Genetically Sequencing Healthy Babies Yielded Surprising Results
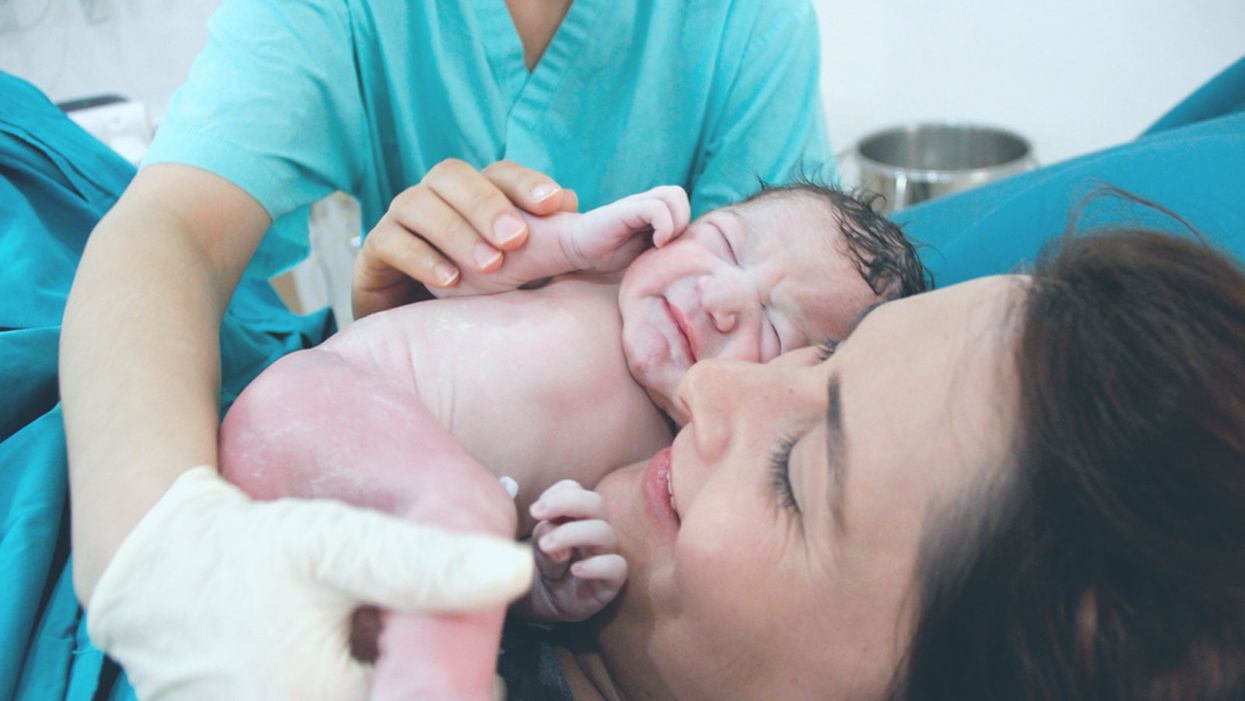

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
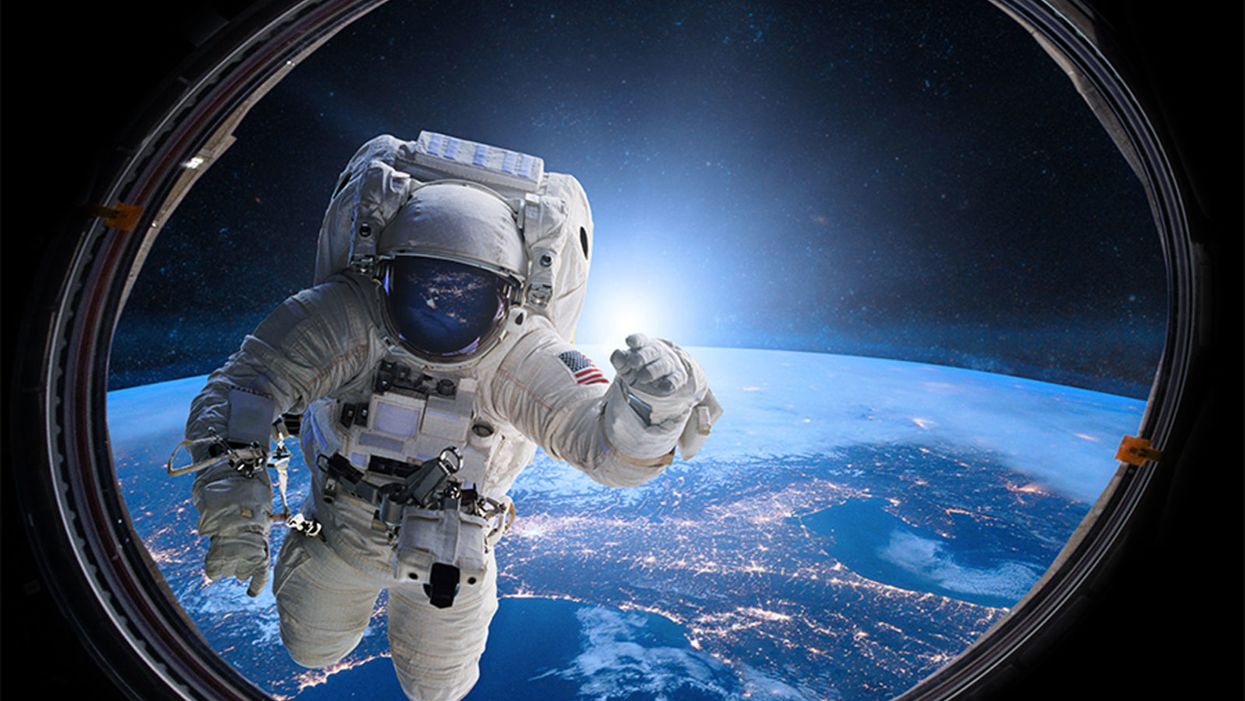
An astronaut peers through a portal in outer space.
What if people could just survive on sunlight like plants?
The admittedly outlandish question occurred to me after reading about how climate change will exacerbate drought, flooding, and worldwide food shortages. Many of these problems could be eliminated if human photosynthesis were possible. Had anyone ever tried it?
Extreme space travel exists at an ethically unique spot that makes human experimentation much more palatable.
I emailed Sidney Pierce, professor emeritus in the Department of Integrative Biology at the University of South Florida, who studies a type of sea slug, Elysia chlorotica, that eats photosynthetic algae, incorporating the algae's key cell structure into itself. It's still a mystery how exactly a slug can operate the part of the cell that converts sunlight into energy, which requires proteins made by genes to function, but the upshot is that the slugs can (and do) live on sunlight in-between feedings.
Pierce says he gets questions about human photosynthesis a couple of times a year, but it almost certainly wouldn't be worth it to try to develop the process in a human. "A high-metabolic rate, large animal like a human could probably not survive on photosynthesis," he wrote to me in an email. "The main reason is a lack of surface area. They would either have to grow leaves or pull a trailer covered with them."
