Genetically Sequencing Healthy Babies Yielded Surprising Results
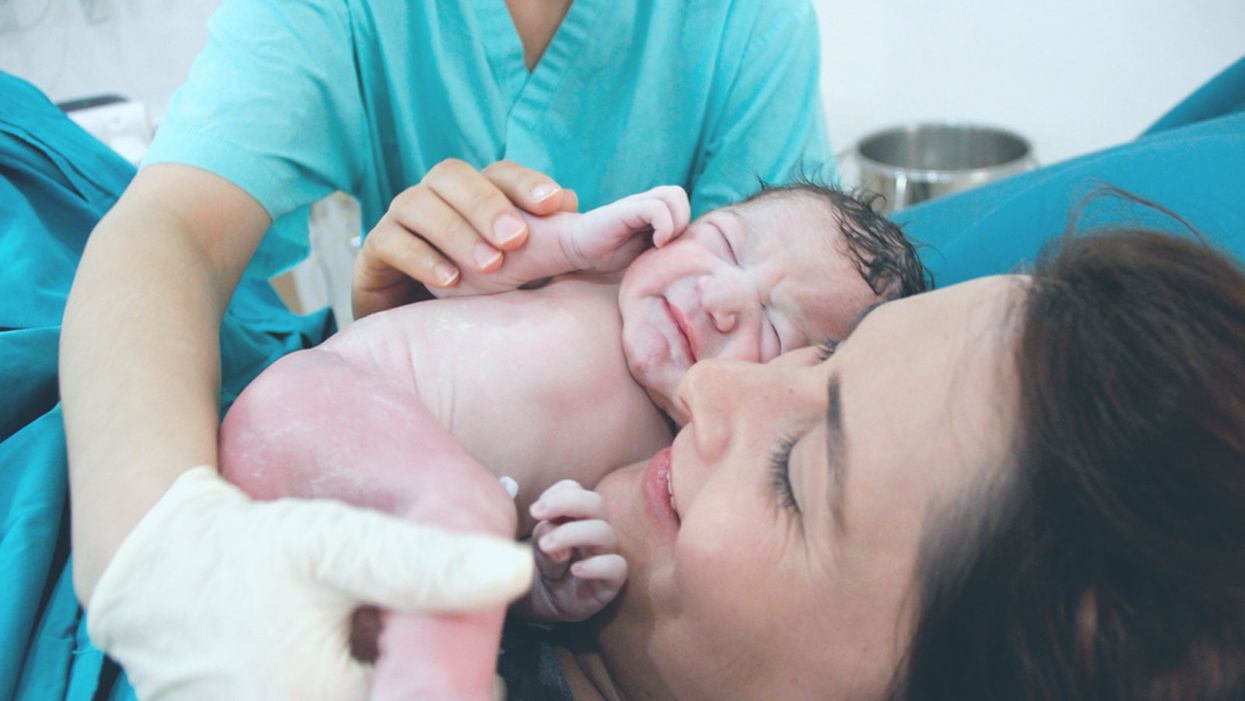

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
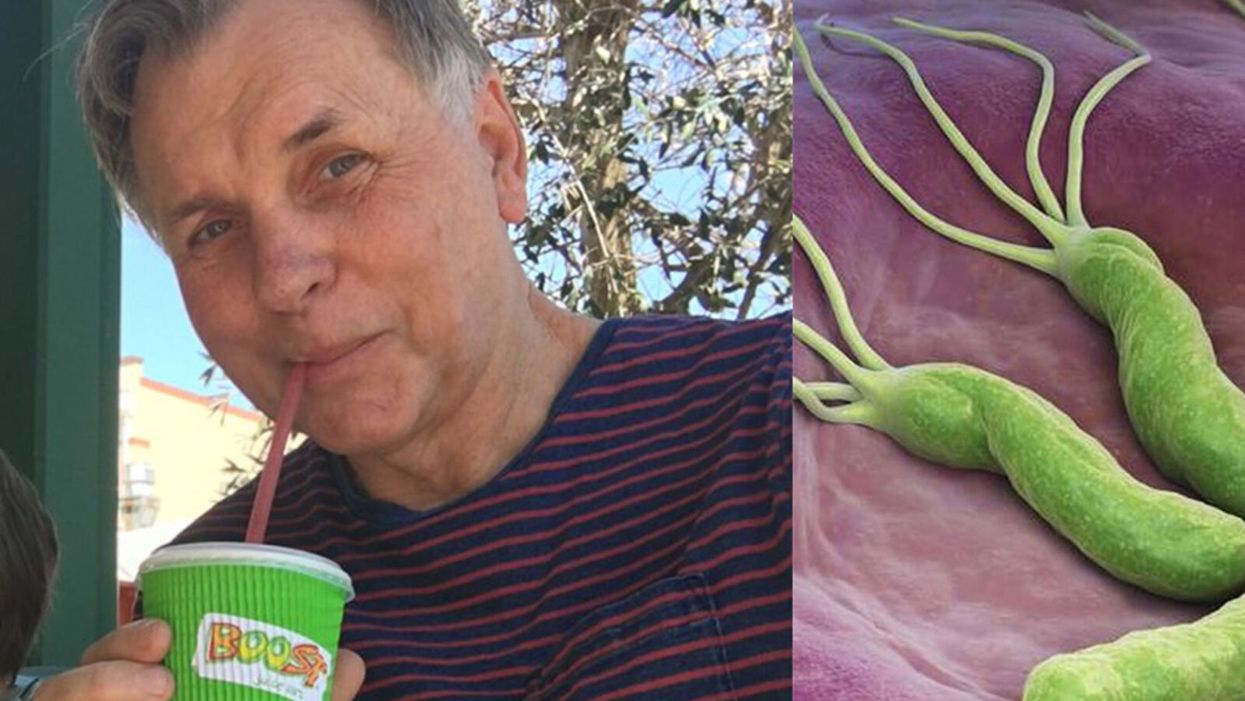
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
Dr. Barry Marshall (along with a collaborator) discovered that the bacteria H. Pylori, pictured, caused stomach ulcers.
