Food Poisoning Outbreaks Are Still A Problem. Powerful Tech Is Fighting Back.

The latest tools, like whole genome sequencing, are allowing food outbreaks to be identified and stopped more quickly.
With the pandemic at the forefront of everyone's minds, many people have wondered if food could be a source of coronavirus transmission. Luckily, that "seems unlikely," according to the CDC, but foodborne illnesses do still sicken a whopping 48 million people per year.
Whole genome sequencing is like "going from an eight-bit image—maybe like what you would see in Minecraft—to a high definition image."
In normal times, when there isn't a historic global health crisis infecting millions and affecting the lives of billions, foodborne outbreaks are real and frightening, potentially deadly, and can cause widespread fear of particular foods. Think of Romaine lettuce spreading E. coli last year— an outbreak that infected more than 500 people and killed eight—or peanut butter spreading salmonella in 2008, which infected 167 people.
The technologies available to detect and prevent the next foodborne disease outbreak have improved greatly over the past 30-plus years, particularly during the past decade, and better, more nimble technologies are being developed, according to experts in government, academia, and private industry. The key to advancing detection of harmful foodborne pathogens, they say, is increasing speed and portability of detection, and the precision of that detection.
Getting to Rapid Results
Researchers at Purdue University have recently developed a lateral flow assay that, with the help of a laser, can detect toxins and pathogenic E. coli. Lateral flow assays are cheap and easy to use; a good example is a home pregnancy test. You place a liquid or liquefied sample on a piece of paper designed to detect a single substance and soon after you get the results in the form of a colored line: yes or no.
"They're a great portable tool for us for food contaminant detection," says Carmen Gondhalekar, a fifth-year biomedical engineering graduate student at Purdue. "But one of the areas where paper-based lateral flow assays could use improvement is in multiplexing capability and their sensitivity."
J. Paul Robinson, a professor in Purdue's Colleges of Veterinary Medicine and Engineering, and Gondhalekar's advisor, agrees. "One of the fundamental problems that we have in detection is that it is hard to identify pathogens in complex samples," he says.
When it comes to foodborne disease outbreaks, you don't always know what substance you're looking for, so an assay made to detect only a single substance isn't always effective. The goal of the project at Purdue is to make assays that can detect multiple substances at once.
These assays would be more complex than a pregnancy test. As detailed in Gondhalekar's recent paper, a laser pulse helps create a spectral signal from the sample on the assay paper, and the spectral signal is then used to determine if any unique wavelengths associated with one of several toxins or pathogens are present in the sample. Though the handheld technology has yet to be built, the idea is that the results would be given on the spot. So someone in the field trying to track the source of a Salmonella infection could, for instance, put a suspected lettuce sample on the assay and see if it has the pathogen on it.
"What our technology is designed to do is to give you a rapid assessment of the sample," says Robinson. "The goal here is speed."
Seeing the Pathogen in "High-Def"
"One in six Americans will get a foodborne illness every year," according to Dr. Heather Carleton, a microbiologist at the Centers for Disease Control and Prevention's Enteric Diseases Laboratory Branch. But not every foodborne outbreak makes the news. In 2017 alone, the CDC monitored between 18 and 37 foodborne poison clusters per week and investigated 200 multi-state clusters. Hardboiled eggs, ground beef, chopped salad kits, raw oysters, frozen tuna, and pre-cut melon are just a taste of the foods that were investigated last year for different strains of listeria, salmonella, and E. coli.
At the heart of the CDC investigations is PulseNet, a national network of laboratories that uses DNA fingerprinting to detect outbreaks at local and regional levels. This is how it works: When a patient gets sick—with symptoms like vomiting and fever, for instance—they will go to a hospital or clinic for treatment. Since we're talking about foodborne illnesses, a clinician will likely take a stool sample from the patient and send it off to a laboratory to see if there is a foodborne pathogen, like salmonella, E. Coli, or another one. If it does contain a potentially harmful pathogen, then a bacterial isolate of that identified sample is sent to a regional public health lab so that whole genome sequencing can be performed.