In short: Plants have already exploited the best tricks for subsisting on photosynthesis, and unless we want to look and act like plants, we won't have much success ourselves. Not that it stopped Pierce from trying to develop human photosynthesis technology anyway: "I even tried to sell it to the Navy back in the day," he told me. "Imagine photosynthetic SEALS."
It turns out, however, that while no one is actively trying to create photosynthetic humans, scientists are considering the ways humans might need to change to adapt to future environments, either here on the rapidly changing Earth or on another planet. Rice University biologist Scott Solomon has written an entire book, Future Humans, in which he explores the environmental pressures that are likely to influence human evolution from this point forward. On Earth, Solomon says, infectious disease will remain a major driver of change. As for Mars, the big two are lower gravity and radiation, the latter of which bombards the Martian surface constantly because the planet has no magnetosphere.
Although he considers this example "pretty out there," Solomon says one possible solution to Mars' magnetic assault could leave humans not photosynthetic green, but orange, thanks to pigments called carotenoids that are responsible for the bright hues of pumpkins and carrots.
"Carotenoids protect against radiation," he says. "Usually only plants and microbes can produce carotenoids, but there's at least one kind of insect, a particular type of aphid, that somehow acquired the gene for making carotenoids from a fungus. We don't exactly know how that happened, but now they're orange... I view that as an example of, hey, maybe humans on Mars will evolve new kinds of pigmentation that will protect us from the radiation there."
