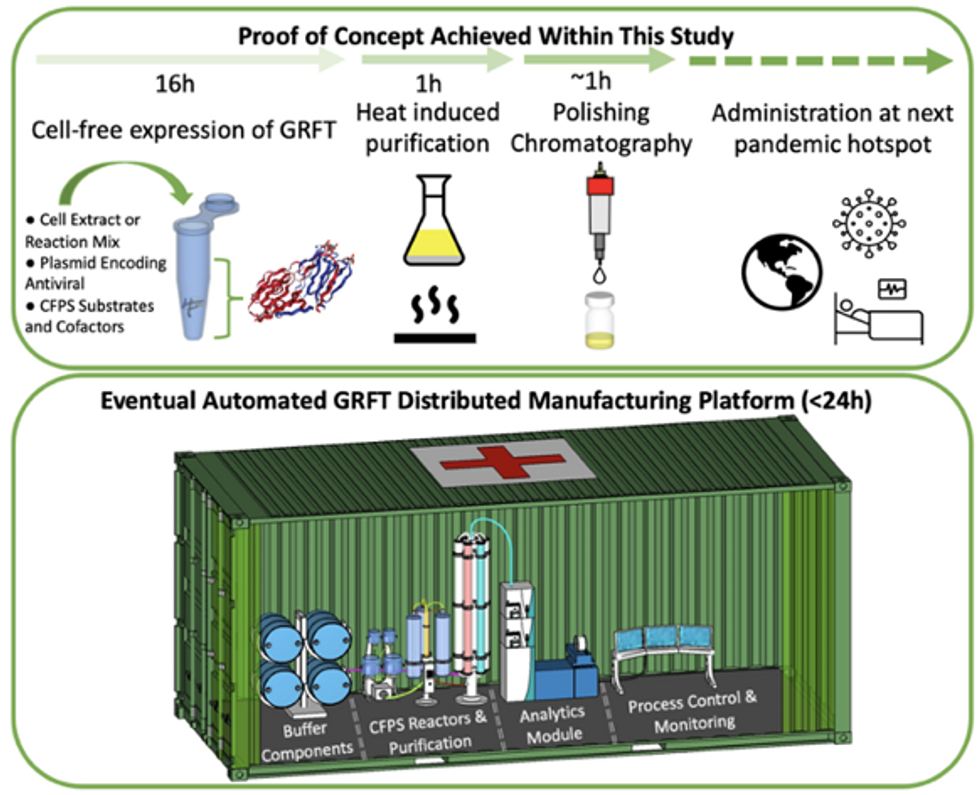
Top: Describes the process used in the study. Bottom: Describes how the new medicines and vaccines could be made at the site of a future viral outbreak.
Image courtesy of Rao and team, sourced from An approach to rapid distributed manufacturing of broad spectrumanti-viral griffithsin using cell-free systems to mitigate pandemics.
Rao’s idea is to perfect the technology to the point that any hospital or pharmacy can load up the media containing molecular factories, mix up the required amino acids, nucleotides and enzymes, and harvest the meds within hours. That will allow making medicines onsite and on demand. “That would be a self-contained production unit, so that you could just ship the production wherever the pandemic is breaking out,” says Swartz.
These units and the meds they produce, will, of course, have to undergo rigorous testing. “The biggest hurdles will be validating these against conventional technology,” Rao says. The biotech industry is risk-averse and prefers the familiar methods. But if this approach works, it may go beyond emergency situations and revolutionize the medicine-making paradigm even outside hospitals and pharmacies. Rao hopes that someday the method might become so mainstream that people may be able to buy and operate such reactors at home. “You can imagine a diabetic patient making insulin that way, or some other drugs,” Rao says. It would work not unlike making baby formula from the mere white powder. Just add water—and some oxygen, too.
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.

