Genetically Sequencing Healthy Babies Yielded Surprising Results

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
Thanks to safety cautions from the COVID-19 pandemic, a strain of influenza has been completely eliminated.
If you were one of the millions who masked up, washed your hands thoroughly and socially distanced, pat yourself on the back—you may have helped change the course of human history.
Scientists say that thanks to these safety precautions, which were introduced in early 2020 as a way to stop transmission of the novel COVID-19 virus, a strain of influenza has been completely eliminated. This marks the first time in human history that a virus has been wiped out through non-pharmaceutical interventions, such as vaccines.
The flu shot, explained
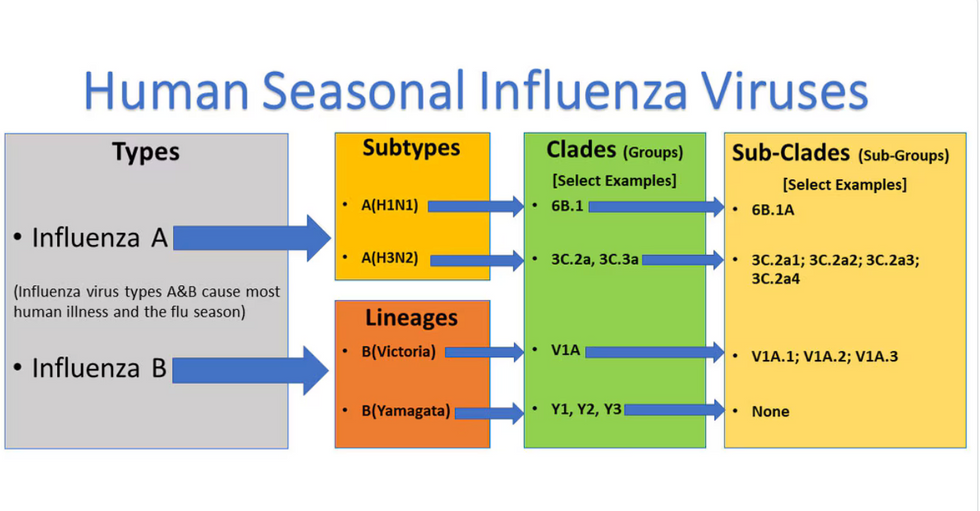
Influenza viruses type A and B are responsible for the majority of human illnesses and the flu season.
Centers for Disease Control
For more than a decade, flu shots have protected against two types of the influenza virus–type A and type B. While there are four different strains of influenza in existence (A, B, C, and D), only strains A, B, and C are capable of infecting humans, and only A and B cause pandemics. In other words, if you catch the flu during flu season, you’re most likely sick with flu type A or B.
Flu vaccines contain inactivated—or dead—influenza virus. These inactivated viruses can’t cause sickness in humans, but when administered as part of a vaccine, they teach a person’s immune system to recognize and kill those viruses when they’re encountered in the wild.
Each spring, a panel of experts gives a recommendation to the US Food and Drug Administration on which strains of each flu type to include in that year’s flu vaccine, depending on what surveillance data says is circulating and what they believe is likely to cause the most illness during the upcoming flu season. For the past decade, Americans have had access to vaccines that provide protection against two strains of influenza A and two lineages of influenza B, known as the Victoria lineage and the Yamagata lineage. But this year, the seasonal flu shot won’t include the Yamagata strain, because the Yamagata strain is no longer circulating among humans.
How Yamagata Disappeared

Flu surveillance data from the Global Initiative on Sharing All Influenza Data (GISAID) shows that the Yamagata lineage of flu type B has not been sequenced since April 2020.
Nature
Experts believe that the Yamagata lineage had already been in decline before the pandemic hit, likely because the strain was naturally less capable of infecting large numbers of people compared to the other strains. When the COVID-19 pandemic hit, the resulting safety precautions such as social distancing, isolating, hand-washing, and masking were enough to drive the virus into extinction completely.
Because the strain hasn’t been circulating since 2020, the FDA elected to remove the Yamagata strain from the seasonal flu vaccine. This will mark the first time since 2012 that the annual flu shot will be trivalent (three-component) rather than quadrivalent (four-component).
Should I still get the flu shot?
The flu shot will protect against fewer strains this year—but that doesn’t mean we should skip it. Influenza places a substantial health burden on the United States every year, responsible for hundreds of thousands of hospitalizations and tens of thousands of deaths. The flu shot has been shown to prevent millions of illnesses each year (more than six million during the 2022-2023 season). And while it’s still possible to catch the flu after getting the flu shot, studies show that people are far less likely to be hospitalized or die when they’re vaccinated.
Another unexpected benefit of dropping the Yamagata strain from the seasonal vaccine? This will possibly make production of the flu vaccine faster, and enable manufacturers to make more vaccines, helping countries who have a flu vaccine shortage and potentially saving millions more lives.
After his grandmother’s dementia diagnosis, one man invented a snack to keep her healthy and hydrated.
Founder Lewis Hornby and his grandmother Pat, sampling Jelly Drops—an edible gummy containing water and life-saving electrolytes.
On a visit to his grandmother’s nursing home in 2016, college student Lewis Hornby made a shocking discovery: Dehydration is a common (and dangerous) problem among seniors—especially those that are diagnosed with dementia.
Hornby’s grandmother, Pat, had always had difficulty keeping up her water intake as she got older, a common issue with seniors. As we age, our body composition changes, and we naturally hold less water than younger adults or children, so it’s easier to become dehydrated quickly if those fluids aren’t replenished. What’s more, our thirst signals diminish naturally as we age as well—meaning our body is not as good as it once was in letting us know that we need to rehydrate. This often creates a perfect storm that commonly leads to dehydration. In Pat’s case, her dehydration was so severe she nearly died.
When Lewis Hornby visited his grandmother at her nursing home afterward, he learned that dehydration especially affects people with dementia, as they often don’t feel thirst cues at all, or may not recognize how to use cups correctly. But while dementia patients often don’t remember to drink water, it seemed to Hornby that they had less problem remembering to eat, particularly candy.

Hornby wanted to create a solution for elderly people who struggled keeping their fluid intake up. He spent the next eighteen months researching and designing a solution and securing funding for his project. In 2019, Hornby won a sizable grant from the Alzheimer’s Society, a UK-based care and research charity for people with dementia and their caregivers. Together, through the charity’s Accelerator Program, they created a bite-sized, sugar-free, edible jelly drop that looked and tasted like candy. The candy, called Jelly Drops, contained 95% water and electrolytes—important minerals that are often lost during dehydration. The final product launched in 2020—and was an immediate success. The drops were able to provide extra hydration to the elderly, as well as help keep dementia patients safe, since dehydration commonly leads to confusion, hospitalization, and sometimes even death.
Not only did Jelly Drops quickly become a favorite snack among dementia patients in the UK, but they were able to provide an additional boost of hydration to hospital workers during the pandemic. In NHS coronavirus hospital wards, patients infected with the virus were regularly given Jelly Drops to keep their fluid levels normal—and staff members snacked on them as well, since long shifts and personal protective equipment (PPE) they were required to wear often left them feeling parched.
In April 2022, Jelly Drops launched in the United States. The company continues to donate 1% of its profits to help fund Alzheimer’s research.