Whole genome sequencing can differentiate "virtually all" strains of foodborne pathogens, no matter the species, according to the FDA.
Whole genome sequencing is a method for reading the entire genome of a bacterial isolate (or from any organism, for that matter). Instead of working with a couple dozen data points, now you're working with millions of base pairs. Carleton likes to describe it as "going from an eight-bit image—maybe like what you would see in Minecraft—to a high definition image," she says. "It's really an evolution of how we detect foodborne illnesses and identify outbreaks."
If the bacterial isolate matches another in the CDC's database, this means there could be a potential outbreak and an investigation may be started, with the goal of tracking the pathogen to its source.
Whole genome sequencing has been a relatively recent shift in foodborne disease detection. For more than 20 years, the standard technique for analyzing pathogens in foodborne disease outbreaks was pulsed-field gel electrophoresis. This method creates a DNA fingerprint for each sample in the form of a pattern of about 15-30 "bands," with each band representing a piece of DNA. Researchers like Carleton can use this fingerprint to see if two samples are from the same bacteria. The problem is that 15-30 bands are not enough to differentiate all isolates. Some isolates whose bands look very similar may actually come from different sources and some whose bands look different may be from the same source. But if you can see the entire DNA fingerprint, then you don't have that issue. That's where whole genome sequencing comes in.
Although the PulseNet team had piloted whole genome sequencing as early as 2013, it wasn't until July of last year that the transition to using whole genome sequencing for all pathogens was complete. Though whole genome sequencing requires far more computing power to generate, analyze, and compare those millions of data points, the payoff is huge.
Stopping Outbreaks Sooner
The U.S. Food and Drug Administration (FDA) acquired their first whole genome sequencers in 2008, according to Dr. Eric Brown, the Director of the Division of Microbiology in the FDA's Office of Regulatory Science. Since then, through their GenomeTrakr program, a network of more than 60 domestic and international labs, the FDA has sequenced and publicly shared more than 400,000 isolates. "The impact of what whole genome sequencing could do to resolve a foodborne outbreak event was no less impactful than when NASA turned on the Hubble Telescope for the first time," says Brown.
Whole genome sequencing has helped identify strains of Salmonella that prior methods were unable to differentiate. In fact, whole genome sequencing can differentiate "virtually all" strains of foodborne pathogens, no matter the species, according to the FDA. This means it takes fewer clinical cases—fewer sick people—to detect and end an outbreak.
And perhaps the largest benefit of whole genome sequencing is that these detailed sequences—the millions of base pairs—can imply geographic location. The genomic information of bacterial strains can be different depending on the area of the country, helping these public health agencies eventually track the source of outbreaks—a restaurant, a farm, a food-processing center.
Coming Soon: "Lab in a Backpack"
Now that whole genome sequencing has become the go-to technology of choice for analyzing foodborne pathogens, the next step is making the process nimbler and more portable. Putting "the lab in a backpack," as Brown says.
The CDC's Carleton agrees. "Right now, the sequencer we use is a fairly big box that weighs about 60 pounds," she says. "We can't take it into the field."
A company called Oxford Nanopore Technologies is developing handheld sequencers. Their devices are meant to "enable the sequencing of anything by anyone anywhere," according to Dan Turner, the VP of Applications at Oxford Nanopore.
"The sooner that we can see linkages…the sooner the FDA gets in action to mitigate the problem and put in some kind of preventative control."
"Right now, sequencing is very much something that is done by people in white coats in laboratories that are set up for that purpose," says Turner. Oxford Nanopore would like to create a new, democratized paradigm.
The FDA is currently testing these types of portable sequencers. "We're very excited about it. We've done some pilots, to be able to do that sequencing in the field. To actually do it at a pond, at a river, at a canal. To do it on site right there," says Brown. "This, of course, is huge because it means we can have real-time sequencing capability to stay in step with an actual laboratory investigation in the field."
