Genetically Sequencing Healthy Babies Yielded Surprising Results
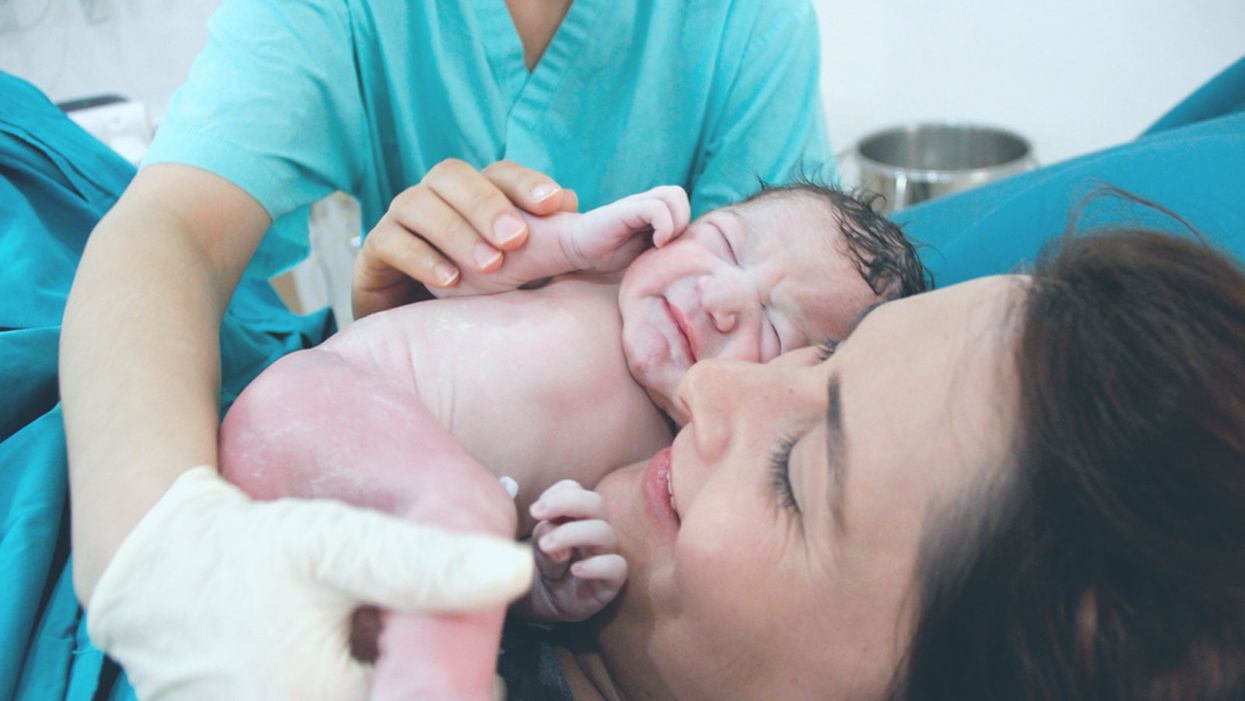

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
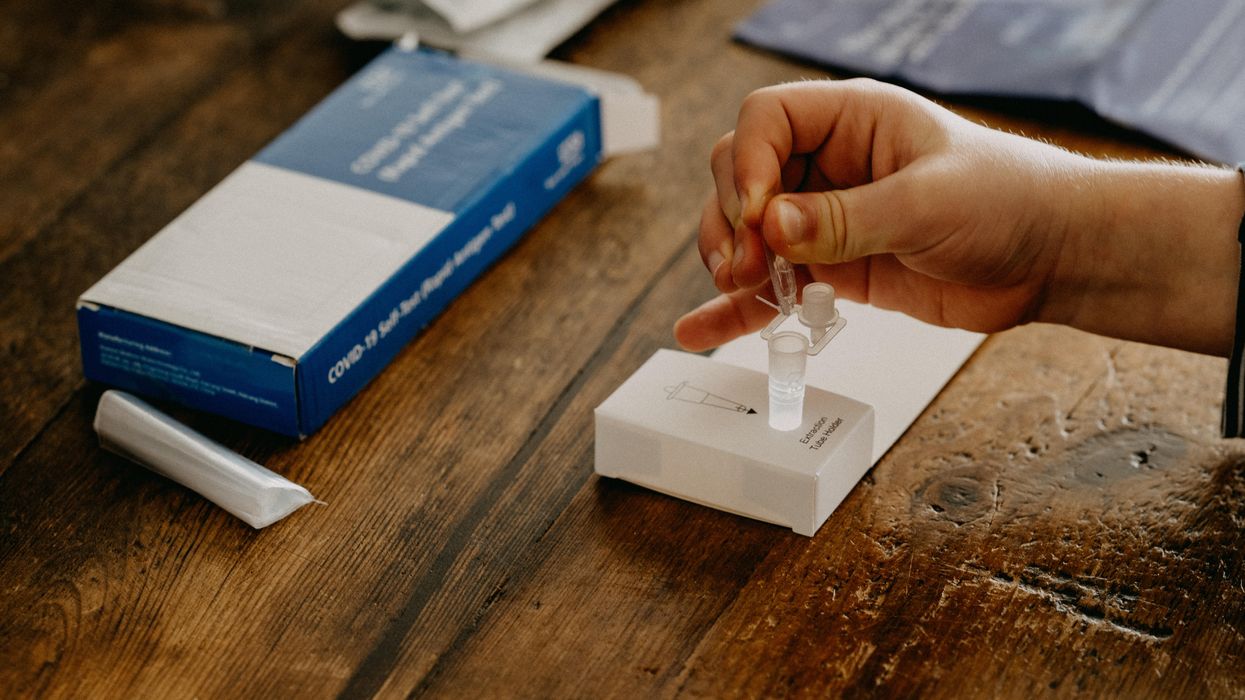
Biohackers Made a Cheap and Effective Home Covid Test -- But No One Is Allowed to Use It
A stock image of a home test for COVID-19.
Last summer, when fast and cheap Covid tests were in high demand and governments were struggling to manufacture and distribute them, a group of independent scientists working together had a bit of a breakthrough.
Working on the Just One Giant Lab platform, an online community that serves as a kind of clearing house for open science researchers to find each other and work together, they managed to create a simple, one-hour Covid test that anyone could take at home with just a cup of hot water. The group tested it across a network of home and professional laboratories before being listed as a semi-finalist team for the XPrize, a competition that rewards innovative solutions-based projects. Then, the group hit a wall: they couldn't commercialize the test.
They wanted to keep their project open source, making it accessible to people around the world, so they decided to forgo traditional means of intellectual property protection and didn't seek patents. (They couldn't afford lawyers anyway). And, as a loose-knit group that was not supported by a traditional scientific institution, working in community labs and homes around the world, they had no access to resources or financial support for manufacturing or distributing their test at scale.
But without ethical and regulatory approval for clinical testing, manufacture, and distribution, they were legally unable to create field tests for real people, leaving their inexpensive, $16-per-test, innovative product languishing behind, while other, more expensive over-the-counter tests made their way onto the market.
Who Are These Radical Scientists?