[Editor's Note: Welcome to Leaps of the Past, a new monthly column that spotlights the fascinating backstory behind a medical or scientific breakthrough from history.]
------
Until about 40 years ago, ulcers were a mysterious – and sometimes deadly – ailment. Found in a person's stomach lining or intestine, ulcers are small sores that cause a variety of painful symptoms, such as vomiting, a burning or aching sensation, internal bleeding and stomach obstruction. Patients with ulcers suffered for years without a cure and sometimes even needed their stomachs completely removed to rid them from pain.
"To gastroenterologists, the concept of a germ causing ulcers was like saying the Earth is flat."
In the early 1980s, the majority of scientists thought that ulcers were caused by stress or poor diet. But a handful of scientists had a different theory: They believed that ulcers were caused by a corkscrew-shaped bacterium called Helicobacter pylori, or H. pylori for short. Robin Warren, a pathologist, and Barry Marshall, an internist, were the two pioneers of this theory, and the two teamed up to study H. pylori at the Royal Perth Hospital in 1981.
The pair started off by trying to culture the bacteria in the stomachs of patients with gastritis, an inflammation of the stomach lining and a precursor to developing an ulcer. Initially, the microbiologists involved in their clinical trial found no trace of the bacteria from patient samples – but after a few weeks, the microbiologists discovered that their lab techs had been throwing away the cultures before H. pylori could grow. "After that, we let the cultures grow longer and found 13 patients with duodenal ulcer," said Marshall in a later interview. "All of them had the bacteria."
Marshall and Warren also cultured H. pylori in the stomachs of patients with stomach cancer. They observed that "everybody with stomach cancer developed it on a background of gastritis. Whenever we found a person without Helicobacter, we couldn't find gastritis either." Marshall and Warren were convinced that H. pylori not only caused gastritis and peptic ulcers, but stomach cancer as well.
But when the team presented their findings at an annual meeting of the Royal Australasian College of Physicians in Perth, they were mostly met with skepticism. "To gastroenterologists, the concept of a germ causing ulcers was like saying the Earth is flat," Marshall said. "The idea was too weird."
Warren started treating his gastritis patients with antibiotics with great success – but other internists remained doubtful, continuing to treat their patients with antacids instead. Making matters more complicated, neither Warren nor Marshall could readily test their theory, since the pair only had lab mice at their disposal and H. pylori infects only humans and non-human primates, such as rhesus monkeys.
So Marshall took an unconventional approach. First, he underwent two tests to get a baseline reading of his stomach, which showed no presence of H. pylori. Then, Marshall took some H. pylori bacteria from a petri dish, mixed it with beef extract to create a broth, and gulped it down. If his theory was correct, a second gastric biopsy would show that his stomach was overrun with H. pylori bacteria, and a second endoscopy would show a painfully inflamed stomach – gastritis.
Less than a week later, Marshall started feeling sick. "I expected to develop an asymptomatic infection," he later said in an interview published in the Canadian Journal of Gastroenterology. "… [but] after five days, I started to have bloating and fullness after the evening meal, and my appetite decreased. My breath was bad and I vomited clear watery liquid, without acid, each morning."