"The timeliness of this information is critical," says Marc Allard, a senior biomedical research officer and Brown's colleague at the FDA. "The sooner that we can see linkages…the sooner the FDA gets in action to mitigate the problem and put in some kind of preventative control."
At the moment, the world is rightly focused on COVID-19. But as the danger of one virus subsides, it's only a matter of time before another pathogen strikes. Hopefully, with new and advancing technology like whole genome sequencing, we can stop the next deadly outbreak before it really gets going.
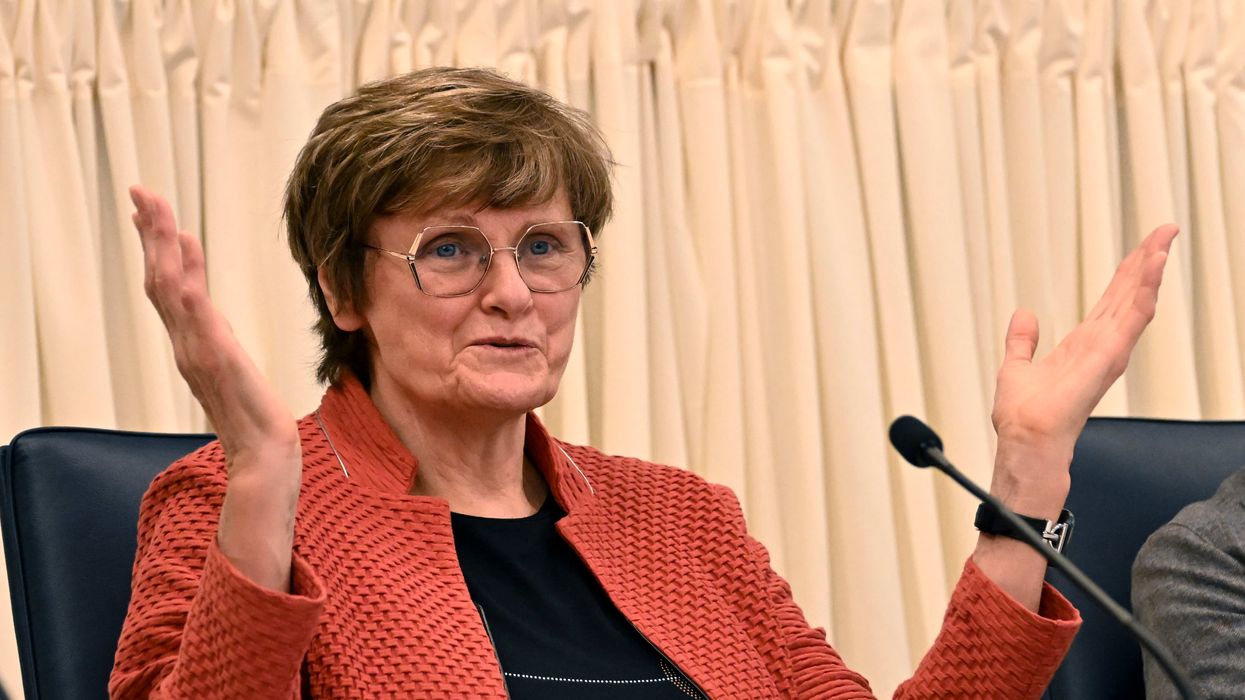
Nobel Prize goes to technology for mRNA vaccines
Katalin Karikó, pictured, and Drew Weissman won the Nobel Prize for advances in mRNA research that led to the first Covid vaccines.