We could wait for an orange human-producing genetic variation to occur naturally, or with new gene editing techniques such as CRISPR-Cas9, we could just directly give astronauts genetic advantages such as carotenoid-producing skin. This may not be as far-off as it sounds: Extreme space travel exists at an ethically unique spot that makes human experimentation much more palatable. If an astronaut already plans to subject herself to the enormous experiment of traveling to, and maybe living out her days on, a dangerous and faraway planet, do we have any obligation to provide all the protection we can?
Probably the most vocal person trying to figure out what genetic protections might help astronauts is Cornell geneticist Chris Mason. His lab has outlined a 10-phase, 500-year plan for human survival, starting with the comparatively modest goal of establishing which human genes are not amenable to change and should be marked with a "Do not disturb" sign.
To be clear, Mason is not actually modifying human beings. Instead, his lab has studied genes in radiation-resistant bacteria, such as the Deinococcus genus. They've expressed proteins called DSUP from tardigrades, tiny water bears that can survive in space, in human cells. They've looked into p53, a gene that is overexpressed in elephants and seems to protect them from cancer. They also developed a protocol to work on the NASA twin study comparing astronauts Scott Kelly, who spent a year aboard the International Space Station, and his brother Mark, who did not, to find out what effects space tends to have on genes in the first place.
In a talk he gave in December, Mason reported that 8.7 percent of Scott Kelly's genes—mostly those associated with immune function, DNA repair, and bone formation—did not return to normal after the astronaut had been home for six months. "Some of these space genes, we could engineer them, activate them, have them be hyperactive when you go to space," he said in that same talk. "When we think about having the hubris to go to a faraway planet...it seems like an almost impossible idea….but I really like people and I want us to survive for a long time, and this is the first step on the stairwell to survive out of the solar system."
What is the most important ability we could give our future selves through science?
There are others performing studies to figure out what capabilities we might bestow on the future-proof superhuman, but none of them are quite as extreme as photosynthesis (although all of them are useful). At Harvard, geneticist George Church wants to engineer cells to be resistant to viruses, such as the common cold and HIV. At Columbia, synthetic biologist Harris Wang is addressing self-sufficient humans more directly—trying to spur kidney cells to produce amino acids that are normally only available from diet.
But perhaps Future Humans author Scott Solomon has the most radical idea. I asked him a version of the classic What would be your superhero power? question: What does he see as the most important ability we could give our future selves through science?
"The empathy gene," he said. "The ability to put yourself in someone else's shoes and see the world as they see it. I think it would solve a lot of our problems."
A model of a human kidney.
Science's dream of creating perfect custom organs on demand as soon as a patient needs one is still a long way off. But tiny versions are already serving as useful research tools and stepping stones toward full-fledged replacements.
Although organoids cannot yet replace kidneys, they are invaluable tools for research.
The Lowdown
Australian researchers have grown hundreds of mini human kidneys in the past few years. Known as organoids, they function much like their full-grown counterparts, minus a few features due to a lack of blood supply.
Cultivated in a petri dish, these kidneys are still a shadow of their human counterparts. They grow no larger than one-sixth of an inch in diameter; fully developed organs are up to five inches in length. They contain no more than a few dozen nephrons, the kidney's individual blood-filtering unit, whereas a fully-grown kidney has about 1 million nephrons. And the dish variety live for just a few weeks.