At his wife's urging, Marshall started on a regimen of antibiotics to kill off the burgeoning bacteria, so a follow-up biopsy showed no signs of H. pylori. A follow-up endoscopy, however, showed "severe active gastritis" along with epithelial damage. This was the smoking gun other clinicians needed to believe that H. pylori caused gastritis and stomach cancer. When they began to treat their gastritis patients with antibiotics, the rate of peptic ulcers in the Australian population diminished by 70 percent.
Today, antibiotics are the standard of care for anyone afflicted with gastritis.
In 2005, Marshall and Warren were awarded the Nobel Prize in Physiology or Medicine for their discovery of H. Pylori and its role in developing gastritis and peptic ulcers. "Thanks to the pioneering discovery by Marshall and Warren, peptic ulcer disease is no longer a chronic, frequently disabling condition, but a disease that can be cured by a short regimen of antibiotics and acid secretion inhibitors," the Nobel Prize Committee said.
Today, antibiotics are the standard of care for anyone afflicted with gastritis – and stomach cancer has been significantly reduced in the Western world.
Would a Broad-Spectrum Antiviral Drug Stop the Pandemic?
An illustration of the novel coronavirus infecting human tissue.
The refocusing of medical research to COVID-19 is unprecedented in human history. Seven months ago, we barely were aware that the virus existed, and now a torrent of new information greets us each day online.
There are many unanswered questions about COVID-19, but perhaps the most fascinating is whether we even need to directly go after the virus itself.
Clinicaltrials.gov, the most commonly used registry for worldwide medical research, listed 1358 clinical trials on the disease, including using scores of different potential drugs and multiple combinations, when I first wrote this sentence. The following day that number of trials had increased to 1409. Laboratory work to prepare for trials presents an even broader and untabulated scope of activity.
Most trials will fail or not be as good as what has been discovered in the interim, but the hope is that a handful of them will yield vaccines for prevention and treatments to attenuate and ultimately cure the deadly infection.
The first impulse is to grab whatever drugs are on the shelf and see if any work against the new foe. We know their safety profiles and they have passed some regulatory hurdles. Remdesivir is the first to register some success against SARS-CoV-2, the virus behind the disease. The FDA has granted it expedited-use status, pending presentation of data that may lead to full approval of the drug.
Most observers see it as a treatment that might help, but not one that by itself is likely to break the back of the pandemic. Part of that is because it is delivered though IV infusion, which requires hospitalization, and as with most antiviral drugs, appears to be most beneficial when started early in disease. "The most effective products are going to be that ones that are developed by actually understanding more about this coronavirus," says Margaret "Peggy" Hamburg, who once led the New York City public health department and later the U.S. Food and Drug Administration.
Combination therapy that uses different drugs to hit a virus at different places in its life cycle have proven to work best in treating HIV and hepatitis C, and likely will be needed with this virus as well. Most viruses are simply too facile at evolving resistance to a single drug, and so require multiple hits to keep them down.
Laboratory work suggests that other drugs, both off-the-shelf and in development, particularly those to treat HIV and hepatitis, might also be of some benefit against SARS-CoV-2. But the number of possible drug combinations is mind-bogglingly large and the capacity to test them all right now is limited.
Broad-Spectrum Antivirals
Viruses are simple quasi-life forms. Effective treatments are more likely to be specific to a given virus, or at best its close relatives. That is unlike bacteria, where broad-spectrum antibiotics often can be used against common elements like the bacterial cell wall, or can disrupt quorum sensing signals that bacteria use to function as biofilms.
More than a decade ago, virologist Benhur Lee's lab at UCLA (now at Mt. Sinai in New York City) stumbled upon a broad-spectrum antiviral approach that seemed to work against all enveloped viruses they tested. The list ranged from the common flu to HIV to Ebola.
Other researchers grabbed this lead to develop a compound that worked quite well in cell cultures, but when they tried it in animals, a frustrating snag emerged; the compound needed to be activated by light. As the greatest medical need is to counter viruses deep inside the body, the research was put on the shelf. So Lee was surprised to learn recently that a company has inquired about rights to develop the compound not as a treatment but as a possible disinfectant. The tale illustrates both the unanticipated difficulties of drug development and that one never knows how knowledge ultimately might be put to use.
Remdesivir is a failed drug for Ebola that has found new life with SARS-CoV-2. It targets polymerase, an enzyme that the virus produces to use host cell machinery to replicate itself, and since the genetic sequence of polymerase is very similar among all of the different coronaviruses, scientists hope that the drug might be useful against known members of the family and others that might emerge in the future.
But nature isn't always that simple. Viral RNA is not a two-dimensional assemblage of genes in a flat line on a table; rather it is a three-dimensional matrix of twists and turns where a single atom change within the polymerase gene or another gene close by might change the orientation of the RNA or a molecular arm within it and block a drug from accessing the targeted binding site on the virus. One drug might need to bind to a large flat surface, while another might be able to slip a dagger-like molecular arm through a space in the matrix to reach its binding target.
