Genetically Sequencing Healthy Babies Yielded Surprising Results

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
Jamie Rettinger with his now fiance Amie Purnel-Davis, who helped him through the clinical trial.
Jamie Rettinger was still in his thirties when he first noticed a tiny streak of brown running through the thumbnail of his right hand. It slowly grew wider and the skin underneath began to deteriorate before he went to a local dermatologist in 2013. The doctor thought it was a wart and tried scooping it out, treating the affected area for three years before finally removing the nail bed and sending it off to a pathology lab for analysis.
"I have some bad news for you; what we removed was a five-millimeter melanoma, a cancerous tumor that often spreads," Jamie recalls being told on his return visit. "I'd never heard of cancer coming through a thumbnail," he says. None of his doctors had ever mentioned it either. "I just thought I was being treated for a wart." But nothing was healing and it continued to bleed.
A few months later a surgeon amputated the top half of his thumb. Lymph node biopsy tested negative for spread of the cancer and when the bandages finally came off, Jamie thought his medical issues were resolved.
Melanoma is the deadliest form of skin cancer. About 85,000 people are diagnosed with it each year in the U.S. and more than 8,000 die of the cancer when it spreads to other parts of the body, according to the Centers for Disease Control and Prevention (CDC).
There are two peaks in diagnosis of melanoma; one is in younger women ages 30-40 and often is tied to past use of tanning beds; the second is older men 60+ and is related to outdoor activity from farming to sports. Light-skinned people have a twenty-times greater risk of melanoma than do people with dark skin.
"When I graduated from medical school, in 2005, melanoma was a death sentence" --Diwakar Davar.
Jamie had a follow up PET scan about six months after his surgery. A suspicious spot on his lung led to a biopsy that came back positive for melanoma. The cancer had spread. Treatment with a monoclonal antibody (nivolumab/Opdivo®) didn't prove effective and he was referred to the UPMC Hillman Cancer Center in Pittsburgh, a four-hour drive from his home in western Ohio.
An alternative monoclonal antibody treatment brought on such bad side effects, diarrhea as often as 15 times a day, that it took more than a week of hospitalization to stabilize his condition. The only options left were experimental approaches in clinical trials.
Early research
"When I graduated from medical school, in 2005, melanoma was a death sentence" with a cure rate in the single digits, says Diwakar Davar, 39, an oncologist at UPMC Hillman Cancer Center who specializes in skin cancer. That began to change in 2010 with introduction of the first immunotherapies, monoclonal antibodies, to treat cancer. The antibodies attach to PD-1, a receptor on the surface of T cells of the immune system and on cancer cells. Antibody treatment boosted the melanoma cure rate to about 30 percent. The search was on to understand why some people responded to these drugs and others did not.
At the same time, there was a growing understanding of the role that bacteria in the gut, the gut microbiome, plays in helping to train and maintain the function of the body's various immune cells. Perhaps the bacteria also plays a role in shaping the immune response to cancer therapy.
One clue came from genetically identical mice. Animals ordered from different suppliers sometimes responded differently to the experiments being performed. That difference was traced to different compositions of their gut microbiome; transferring the microbiome from one animal to another in a process known as fecal transplant (FMT) could change their responses to disease or treatment.
When researchers looked at humans, they found that the patients who responded well to immunotherapies had a gut microbiome that looked like healthy normal folks, but patients who didn't respond had missing or reduced strains of bacteria.
Davar and his team knew that FMT had a very successful cure rate in treating the gut dysbiosis of Clostridioides difficile, a persistant intestinal infection, and they wondered if a fecal transplant from a patient who had responded well to cancer immunotherapy treatment might improve the cure rate of patients who did not originally respond to immunotherapies for melanoma.
