Genetically Sequencing Healthy Babies Yielded Surprising Results

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
A company in England has made a test that picks out the compounds from breath that reveal if people have liver disease.
Every year, around two million people worldwide die of liver disease. While some people inherit the disease, it’s most commonly caused by hepatitis, obesity and alcoholism. These underlying conditions kill liver cells, causing scar tissue to form until eventually the liver cannot function properly. Since 1979, deaths due to liver disease have increased by 400 percent.
The sooner the disease is detected, the more effective treatment can be. But once symptoms appear, the liver is already damaged. Around 50 percent of cases are diagnosed only after the disease has reached the final stages, when treatment is largely ineffective.
To address this problem, Owlstone Medical, a biotech company in England, has developed a breath test that can detect liver disease earlier than conventional approaches. Human breath contains volatile organic compounds (VOCs) that change in the first stages of liver disease. Owlstone’s breath test can reliably collect, store and detect VOCs, while picking out the specific compounds that reveal liver disease.
“There’s a need to screen more broadly for people with early-stage liver disease,” says Owlstone’s CEO Billy Boyle. “Equally important is having a test that's non-invasive, cost effective and can be deployed in a primary care setting.”
The standard tool for detection is a biopsy. It is invasive and expensive, making it impractical to use for people who aren't yet symptomatic. Meanwhile, blood tests are less invasive, but they can be inaccurate and can’t discriminate between different stages of the disease.
In the past, breath tests have not been widely used because of the difficulties of reliably collecting and storing breath. But Owlstone’s technology could help change that.
The team is testing patients in the early stages of advanced liver disease, or cirrhosis, to identify and detect these biomarkers. In an initial study, Owlstone’s breathalyzer was able to pick out patients who had early cirrhosis with 83 percent sensitivity.
Boyle’s work is personally motivated. His wife died of colorectal cancer after she was diagnosed with a progressed form of the disease. “That was a big impetus for me to see if this technology could work in early detection,” he says. “As a company, Owlstone is interested in early detection across a range of diseases because we think that's a way to save lives and a way to save costs.”
How it works
In the past, breath tests have not been widely used because of the difficulties of reliably collecting and storing breath. But Owlstone’s technology could help change that.
Study participants breathe into a mouthpiece attached to a breath sampler developed by Owlstone. It has cartridges are designed and optimized to collect gases. The sampler specifically targets VOCs, extracting them from atmospheric gases in breath, to ensure that even low levels of these compounds are captured.
The sampler can store compounds stably before they are assessed through a method called mass spectrometry, in which compounds are converted into charged atoms, before electromagnetic fields filter and identify even the tiniest amounts of charged atoms according to their weight and charge.
The top four compounds in our breath
In an initial study, Owlstone captured VOCs in breath to see which ones could help them tell the difference between people with and without liver disease. They tested the breath of 46 patients with liver disease - most of them in the earlier stages of cirrhosis - and 42 healthy people. Using this data, they were able to create a diagnostic model. Individually, compounds like 2-Pentanone and limonene performed well as markers for liver disease. Owlstone achieved even better performance by examining the levels of the top four compounds together, distinguishing between liver disease cases and controls with 95 percent accuracy.
“It was a good proof of principle since it looks like there are breath biomarkers that can discriminate between diseases,” Boyle says. “That was a bit of a stepping stone for us to say, taking those identified, let’s try and dose with specific concentrations of probes. It's part of building the evidence and steering the clinical trials to get to liver disease sensitivity.”
Sabine Szunerits, a professor of chemistry in Institute of Electronics at the University of Lille, sees the potential of Owlstone’s technology.
“Breath analysis is showing real promise as a clinical diagnostic tool,” says Szunerits, who has no ties with the company. “Owlstone Medical’s technology is extremely effective in collecting small volatile organic biomarkers in the breath. In combination with pattern recognition it can give an answer on liver disease severity. I see it as a very promising way to give patients novel chances to be cured.”
