Genetically Sequencing Healthy Babies Yielded Surprising Results

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
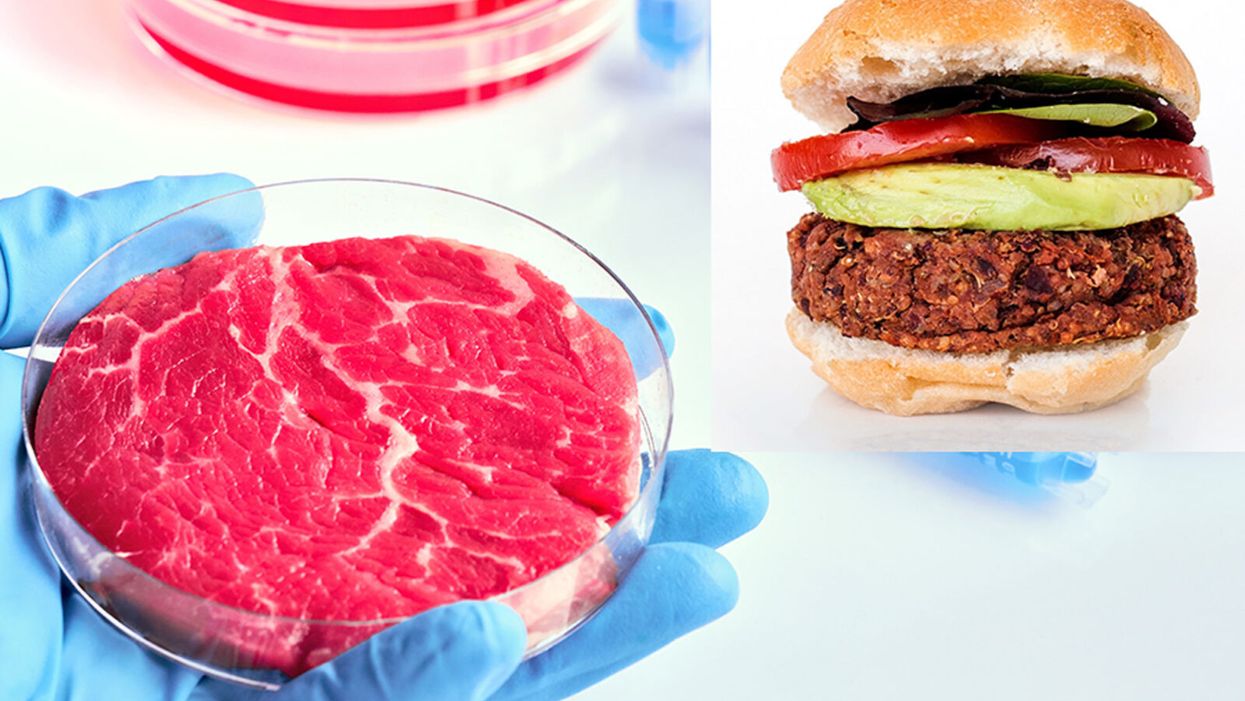
While lab-grown meat is not yet ready to crisis-proof the food supply chain, it offers key benefits that could make it an important solution beyond this pandemic.
The coronavirus pandemic exposed significant weaknesses in the country's food supply chain. Grocery store meat counters were bare. Transportation interruptions influenced supply. Finding beef, poultry, and pork at the store has been, in some places, as challenging as finding toilet paper.
In traditional agriculture models, it takes at least three months to raise chicken, six to nine months for pigs, and 18 months for cattle.
It wasn't a lack of supply -- millions of animals were in the pipeline.
"There's certainly enough food out there, but it can't get anywhere because of the way our system is set up," said Amy Rowat, an associate professor of integrative biology and physiology at UCLA. "Having a more self-contained, self-sufficient way to produce meat could make the supply chain more robust."
Cultured meat could be one way of making the meat supply chain more resilient despite disruptions due to pandemics such as COVID-19. But is the country ready to embrace lab-grown food?
According to a Good Food Institute study, GenZ is almost twice as likely to embrace meat alternatives for reasons related to social and environmental awareness, even prior to the pandemic. That's because this group wants food choices that reflect their values around food justice, equity, and animal welfare.
Largely, the interest in protein alternatives has been plant-based foods. However, factors directly related to COVID-19 may accelerate consumer interest in the scaling up of cell-grown products, according to Liz Specht, the associate director of science and technology at The Good Food Institute. The latter is a nonprofit organization that supports scientists, investors, and entrepreneurs working to develop food alternatives to conventional animal products.
While lab-grown food isn't ready yet to definitively crisis-proof the food supply chain, experts say it offers promise.