Independent, decentralized biomedical research has come of age. Also sometimes called DIYbio, biohacking, or community biology, depending on whom you ask, open research is today a global movement with thousands of members, from scientists with advanced degrees to middle-grade students. Their motivations and interests vary across a wide spectrum, but transparency and accessibility are key to the ethos of the movement. Teams are agile, focused on shoestring-budget R&D, and aim to disrupt business as usual in the ivory towers of the scientific establishment.
Ethics oversight is critical to ensuring that research is conducted responsibly, even by biohackers.
Initiatives developed within the community, such as Open Insulin, which hopes to engineer processes for affordable, small-batch insulin production, "Slybera," a provocative attempt to reverse engineer a $1 million dollar gene therapy, and the hundreds of projects posted on the collaboration platform Just One Giant Lab during the pandemic, all have one thing in common: to pursue testing in humans, they need an ethics oversight mechanism.
These groups, most of which operate collaboratively in community labs, homes, and online, recognize that some sort of oversight or guidance is useful—and that it's the right thing to do.
But also, and perhaps more immediately, they need it because federal rules require ethics oversight of any biomedical research that's headed in the direction of the consumer market. In addition, some individuals engaged in this work do want to publish their research in traditional scientific journals, which—you guessed it—also require that research has undergone an ethics evaluation. Ethics oversight is critical to ensuring that research is conducted responsibly, even by biohackers.
Bridging the Ethics Gap
The problem is that traditional oversight mechanisms, such as institutional review boards at government or academic research institutions, as well as the private boards utilized by pharmaceutical companies, are not accessible to most independent researchers. Traditional review boards are either closed to the public, or charge fees that are out of reach for many citizen science initiatives. This has created an "ethics gap" in nontraditional scientific research.