Improving the breath sampling process
Challenges remain. With more than one thousand VOCs found in the breath, it can be difficult to identify markers for liver disease that are consistent across many patients.
Julian Gardner is a professor of electrical engineering at Warwick University who researches electronic sensing devices. “Everyone’s breath has different levels of VOCs and different ones according to gender, diet, age etc,” Gardner says. “It is indeed very challenging to selectively detect the biomarkers in the breath for liver disease.”
So Owlstone is putting chemicals in the body that they know interact differently with patients with liver disease, and then using the breath sampler to measure these specific VOCs. The chemicals they administer are called Exogenous Volatile Organic Compound) probes, or EVOCs.
Most recently, they used limonene as an EVOC probe, testing 29 patients with early cirrhosis and 29 controls. They gave the limonene to subjects at specific doses to measure how its concentrations change in breath. The aim was to try and see what was happening in their livers.
“They are proposing to use drugs to enhance the signal as they are concerned about the sensitivity and selectivity of their method,” Gardner says. “The approach of EVOC probes is probably necessary as you can then eliminate the person-to-person variation that will be considerable in the soup of VOCs in our breath.”
Through these probes, Owlstone could identify patients with liver disease with 83 percent sensitivity. By targeting what they knew was a disease mechanism, they were able to amplify the signal. The company is starting a larger clinical trial, and the plan is to eventually use a panel of EVOC probes to make sure they can see diverging VOCs more clearly.
“I think the approach of using probes to amplify the VOC signal will ultimately increase the specificity of any VOC breath tests, and improve their practical usability,” says Roger Yazbek, who leads the South Australian Breath Analysis Research (SABAR) laboratory in Flinders University. “Whilst the findings are interesting, it still is only a small cohort of patients in one location.”
The future of breath diagnosis
Owlstone wants to partner with pharmaceutical companies looking to learn if their drugs have an effect on liver disease. They’ve also developed a microchip, a miniaturized version of mass spectrometry instruments, that can be used with the breathalyzer. It is less sensitive but will enable faster detection.
Boyle says the company's mission is for their tests to save 100,000 lives. "There are lots of risks and lots of challenges. I think there's an opportunity to really establish breath as a new diagnostic class.”
Doctors worry that fungal pathogens may cause the next pandemic.
Bacterial antibiotic resistance has been a concern in the medical field for several years. Now a new, similar threat is arising: drug-resistant fungal infections. The Centers for Disease Control and Prevention considers antifungal and antimicrobial resistance to be among the world’s greatest public health challenges.
One particular type of fungal infection caused by Candida auris is escalating rapidly throughout the world. And to make matters worse, C. auris is becoming increasingly resistant to current antifungal medications, which means that if you develop a C. auris infection, the drugs your doctor prescribes may not work. “We’re effectively out of medicines,” says Thomas Walsh, founding director of the Center for Innovative Therapeutics and Diagnostics, a translational research center dedicated to solving the antimicrobial resistance problem. Walsh spoke about the challenges at a Demy-Colton Virtual Salon, one in a series of interactive discussions among life science thought leaders.
Although C. auris typically doesn’t sicken healthy people, it afflicts immunocompromised hospital patients and may cause severe infections that can lead to sepsis, a life-threatening condition in which the overwhelmed immune system begins to attack the body’s own organs. Between 30 and 60 percent of patients who contract a C. auris infection die from it, according to the CDC. People who are undergoing stem cell transplants, have catheters or have taken antifungal or antibiotic medicines are at highest risk. “We’re coming to a perfect storm of increasing resistance rates, increasing numbers of immunosuppressed patients worldwide and a bug that is adapting to higher temperatures as the climate changes,” says Prabhavathi Fernandes, chair of the National BioDefense Science Board.
Most Candida species aren’t well-adapted to our body temperatures so they aren’t a threat. C. auris, however, thrives at human body temperatures.
Although medical professionals aren’t concerned at this point about C. auris evolving to affect healthy people, they worry that its presence in hospitals can turn routine surgeries into life-threatening calamities. “It’s coming,” says Fernandes. “It’s just a matter of time.”