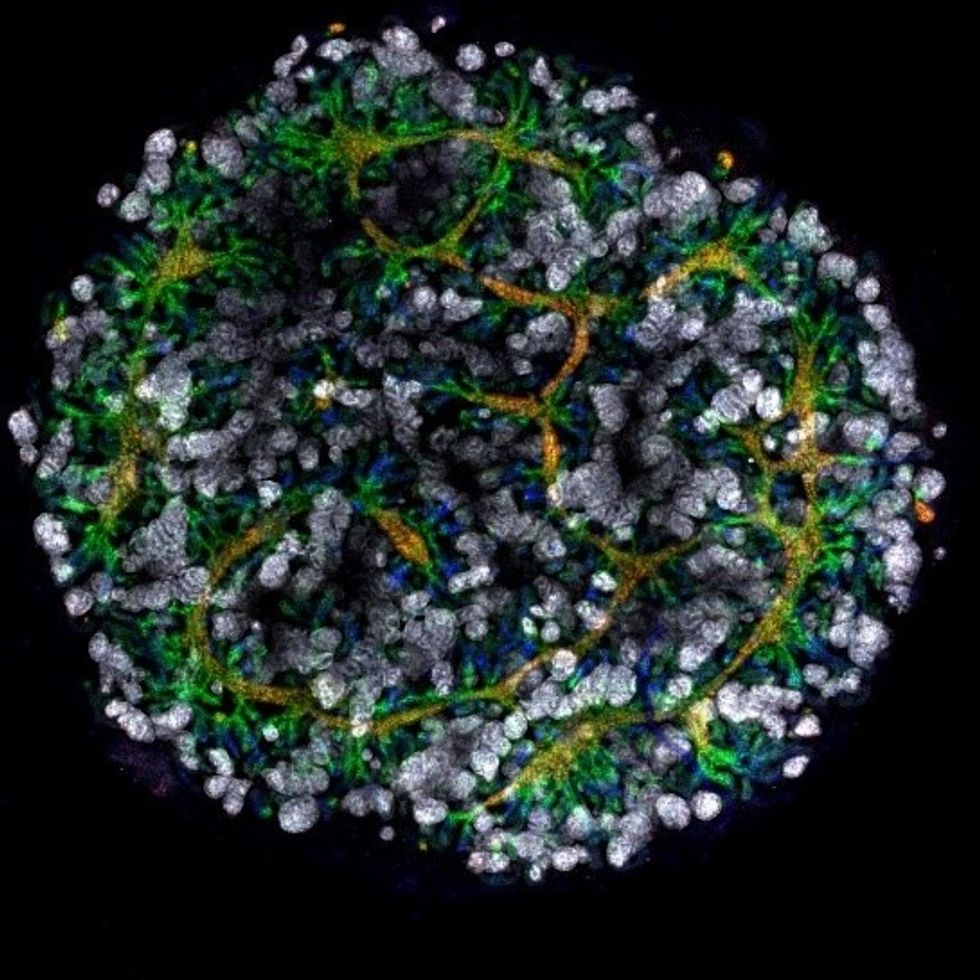
An organoid kidney created by the Murdoch Children's Institute in Melbourne, Australia.
Photo Credit: Shahnaz Khan.
But Melissa Little, head of the kidney research laboratory at the Murdoch Children's Institute in Melbourne, says these organoids are invaluable tools for research. Although renal failure is rare in children, more than half of those who suffer from such a disorder inherited it.
The mini kidneys enable scientists to better understand the progression of such disorders because they can be grown with a patient's specific genetic condition.
Mature stem cells can be extracted from a patient's blood sample and then reprogrammed to become like embryonic cells, able to turn into any type of cell in the body. It's akin to walking back the clock so that the cells regain unlimited potential for development. (The Japanese scientist who pioneered this technique was awarded the Nobel Prize in 2012.) These "induced pluripotent stem cells" can then be chemically coaxed to grow into mini kidneys that have the patient's genetic disorder.
"The (genetic) defects are quite clear in the organoids, and they can be monitored in the dish," Little says. To date, her research team has created organoids from 20 different stem cell lines.
Medication regimens can also be tested on the organoids, allowing specific tailoring for each patient. For now, such testing remains restricted to mice, but Little says it eventually will be done on human organoids so that the results can more accurately reflect how a given patient will respond to particular drugs.
Next Steps
Although these organoids cannot yet replace kidneys, Little says they may plug a huge gap in renal care by assisting in developing new treatments for chronic conditions. Currently, most patients with a serious kidney disorder see their options narrow to dialysis or organ transplantation. The former not only requires multiple sessions a week, but takes a huge toll on patient health.
Ten percent of older patients on dialysis die every year in the U.S. Aside from the physical trauma of organ transplantation, finding a suitable donor outside of a family member can be difficult.
"This is just another great example of the potential of pluripotent stem cells."
Meanwhile, the ongoing creation of organoids is supplying Little and her colleagues with enough information to create larger and more functional organs in the future. According to Little, researchers in the Netherlands, for example, have found that implanting organoids in mice leads to the creation of vascular growth, a potential pathway toward creating bigger and better kidneys.
And while Little acknowledges that creating a fully-formed custom organ is the ultimate goal, the mini organs are an important bridge step.
"This is just another great example of the potential of pluripotent stem cells, and I am just passionate to see it do some good."