Biohackers are seen in some ways as the direct descendents of "white hat" computer hackers, or those focused on calling out security holes and contributing solutions to technical problems within self-regulating communities. In the case of health and biotechnology, those problems include both the absence of treatments and the availability of only expensive treatments for certain conditions. As the DIYbio community grows, there needs to be a way to provide assurance that, when the work is successful, the public is able to benefit from it eventually. The team that developed the one-hour Covid test found a potential commercial partner and so might well overcome the oversight hurdle, but it's been 14 months since they developed the test--and counting.
In short, without some kind of oversight mechanism for the work of independent biomedical researchers, the solutions they innovate will never have the opportunity to reach consumers.
In a new paper in the journal Citizen Science: Theory & Practice, we consider the issue of the ethics gap and ask whether ethics oversight is something nontraditional researchers want, and if so, what forms it might take. Given that individuals within these communities sometimes vehemently disagree with each other, is consensus on these questions even possible?
We learned that there is no "one size fits all" solution for ethics oversight of nontraditional research. Rather, the appropriateness of any oversight model will depend on each initiative's objectives, needs, risks, and constraints.
We also learned that nontraditional researchers are generally willing (and in some cases eager) to engage with traditional scientific, legal, and bioethics experts on ethics, safety, and related questions.
We suggest that these experts make themselves available to help nontraditional researchers build infrastructure for ethics self-governance and identify when it might be necessary to seek outside assistance.
Independent biomedical research has promise, but like any emerging science, it poses novel ethical questions and challenges. Existing research ethics and oversight frameworks may not be well-suited to answer them in every context, so we need to think outside the box about what we can create for the future. That process should begin by talking to independent biomedical researchers about their activities, priorities, and concerns with an eye to understanding how best to support them.
Christi Guerrini, JD, MPH studies biomedical citizen science and is an Associate Professor at Baylor College of Medicine. Alex Pearlman, MA, is a science journalist and bioethicist who writes about emerging issues in biotechnology. They have recently launched outlawbio.org, a place for discussion about nontraditional research.
Sept. 13th Event: Delta, Vaccines, and Breakthrough Infections
A visualization of the virus that causes COVID-19.
This virtual event will convene leading scientific and medical experts to address the public's questions and concerns about COVID-19 vaccines, Delta, and breakthrough infections. Audience Q&A will follow the panel discussion. Your questions can be submitted in advance at the registration link.
DATE:
Monday, September 13th, 2021
12:30 p.m. - 1:45 p.m. EDT
REGISTER:

Dr. Amesh Adalja, M.D., FIDSA, Senior Scholar, Johns Hopkins Center for Health Security; Adjunct Assistant Professor, Johns Hopkins Bloomberg School of Public Health; Affiliate of the Johns Hopkins Center for Global Health. His work is focused on emerging infectious disease, pandemic preparedness, and biosecurity.

Dr. Nahid Bhadelia, M.D., MALD, Founding Director, Boston University Center for Emerging Infectious Diseases Policy and Research (CEID); Associate Director, National Emerging Infectious Diseases Laboratories (NEIDL), Boston University; Associate Professor, Section of Infectious Diseases, Boston University School of Medicine. She is an internationally recognized leader in highly communicable and emerging infectious diseases (EIDs) with clinical, field, academic, and policy experience in pandemic preparedness.

Dr. Akiko Iwasaki, Ph.D., Waldemar Von Zedtwitz Professor of Immunobiology and Molecular, Cellular and Developmental Biology and Professor of Epidemiology (Microbial Diseases), Yale School of Medicine; Investigator, Howard Hughes Medical Institute. Her laboratory researches how innate recognition of viral infections lead to the generation of adaptive immunity, and how adaptive immunity mediates protection against subsequent viral challenge.
 Dr. Marion Pepper, Ph.D., Associate Professor, Department of Immunology, University of Washington. Her lab studies how cells of the adaptive immune system, called CD4+ T cells and B cells, form immunological memory by visualizing their differentiation, retention, and function.
Dr. Marion Pepper, Ph.D., Associate Professor, Department of Immunology, University of Washington. Her lab studies how cells of the adaptive immune system, called CD4+ T cells and B cells, form immunological memory by visualizing their differentiation, retention, and function.
This event is the third of a four-part series co-hosted by Leaps.org, the Aspen Institute Science & Society Program, and the Sabin–Aspen Vaccine Science & Policy Group, with generous support from the Gordon and Betty Moore Foundation and the Howard Hughes Medical Institute.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.