An emerging global threat
“Fungi are found in the environment,” explains Fernandes, so Candida spores can easily wind up on people’s skin. In hospitals, they can be transferred from contact with healthcare workers or contaminated surfaces. Most Candida species aren’t well-adapted to our body temperatures so they aren’t a threat. C. auris, however, thrives at human body temperatures. It can enter the body during medical treatments that break the skin—and cause an infection. Overall, fungal infections cost some $48 billion in the U.S. each year. And infection rates are increasing because, in an ironic twist, advanced medical therapies are enabling severely ill patients to live longer and, therefore, be exposed to this pathogen.
The first-ever case of a C. auris infection was reported in Japan in 2009, although an analysis of Candida samples dated the earliest strain to a 1996 sample from South Korea. Since then, five separate varieties – called clades, which are similar to strains among bacteria – developed independently in different geographies: South Asia, East Asia, South Africa, South America and, recently, Iran. So far, C. auris infections have been reported in 35 countries.
In the U.S., the first infection was reported in 2016, and the CDC started tracking it nationally two years later. During that time, 5,654 cases have been reported to the CDC, which only tracks U.S. data.
What’s more notable than the number of cases is their rate of increase. In 2016, new cases increased by 175 percent and, on average, they have approximately doubled every year. From 2016 through 2022, the number of infections jumped from 63 to 2,377, a roughly 37-fold increase.
“This reminds me of what we saw with epidemics from 2013 through 2020… with Ebola, Zika and the COVID-19 pandemic,” says Robin Robinson, CEO of Spriovas and founding director of the Biomedical Advanced Research and Development Authority (BARDA), which is part of the U.S. Department of Health and Human Services. These epidemics started with a hockey stick trajectory, Robinson says—a gradual growth leading to a sharp spike, just like the shape of a hockey stick.
Another challenge is that right now medics don’t have rapid diagnostic tests for fungal infections. Currently, patients are often misdiagnosed because C. auris resembles several other easily treated fungi. Or they are diagnosed long after the infection begins and is harder to treat.
The problem is that existing diagnostics tests can only identify C. auris once it reaches the bloodstream. Yet, because this pathogen infects bodily tissues first, it should be possible to catch it much earlier before it becomes life-threatening. “We have to diagnose it before it reaches the bloodstream,” Walsh says.
The most alarming fact is that some Candida infections no longer respond to standard therapeutics.
“We need to focus on rapid diagnostic tests that do not rely on a positive blood culture,” says John Sperzel, president and CEO of T2 Biosystems, a company specializing in diagnostics solutions. Blood cultures typically take two to three days for the concentration of Candida to become large enough to detect. The company’s novel test detects about 90 percent of Candida species within three to five hours—thanks to its ability to spot minute quantities of the pathogen in blood samples instead of waiting for them to incubate and proliferate.