When Drew Weissman received a call from Katalin Karikó in the early morning hours this past Monday, he assumed his longtime research partner was calling to share a nascent, nagging idea. Weissman, a professor of medicine at the Perelman School of Medicine at the University of Pennsylvania, and Karikó, a professor at Szeged University and an adjunct professor at UPenn, both struggle with sleep disturbances. Thus, middle-of-the-night discourses between the two, often over email, has been a staple of their friendship. But this time, Karikó had something more pressing and exciting to share: They had won the 2023 Nobel Prize in Physiology or Medicine.
The work for which they garnered the illustrious award and its accompanying $1,000,000 cash windfall was completed about two decades ago, wrought through long hours in the lab over many arduous years. But humanity collectively benefited from its life-saving outcome three years ago, when both Moderna and Pfizer/BioNTech’s mRNA vaccines against COVID were found to be safe and highly effective at preventing severe disease. Billions of doses have since been given out to protect humans from the upstart viral scourge.
“I thought of going somewhere else, or doing something else,” said Katalin Karikó. “I also thought maybe I’m not good enough, not smart enough. I tried to imagine: Everything is here, and I just have to do better experiments.”
Unlocking the power of mRNA
Weissman and Karikó unlocked mRNA vaccines for the world back in the early 2000s when they made a key breakthrough. Messenger RNA molecules are essentially instructions for cells’ ribosomes to make specific proteins, so in the 1980s and 1990s, researchers started wondering if sneaking mRNA into the body could trigger cells to manufacture antibodies, enzymes, or growth agents for protecting against infection, treating disease, or repairing tissues. But there was a big problem: injecting this synthetic mRNA triggered a dangerous, inflammatory immune response resulting in the mRNA’s destruction.
While most other researchers chose not to tackle this perplexing problem to instead pursue more lucrative and publishable exploits, Karikó stuck with it. The choice sent her academic career into depressing doldrums. Nobody would fund her work, publications dried up, and after six years as an assistant professor at the University of Pennsylvania, Karikó got demoted. She was going backward.
“I thought of going somewhere else, or doing something else,” Karikó told Stat in 2020. “I also thought maybe I’m not good enough, not smart enough. I tried to imagine: Everything is here, and I just have to do better experiments.”
A tale of tenacity
Collaborating with Drew Weissman, a new professor at the University of Pennsylvania, in the late 1990s helped provide Karikó with the tenacity to continue. Weissman nurtured a goal of developing a vaccine against HIV-1, and saw mRNA as a potential way to do it.
“For the 20 years that we’ve worked together before anybody knew what RNA is, or cared, it was the two of us literally side by side at a bench working together,” Weissman said in an interview with Adam Smith of the Nobel Foundation.
In 2005, the duo made their 2023 Nobel Prize-winning breakthrough, detailing it in a relatively small journal, Immunity. (Their paper was rejected by larger journals, including Science and Nature.) They figured out that chemically modifying the nucleoside bases that make up mRNA allowed the molecule to slip past the body’s immune defenses. Karikó and Weissman followed up that finding by creating mRNA that’s more efficiently translated within cells, greatly boosting protein production. In 2020, scientists at Moderna and BioNTech (where Karikó worked from 2013 to 2022) rushed to craft vaccines against COVID, putting their methods to life-saving use.
The future of vaccines
Buoyed by the resounding success of mRNA vaccines, scientists are now hurriedly researching ways to use mRNA medicine against other infectious diseases, cancer, and genetic disorders. The now ubiquitous efforts stand in stark contrast to Karikó and Weissman’s previously unheralded struggles years ago as they doggedly worked to realize a shared dream that so many others shied away from. Katalin Karikó and Drew Weissman were brave enough to walk a scientific path that very well could have ended in a dead end, and for that, they absolutely deserve their 2023 Nobel Prize.
This article originally appeared on Big Think, home of the brightest minds and biggest ideas of all time.

Scientists turn pee into power in Uganda
With conventional fuel cells as their model, researchers learned to use similar chemical reactions to make a fuel from microbes in pee.