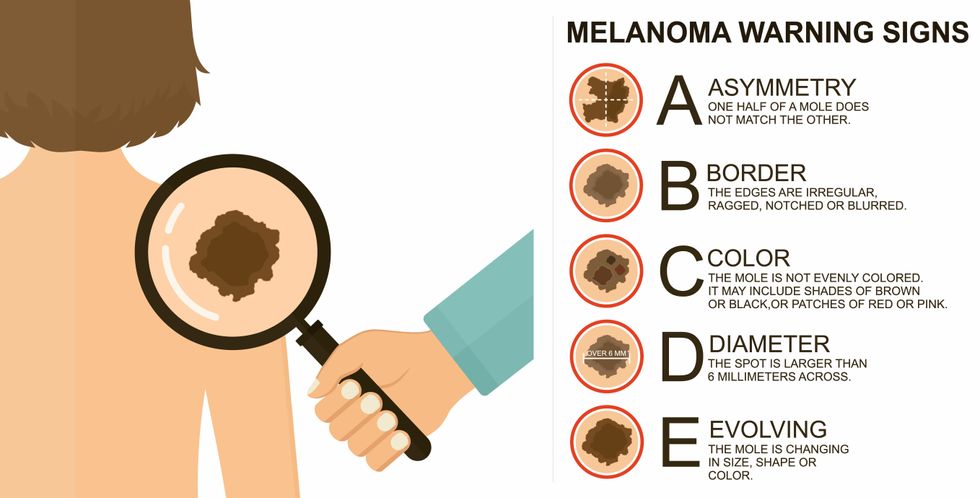
The ABCDE of melanoma detection
Adobe Stock
Clinical trial
"It was pretty weird, I was totally blasted away. Who had thought of this?" Jamie first thought when the hypothesis was explained to him. But Davar's explanation that the procedure might restore some of the beneficial bacterial his gut was lacking, convinced him to try. He quickly signed on in October 2018 to be the first person in the clinical trial.
Fecal donations go through the same safety procedures of screening for and inactivating diseases that are used in processing blood donations to make them safe for transfusion. The procedure itself uses a standard hollow colonoscope designed to screen for colon cancer and remove polyps. The transplant is inserted through the center of the flexible tube.
Most patients are sedated for procedures that use a colonoscope but Jamie doesn't respond to those drugs: "You can't knock me out. I was watching them on the TV going up my own butt. It was kind of unreal at that point," he says. "There were about twelve people in there watching because no one had seen this done before."
A test two weeks after the procedure showed that the FMT had engrafted and the once-missing bacteria were thriving in his gut. More importantly, his body was responding to another monoclonal antibody (pembrolizumab/Keytruda®) and signs of melanoma began to shrink. Every three months he made the four-hour drive from home to Pittsburgh for six rounds of treatment with the antibody drug.
"We were very, very lucky that the first patient had a great response," says Davar. "It allowed us to believe that even though we failed with the next six, we were on the right track. We just needed to tweak the [fecal] cocktail a little better" and enroll patients in the study who had less aggressive tumor growth and were likely to live long enough to complete the extensive rounds of therapy. Six of 15 patients responded positively in the pilot clinical trial that was published in the journal Science.
Davar believes they are beginning to understand the biological mechanisms of why some patients initially do not respond to immunotherapy but later can with a FMT. It is tied to the background level of inflammation produced by the interaction between the microbiome and the immune system. That paper is not yet published.
Surviving cancer
It has been almost a year since the last in his series of cancer treatments and Jamie has no measurable disease. He is cautiously optimistic that his cancer is not simply in remission but is gone for good. "I'm still scared every time I get my scans, because you don't know whether it is going to come back or not. And to realize that it is something that is totally out of my control."
"It was hard for me to regain trust" after being misdiagnosed and mistreated by several doctors he says. But his experience at Hillman helped to restore that trust "because they were interested in me, not just fixing the problem."
He is grateful for the support provided by family and friends over the last eight years. After a pause and a sigh, the ruggedly built 47-year-old says, "If everyone else was dead in my family, I probably wouldn't have been able to do it."
"I never hesitated to ask a question and I never hesitated to get a second opinion." But Jamie acknowledges the experience has made him more aware of the need for regular preventive medical care and a primary care physician. That person might have caught his melanoma at an earlier stage when it was easier to treat.
Davar continues to work on clinical studies to optimize this treatment approach. Perhaps down the road, screening the microbiome will be standard for melanoma and other cancers prior to using immunotherapies, and the FMT will be as simple as swallowing a handful of freeze-dried capsules off the shelf rather than through a colonoscopy. Earlier this year, the Food and Drug Administration approved the first oral fecal microbiota product for C. difficile, hopefully paving the way for more.