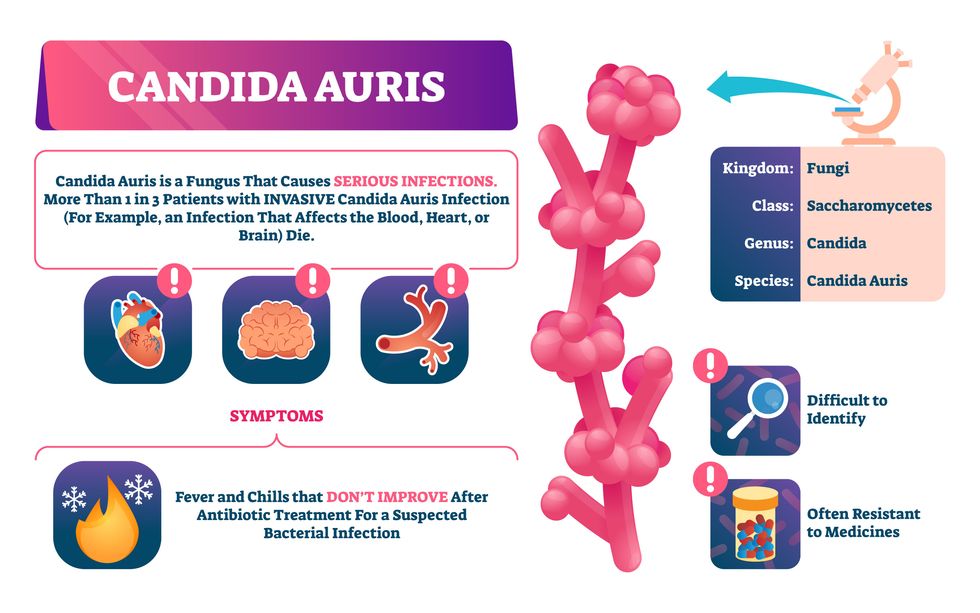
Unlike other Candida species C. auris thrives at human body temperatures
Adobe Stock
Tackling the resistance challenge
The most alarming fact is that some Candida infections no longer respond to standard therapeutics. The number of cases that stopped responding to echinocandin, the first-line therapy for most Candida infections, tripled in 2020, according to a study by the CDC.
Now, each of the first four clades shows varying levels of resistance to all three commonly prescribed classes of antifungal medications, such as azoles, echinocandins, and polyenes. For example, 97 percent of infections from C. auris Clade I are resistant to fluconazole, 54 percent to voriconazole and 30 percent of amphotericin. Nearly half are resistant to multiple antifungal drugs. Even with Clade II fungi, which has the least resistance of all the clades, 11 to 14 percent have become resistant to fluconazole.
Anti-fungal therapies typically target specific chemical compounds present on fungi’s cell membranes, but not on human cells—otherwise the medicine would cause damage to our own tissues. Fluconazole and other azole antifungals target a compound called ergosterol, preventing the fungal cells from replicating. Over the years, however, C. auris evolved to resist it, so existing fungal medications don’t work as well anymore.
A newer class of drugs called echinocandins targets a different part of the fungal cell. “The echinocandins – like caspofungin – inhibit (a part of the fungi) involved in making glucan, which is an essential component of the fungal cell wall and is not found in human cells,” Fernandes says. New antifungal treatments are needed, she adds, but there are only a few magic bullets that will hit just the fungus and not the human cells.
Research to fight infections also has been challenged by a lack of government support. That is changing now that BARDA is requesting proposals to develop novel antifungals. “The scope includes C. auris, as well as antifungals following a radiological/nuclear emergency, says BARDA spokesperson Elleen Kane.
The remaining challenge is the number of patients available to participate in clinical trials. Large numbers are needed, but the available patients are quite sick and often die before trials can be completed. Consequently, few biopharmaceutical companies are developing new treatments for C. auris.
ClinicalTrials.gov reports only two drugs in development for invasive C. auris infections—those than can spread throughout the body rather than localize in one particular area, like throat or vaginal infections: ibrexafungerp by Scynexis, Inc., fosmanogepix, by Pfizer.
Scynexis’ ibrexafungerp appears active against C. auris and other emerging, drug-resistant pathogens. The FDA recently approved it as a therapy for vaginal yeast infections and it is undergoing Phase III clinical trials against invasive candidiasis in an attempt to keep the infection from spreading.
“Ibreafungerp is structurally different from other echinocandins,” Fernandes says, because it targets a different part of the fungus. “We’re lucky it has activity against C. auris.”
Pfizer’s fosmanogepix is in Phase II clinical trials for patients with invasive fungal infections caused by multiple Candida species. Results are showing significantly better survival rates for people taking fosmanogepix.
Although C. auris does pose a serious threat to healthcare worldwide, scientists try to stay optimistic—because they recognized the problem early enough, they might have solutions in place before the perfect storm hits. “There is a bit of hope,” says Robinson. “BARDA has finally been able to fund the development of new antifungal agents and, hopefully, this year we can get several new classes of antifungals into development.”