That is why a broad-spectrum antiviral is so hard to develop, and why researchers continue to work on a wide variety of compounds that target polymerase as a binding site.
Additionally, it has taken us decades to begin to recognize the unintended consequences of broad-spectrum rather than narrowly targeted antibiotics on the gut microbiome and our overall health. Will a similar issue potentially arise in using a broad-spectrum antiviral?
"Off-target side effects are always of concern with drugs, and antivirals are no exception," says Yale University microbiologist Ben Chen. He believes that "most" bacteriophages, the viruses that infect bacteria and likely help to maintain stability in the gut microbial ecosystem, will shrug off such a drug. However, a few families of phages share polymerases that are similar to those found in coronaviruses. While the immediate need for treatment is great, we will have to keep a sharp eye out for unanticipated activity in the body's ecosystem from new drugs.
Is an Antiviral Needed?
There are many unanswered questions about COVID-19, but perhaps the most fascinating is whether we even need to directly go after the virus itself. Mounting evidence indicates that up to half the people who contract the infection don't seem to experience significant symptoms and their immune system seems to clear the virus.
The most severe cases of COVID-19 appear to result from an overactive immune response that damages surrounding tissue. Perhaps downregulating that response will be sufficient to reduce the disease burden. Several studies are underway using approved antibodies that modulate an overly active immune response.
One of the most surprising findings to date involves the monoclonal antibody leronlimab. It was originally developed to treat HIV infection and works modestly well there, but other drugs are better and its future likely will be mainly to treat patients who have developed resistance to those other drugs.
The response has been amazingly different in patients in the U.S. with COVID-19 who were given emergency access to leronlimab – two injections a week apart, though the company believes that four might be better. The immune response and inflammatory cytokines declined significantly, T cell counts were maintained, and surprisingly the amount of virus in the blood declined too. Data from the first ten patients is available in a preprint while the paper undergoes peer review for publication. Data from an additional fifty patients will be added.
"We got lucky and hit the bulls' eye from a mile away," says Jay Lalezari, the chief science officer of Cytodyn, the company behind leronlimab. Dr. Jay, as he is widely known in San Francisco, built an adoring fan base running many of the early-phase drug studies for treating HIV. While touting leronlimab, Lalezari suspects it might best be used as part of a combination therapy.
The small, under-capitalized firm is struggling for attention in the vast pool of therapies proposed to treat COVID-19. It faces the added challenge of gaining acceptance because it is based on a different approach and mechanism of action, which involves a signaling molecule important to immune cell migration, than what most researchers and the FDA anticipate as being relevant to counter SARS-CoV-2.
Common Issues
All of the therapeutics under development will face some common sets of issues. One is the pressure to have results yesterday, because people are dying. The rush to disseminate information "make me worry that certain things will become entrenched as truth, even in the scientific community, without the actual scientific documentation that ordinarily scientists would demand," says Hamburg.
"It is becoming increasingly clear that the biggest problem for drug and vaccine makers is not which therapeutics or vaccine platform to pursue."
Lack of standardization in assays and laboratory operations makes it difficult to compare results between labs studying SARS-CoV-2. In the long run, this will slow down the iterative process of research that builds upon what has gone before. And the shut down of supply chains, from chemicals to cell lines to animals to air shipment, has the potential to further hobble research.
Almost all researchers consult with the FDA in putting together their clinical trials. But the agency is overwhelmed with the surge of activity in the field, and is even less capable of handling novel approaches that fall outside of its standard guidance.
"It is becoming increasingly clear that the biggest problem for drug and vaccine makers is not which therapeutics or vaccine platform to pursue. It is that conventional clinical development paths are far too lengthy and cumbersome to address the current public health threat," John Hodgson wrote in Nature Biotechnology.
Another complicating factor with this virus is the broad range of organ and tissue types it can infect. That has implications for potential therapies, which often vary in their ability to enter different tissues. At a minimum, it complicates the drug development process.
Remdesivir has become the de facto standard of care. Ideally, clinical trials are conducted using the existing standard of care rather than a placebo as the control group. But shortages of the drug make that difficult and further inhibit learning what is the best treatment regimen for regular clinical care.
"Understandably, we all really want to respond to COVID-19 in a much, much more accelerated fashion," says Hamburg. But ultimately that depends upon "the reality of understanding the nature of the disease. And that is going to take a bit more time than we might like or wish."
[This article was originally published on June 8th, 2020 as part of a standalone magazine called GOOD10: The Pandemic Issue. Produced as a partnership among LeapsMag, The Aspen Institute, and GOOD, the magazine is available for free online.]