An older version of this hit article was first published on May 18, 2021
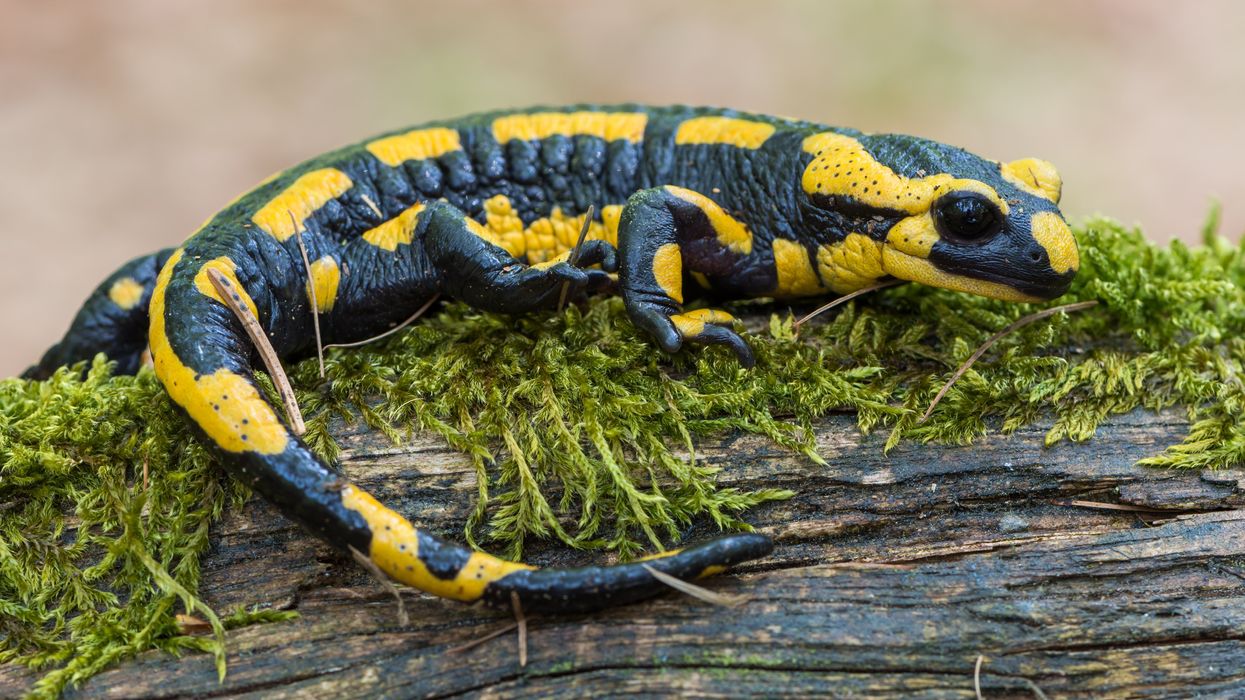
All organisms can repair damaged tissue, but none do it better than salamanders and newts. A surprising area of science could tell us how they manage this feat - and perhaps even help us develop a similar ability.
All organisms have the capacity to repair or regenerate tissue damage. None can do it better than salamanders or newts, which can regenerate an entire severed limb.
That feat has amazed and delighted man from the dawn of time and led to endless attempts to understand how it happens – and whether we can control it for our own purposes. An exciting new clue toward that understanding has come from a surprising source: research on the decline of cells, called cellular senescence.
Senescence is the last stage in the life of a cell. Whereas some cells simply break up or wither and die off, others transition into a zombie-like state where they can no longer divide. In this liminal phase, the cell still pumps out many different molecules that can affect its neighbors and cause low grade inflammation. Senescence is associated with many of the declining biological functions that characterize aging, such as inflammation and genomic instability.
Oddly enough, newts are one of the few species that do not accumulate senescent cells as they age, according to research over several years by Maximina Yun. A research group leader at the Center for Regenerative Therapies Dresden and the Max Planck Institute of Molecular and Cell Biology and Genetics, in Dresden, Germany, Yun discovered that senescent cells were induced at some stages of regeneration of the salamander limb, “and then, as the regeneration progresses, they disappeared, they were eliminated by the immune system,” she says. “They were present at particular times and then they disappeared.”