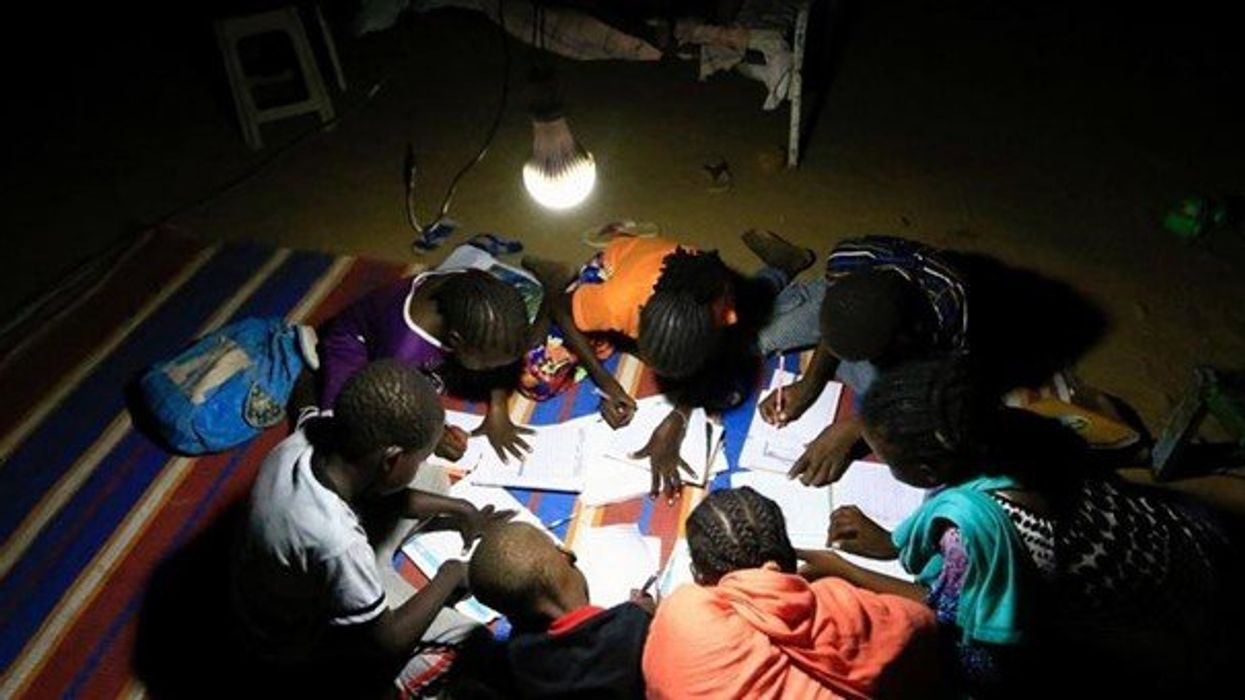
At the edge of a dirt road flanked by trees and green mountains outside the town of Kisoro, Uganda, sits the concrete building that houses Sesame Girls School, where girls aged 11 to 19 can live, learn and, at least for a while, safely use a toilet. In many developing regions, toileting at night is especially dangerous for children. Without electrical power for lighting, kids may fall into the deep pits of the latrines through broken or unsteady floorboards. Girls are sometimes assaulted by men who hide in the dark.
For the Sesame School girls, though, bright LED lights, connected to tiny gadgets, chased the fears away. They got to use new, clean toilets lit by the power of their own pee. Some girls even used the light provided by the latrines to study.
Urine, whether animal or human, is more than waste. It’s a cheap and abundant resource. Each day across the globe, 8.1 billion humans make 4 billion gallons of pee. Cows, pigs, deer, elephants and other animals add more. By spending money to get rid of it, we waste a renewable resource that can serve more than one purpose. Microorganisms that feed on nutrients in urine can be used in a microbial fuel cell that generates electricity – or "pee power," as the Sesame girls called it.