Matching Supply and Demand
Companies developing cell-grown meat claim it can take as few as two months to develop a cell into an edible product, according to Anthony Chow, CFA at Agronomics Limited, an investment company focused on meat alternatives. Tissue is taken from an animal and placed in a culture that contains nutrients and proteins the cells need to grow and expand. He cites a Good Food Institute report that claims a 2.5-millimeter sample can grow three and a half tons of meat in 40 days, allowing for exponential growth when needed.
In traditional agriculture models, it takes at least three months to raise chicken, six to nine months for pigs, and 18 months for cattle. To keep enough maturing animals in the pipeline, farms must plan the number of animals to raise months -- even years -- in advance. Lab-grown meat advocates say that because cultured meat supplies can be flexible, it theoretically allows for scaling up or down in significantly less time.
"Supply and demand has drastically changed in some way around the world and cultivated meat processing would be able to adapt much quicker than conventional farming," Chow said.
Scaling Up
Lab-grown meat may provide an eventual solution, but not in the immediate future, said Paul Mozdziak, a professor of physiology at North Carolina State University who researches animal cell culture techniques, transgenic animal production, and muscle biology.
"The challenge is in culture media," he said. "It's going to take some innovation to get the cells to grow at quantities that are going to be similar to what you can get from an animal. These are questions that everybody in the space is working on."
Chow says some of the most advanced cultured meat companies, such as BlueNal, anticipate introducing products to the market midway through next year. However, he thinks COVID-19 has slowed the process. Once introduced, they will be at a premium price, most likely available at restaurants before they hit grocery store shelves.
"I think in five years' time it will be in a different place," he said. "I don't think that this will have relevance for this pandemic, but certainly beyond that."
"Plant-based meats may be perceived as 'alternatives' to meat, whereas lab-grown meat is producing the same meat, just in a much more efficient manner, without the environmental implications."
Of course, all the technological solutions in the world won't solve the problem unless people are open-minded about embracing them. At least for now, a lab-grown burger or bluefin tuna might still be too strange for many people, especially in the U.S.
For instance, a 2019 article published by "Frontiers in Sustainable Food Systems" reflects results from a study of 3,030 consumers showing that 29 percent of U.S. customers, 59 percent of Chinese consumers, and 56 percent of Indian consumers were either 'very' or 'extremely likely' to try cultivated meat.
"Lab-grown meat is genuine meat, at the cellular level, and therefore will match conventional meat with regard to its nutritional content and overall sensory experience. It could be argued that plant-based meat will never be able to achieve this," says Laura Turner, who works with Chow at Agronomics Limited. "Plant-based meats may be perceived as 'alternatives' to meat, whereas lab-grown meat is producing the same meat, just in a much more efficient manner, without the environmental implications."
A Solution Beyond This Pandemic
The coronavirus has done more than raise awareness of the fragility of food supply chains. It has also been a wakeup call for consumers and policy makers that it is time to radically rethink our meat, Specht says. Those factors have elevated the profile of lab-grown meat.
"I think the economy is getting a little bit more steam and if I was an investor, I would be getting excited about it," adds Mozdziak.
Beyond crises, Mozdziak explains that as affluence continues to increase globally, meat consumption increases exponentially. Yet farm animals can only grow so quickly and traditional farming won't be able to keep up.
"Even Tyson is saying that by 2050, there's not going to be enough capacity in the animal meat space to meet demand," he notes. "If we don't look at some innovative technologies, how are we going to overcome that?"
A COVID-19 moonshot program challenged chemists around the world to submit ideas for how best to design a drug to target the virus.
By mid-March, Alpha Lee was growing restless. A pioneer of AI-driven drug discovery, Lee leads a team of researchers at the University of Cambridge, but his lab had been closed amidst the government-initiated lockdowns spreading inexorably across Europe.
If the Moonshot proves successful, they hope it could serve as a future benchmark for finding new medicines for chronic diseases.
Having spoken to his collaborators across the globe – many of whom were seeing their own experiments and research projects postponed indefinitely due to the pandemic – he noticed a similar sense of frustration and helplessness in the face of COVID-19.
While there was talk of finding a novel treatment for the virus, Lee was well aware the process was likely to be long and laborious. Traditional methods of drug discovery risked suffering the same fate as the efforts to find a cure for SARS in the early 2000, which took years and were ultimately abandoned long before a drug ever reached the market.
To avoid such an outcome, Lee was convinced that global collaboration was required. Together with a collection of scientists in the UK, US and Israel, he launched the 'COVID Moonshot' – a project which encouraged chemists worldwide to share their ideas for potential drug designs. If the Moonshot proves successful, they hope it could serve as a future benchmark for finding new medicines for chronic diseases.