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states.
Previous research on senescence in aging had suggested, logically enough, that applying those cells to the stump of a newly severed salamander limb would slow or even stop its regeneration. But Yun stood that idea on its head. She theorized that senescent cells might also play a role in newt limb regeneration, and she tested it by both adding and removing senescent cells from her animals. It turned out she was right, as the newt limbs grew back faster than normal when more senescent cells were included.
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states, which could then be turned into progenitors, a cell type in between stem cells and specialized cells, needed to regrow the muscle tissue of the missing limb. “We think that this ability to dedifferentiate is intrinsically a big part of why salamanders can regenerate all these very complex structures, which other organisms cannot,” she explains.
Yun sees regeneration as a two part problem. First, the cells must be able to sense that their neighbors from the lost limb are not there anymore. Second, they need to be able to produce the intermediary progenitors for regeneration, , to form what is missing. “Molecularly, that must be encoded like a 3D map,” she says, otherwise the new tissue might grow back as a blob, or liver, or fin instead of a limb.
Wound healing
Another recent study, this time at the Mayo Clinic, provides evidence supporting the role of senescent cells in regeneration. Looking closely at molecules that send information between cells in the wound of a mouse, the researchers found that senescent cells appeared near the start of the healing process and then disappeared as healing progressed. In contrast, persistent senescent cells were the hallmark of a chronic wound that did not heal properly. The function and significance of senescence cells depended on both the timing and the context of their environment.
The paper suggests that senescent cells are not all the same. That has become clearer as researchers have been able to identify protein markers on the surface of some senescent cells. The patterns of these proteins differ for some senescent cells compared to others. In biology, such physical differences suggest functional differences, so it is becoming increasingly likely there are subsets of senescent cells with differing functions that have not yet been identified.
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes.
Scientists initially thought that senescent cells couldn’t play a role in regeneration because they could no longer reproduce, says Anthony Atala, a practicing surgeon and bioengineer who leads the Wake Forest Institute for Regenerative Medicine in North Carolina. But Yun’s study points in the other direction. “What this paper shows clearly is that these cells have the potential to be involved in tissue regeneration [in newts]. The question becomes, will these cells be able to do the same in humans.”
As our knowledge of senescent cells increases, Atala thinks we need to embrace a new analogy to help understand them: humans in retirement. They “have acquired a lot of wisdom throughout their whole life and they can help younger people and mentor them to grow to their full potential. We're seeing the same thing with these cells,” he says. They are no longer putting energy into their own reproduction, but the signaling molecules they secrete “can help other cells around them to regenerate.”
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes. If so, it seems that our genes are unable to express this ability, perhaps as part of a tradeoff in acquiring other traits. It is a fertile area of research.
Dedifferentiation is likely to become an important process in the field of regenerative medicine. One extreme example: a lab has been able to turn back the clock and reprogram adult male skin cells into female eggs, a potential milestone in reproductive health. It will be more difficult to control just how far back one wishes to go in the cell's dedifferentiation – part way or all the way back into a stem cell – and then direct it down a different developmental pathway. Yun is optimistic we can learn these tricks from newts.
Senolytics
A growing field of research is using drugs called senolytics to remove senescent cells and slow or even reverse disease of aging.
“Senolytics are great, but senolytics target different types of senescence,” Yun says. “If senescent cells have positive effects in the context of regeneration, of wound healing, then maybe at the beginning of the regeneration process, you may not want to take them out for a little while.”
“If you look at pretty much all biological systems, too little or too much of something can be bad, you have to be in that central zone” and at the proper time, says Atala. “That's true for proteins, sugars, and the drugs that you take. I think the same thing is true for these cells. Why would they be different?”
Our growing understanding that senescence is not a single thing but a variety of things likely means that effective senolytic drugs will not resemble a single sledge hammer but more a carefully manipulated scalpel where some types of senescent cells are removed while others are added. Combinations and timing could be crucial, meaning the difference between regenerating healthy tissue, a scar, or worse.