Plus, urine contains water, phosphorus, potassium and nitrogen, the key ingredients plants need to grow and survive. Human urine could replace about 25 percent of current nitrogen and phosphorous fertilizers worldwide and could save water for gardens and crops. The average U.S. resident flushes a toilet bowl containing only pee and paper about six to seven times a day, which adds up to about 3,500 gallons of water down per year. Plus cows in the U.S. produce 231 gallons of the stuff each year.
Pee power
A conventional fuel cell uses chemical reactions to produce energy, as electrons move from one electrode to another to power a lightbulb or phone. Ioannis Ieropoulos, a professor and chair of Environmental Engineering at the University of Southampton in England, realized the same type of reaction could be used to make a fuel from microbes in pee.
Bacterial species like Shewanella oneidensis and Pseudomonas aeruginosa can consume carbon and other nutrients in urine and pop out electrons as a result of their digestion. In a microbial fuel cell, one electrode is covered in microbes, immersed in urine and kept away from oxygen. Another electrode is in contact with oxygen. When the microbes feed on nutrients, they produce the electrons that flow through the circuit from one electrod to another to combine with oxygen on the other side. As long as the microbes have fresh pee to chomp on, electrons keep flowing. And after the microbes are done with the pee, it can be used as fertilizer.
These microbes are easily found in wastewater treatment plants, ponds, lakes, rivers or soil. Keeping them alive is the easy part, says Ieropoulos. Once the cells start producing stable power, his group sequences the microbes and keeps using them.
Like many promising technologies, scaling these devices for mass consumption won’t be easy, says Kevin Orner, a civil engineering professor at West Virginia University. But it’s moving in the right direction. Ieropoulos’s device has shrunk from the size of about three packs of cards to a large glue stick. It looks and works much like a AAA battery and produce about the same power. By itself, the device can barely power a light bulb, but when stacked together, they can do much more—just like photovoltaic cells in solar panels. His lab has produced 1760 fuel cells stacked together, and with manufacturing support, there’s no theoretical ceiling, he says.
Although pure urine produces the most power, Ieropoulos’s devices also work with the mixed liquids of the wastewater treatment plants, so they can be retrofit into urban wastewater utilities.
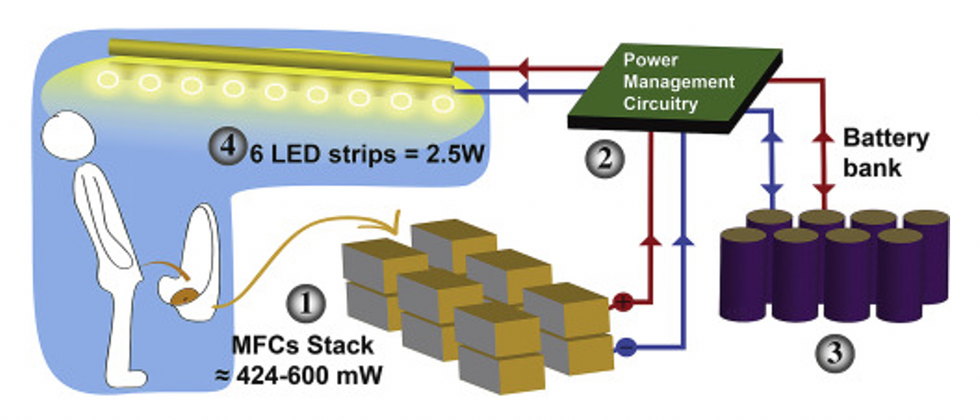
This image shows how the pee-powered system works. Pee feeds bacteria in the stack of fuel cells (1), which give off electrons (2) stored in parallel cylindrical cells (3). These cells are connected to a voltage regulator (4), which smooths out the electrical signal to ensure consistent power to the LED strips lighting the toilet.
Courtesy Ioannis Ieropoulos
Key to the long-term success of any urine reclamation effort, says Orner, is avoiding what he calls “parachute engineering”—when well-meaning scientists solve a problem with novel tech and then abandon it. “The way around that is to have either the need come from the community or to have an organization in a community that is committed to seeing a project operate and maintained,” he says.