Solving a Complex Jigsaw
In February, ShanghaiTech University published the first detailed snapshots of the SARS-CoV-2 coronavirus's proteins using a technique called X-ray crystallography. In particular, they revealed a high-resolution profile of the virus's main protease – the part of its structure that enables it to replicate inside a host – and the main drug target. The images were tantalizing.
"We could see all the tiny pieces sitting in the structure like pieces of a jigsaw," said Lee. "All we needed was for someone to come up with the best idea of joining these pieces together with a drug. Then you'd be left with a strong molecule which sits in the protease, and stops it from working, killing the virus in the process."
Normally, ideas for how best to design such a drug would be kept as carefully guarded secrets within individual labs and companies due to their potential value. But as a result, the steady process of trial and error to reach an optimum design can take years to come to fruition.
However, given the scale of the global emergency, Lee felt that the scientific community would be open to collective brainstorming on a mass scale. "Big Pharma usually wouldn't necessarily do this, but time is of the essence here," he said. "It was a case of, 'Let's just rethink every drug discovery stage to see -- ok, how can we go as fast as we can?'"
On March 13, he launched the COVID moonshot, calling for chemists around the globe to come up with the most creative ideas they could think of, on their laptops at home. No design was too weird or wacky to be considered, and crucially nothing would be patented. The entire project would be done on a not-for-profit basis, meaning that any drug that makes it to market will have been created simply for the good of humanity.
It caught fire: Within just two weeks, more than 2,300 potential drug designs had been submitted. By the middle of July, over 10,000 had been received from scientists around the globe.
The Road Toward Clinical Trials
With so many designs to choose from, the team has been attempting to whittle them down to a shortlist of the most promising. Computational drug discovery experts at Diamond and the Weizmann Institute of Science in Rehovot, Israel, have enabled the Moonshot team to develop algorithms for predicting how quick and easy each design would be to make, and to predict how well each proposed drug might bind to the virus in real life.
The latter is an approach known as computational covalent docking and has previously been used in cancer research. "This was becoming more popular even before COVID-19, with several covalent drugs approved by the FDA in recent years," said Nir London, professor of organic chemistry at the Weizmann Institute, and one of the Moonshot team members. "However, all of these were for oncology. A covalent drug against SARS-CoV-2 will certainly highlight covalent drug-discovery as a viable option."
Through this approach, the team have selected 850 compounds to date, which they have manufactured and tested in various preclinical trials already. Fifty of these compounds - which appear to be especially promising when it comes to killing the virus in a test tube – are now being optimized further.
Lee is hoping that at least one of these potential drugs will be shown to be effective in curing animals of COVID-19 within the next six months, a step that would allow the Moonshot team to reach out to potential pharmaceutical partners to test their compounds in humans.
Future Implications
If the project does succeed, some believe it could open the door to scientific crowdsourcing as a future means of generating novel medicine ideas for other diseases. Frank von Delft, professor of protein science and structural biology at the University of Oxford's Nuffield Department of Medicine, described it as a new form of 'citizen science.'
"There's a vast resource of expertise and imagination that is simply dying to be tapped into," he said.
Others are slightly more skeptical, pointing out that the uniqueness of the current crisis has meant that many scientists were willing to contribute ideas without expecting any future compensation in return. This meant that it was easy to circumvent the traditional hurdles that prevent large-scale global collaborations from happening – namely how to decide who will profit from the final product and who will hold the intellectual property (IP) rights.
"I think it is too early to judge if this is a viable model for future drug discovery," says London. "I am not sure that without the existential threat we would have seen so many contributions, and so many people and institutions willing to waive compensation and future royalties. Many scientists found themselves at home, frustrated that they don't have a way to contribute to the fight against COVID-19, and this project gave them an opportunity. Plus many can get behind the fact that this project has no associated IP and no one will get rich off of this effort. This breaks down a lot of the typical barriers and red-tape for wider collaboration."
"If a drug would sprout from one of these crowdsourced ideas, it would serve as a very powerful argument to consider this mode of drug discovery further in the future."
However the Moonshot team believes that if they can succeed, it will at the very least send a strong statement to policy makers and the scientific community that greater efforts should be made to make such large-scale collaborations more feasible.
"All across the scientific world, we've seen unprecedented adoption of open-science, collaboration and collegiality during this crisis, perhaps recognizing that only a coordinated global effort could address this global challenge," says London. "If a drug would sprout from one of these crowdsourced ideas, it would serve as a very powerful argument to consider this mode of drug discovery further in the future."
[An earlier version of this article was published on June 8th, 2020 as part of a standalone magazine called GOOD10: The Pandemic Issue. Produced as a partnership among LeapsMag, The Aspen Institute, and GOOD, the magazine is available for free online.]