Success with urine reclamation also depends on the economy. “If energy prices are low, it may not make sense to recover energy,” says Orner. “But right now, fertilizer prices worldwide are generally pretty high, so it may make sense to recover fertilizer and nutrients.” There are obstacles, too, such as few incentives for builders to incorporate urine recycling into new construction. And any hiccups like leaks or waste seepage will cost builders money and reputation. Right now, Orner says, the risks are just too high.
Despite the challenges, Ieropoulos envisions a future in which urine is passed through microbial fuel cells at wastewater treatment plants, retrofitted septic tanks, and building basements, and is then delivered to businesses to use as agricultural fertilizers. Although pure urine produces the most power, Ieropoulos’s devices also work with the mixed liquids of the wastewater treatment plants, so they can be retrofitted into urban wastewater utilities where they can make electricity from the effluent. And unlike solar cells, which are a common target of theft in some areas, nobody wants to steal a bunch of pee.
When Ieropoulos’s team returned to wrap up their pilot project 18 months later, the school’s director begged them to leave the fuel cells in place—because they made a major difference in students’ lives. “We replaced it with a substantial photovoltaic panel,” says Ieropoulos, They couldn’t leave the units forever, he explained, because of intellectual property reasons—their funders worried about theft of both the technology and the idea. But the photovoltaic replacement could be stolen, too, leaving the girls in the dark.
The story repeated itself at another school, in Nairobi, Kenya, as well as in an informal settlement in Durban, South Africa. Each time, Ieropoulos vowed to return. Though the pandemic has delayed his promise, he is resolute about continuing his work—it is a moral and legal obligation. “We've made a commitment to ourselves and to the pupils,” he says. “That's why we need to go back.”
Urine as fertilizer
Modern day industrial systems perpetuate the broken cycle of nutrients. When plants grow, they use up nutrients the soil. We eat the plans and excrete some of the nutrients we pass them into rivers and oceans. As a result, farmers must keep fertilizing the fields while our waste keeps fertilizing the waterways, where the algae, overfertilized with nitrogen, phosphorous and other nutrients grows out of control, sucking up oxygen that other marine species need to live. Few global communities remain untouched by the related challenges this broken chain create: insufficient clean water, food, and energy, and too much human and animal waste.
The Rich Earth Institute in Vermont runs a community-wide urine nutrient recovery program, which collects urine from homes and businesses, transports it for processing, and then supplies it as fertilizer to local farms.
One solution to this broken cycle is reclaiming urine and returning it back to the land. The Rich Earth Institute in Vermont is one of several organizations around the world working to divert and save urine for agricultural use. “The urine produced by an adult in one day contains enough fertilizer to grow all the wheat in one loaf of bread,” states their website.
Notably, while urine is not entirely sterile, it tends to harbor fewer pathogens than feces. That’s largely because urine has less organic matter and therefore less food for pathogens to feed on, but also because the urinary tract and the bladder have built-in antimicrobial defenses that kill many germs. In fact, the Rich Earth Institute says it’s safe to put your own urine onto crops grown for home consumption. Nonetheless, you’ll want to dilute it first because pee usually has too much nitrogen and can cause “fertilizer burn” if applied straight without dilution. Other projects to turn urine into fertilizer are in progress in Niger, South Africa, Kenya, Ethiopia, Sweden, Switzerland, The Netherlands, Australia, and France.
Eleven years ago, the Institute started a program that collects urine from homes and businesses, transports it for processing, and then supplies it as fertilizer to local farms. By 2021, the program included 180 donors producing over 12,000 gallons of urine each year. This urine is helping to fertilize hay fields at four partnering farms. Orner, the West Virginia professor, sees it as a success story. “They've shown how you can do this right--implementing it at a community level scale."

