Genetically Sequencing Healthy Babies Yielded Surprising Results

A newborn and mother in the hospital - first touch. (© martin81/Shutterstock)
Today in Melrose, Massachusetts, Cora Stetson is the picture of good health, a bubbly precocious 2-year-old. But Cora has two separate mutations in the gene that produces a critical enzyme called biotinidase and her body produces only 40 percent of the normal levels of that enzyme.
In the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach.
That's enough to pass conventional newborn (heelstick) screening, but may not be enough for normal brain development, putting baby Cora at risk for seizures and cognitive impairment. But thanks to an experimental study in which Cora's DNA was sequenced after birth, this condition was discovered and she is being treated with a safe and inexpensive vitamin supplement.
Stories like these are beginning to emerge from the BabySeq Project, the first clinical trial in the world to systematically sequence healthy newborn infants. This trial was led by my research group with funding from the National Institutes of Health. While still controversial, it is pointing the way to a future in which adults, or even newborns, can receive comprehensive genetic analysis in order to determine their risk of future disease and enable opportunities to prevent them.
Some believe that medicine is still not ready for genomic population screening, but others feel it is long overdue. After all, the sequencing of the Human Genome Project was completed in 2003, and with this milestone, it became feasible to sequence and interpret the genome of any human being. The costs have come down dramatically since then; an entire human genome can now be sequenced for about $800, although the costs of bioinformatic and medical interpretation can add another $200 to $2000 more, depending upon the number of genes interrogated and the sophistication of the interpretive effort.

Two-year-old Cora Stetson, whose DNA sequencing after birth identified a potentially dangerous genetic mutation in time for her to receive preventive treatment.
(Photo courtesy of Robert Green)
The ability to sequence the human genome yielded extraordinary benefits in scientific discovery, disease diagnosis, and targeted cancer treatment. But the ability of genomes to detect health risks in advance, to actually predict the medical future of an individual, has been mired in controversy and slow to manifest. In particular, the oft-cited vision that healthy infants could be genetically tested at birth in order to predict and prevent the diseases they would encounter, has proven to be far tougher to implement than anyone anticipated.
But in the last few years, the dream of predicting and preventing diseases through genomics, starting in childhood, is finally within reach. Why did it take so long? And what remains to be done?
Great Expectations
Part of the problem was the unrealistic expectations that had been building for years in advance of the genomic science itself. For example, the 1997 film Gattaca portrayed a near future in which the lifetime risk of disease was readily predicted the moment an infant is born. In the fanfare that accompanied the completion of the Human Genome Project, the notion of predicting and preventing future disease in an individual became a powerful meme that was used to inspire investment and public support for genomic research long before the tools were in place to make it happen.
Another part of the problem was the success of state-mandated newborn screening programs that began in the 1960's with biochemical tests of the "heel-stick" for babies with metabolic disorders. These programs have worked beautifully, costing only a few dollars per baby and saving thousands of infants from death and severe cognitive impairment. It seemed only logical that a new technology like genome sequencing would add power and promise to such programs. But instead of embracing the notion of newborn sequencing, newborn screening laboratories have thus far rejected the entire idea as too expensive, too ambiguous, and too threatening to the comfortable constituency that they had built within the public health framework.
"What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Creating the Evidence Base for Preventive Genomics
Despite a number of obstacles, there are researchers who are exploring how to achieve the original vision of genomic testing as a tool for disease prediction and prevention. For example, in our NIH-funded MedSeq Project, we were the first to ask the question: "What can you find when you look as deeply as possible into the medical genomes of healthy individuals?"
Most people do not understand that genetic information comes in four separate categories: 1) dominant mutations putting the individual at risk for rare conditions like familial forms of heart disease or cancer, (2) recessive mutations putting the individual's children at risk for rare conditions like cystic fibrosis or PKU, (3) variants across the genome that can be tallied to construct polygenic risk scores for common conditions like heart disease or type 2 diabetes, and (4) variants that can influence drug metabolism or predict drug side effects such as the muscle pain that occasionally occurs with statin use.
The technological and analytical challenges of our study were formidable, because we decided to systematically interrogate over 5000 disease-associated genes and report results in all four categories of genetic information directly to the primary care physicians for each of our volunteers. We enrolled 200 adults and found that everyone who was sequenced had medically relevant polygenic and pharmacogenomic results, over 90 percent carried recessive mutations that could have been important to reproduction, and an extraordinary 14.5 percent carried dominant mutations for rare genetic conditions.
A few years later we launched the BabySeq Project. In this study, we restricted the number of genes to include only those with child/adolescent onset that could benefit medically from early warning, and even so, we found 9.4 percent carried dominant mutations for rare conditions.
At first, our interpretation around the high proportion of apparently healthy individuals with dominant mutations for rare genetic conditions was simple – that these conditions had lower "penetrance" than anticipated; in other words, only a small proportion of those who carried the dominant mutation would get the disease. If this interpretation were to hold, then genetic risk information might be far less useful than we had hoped.
Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
But then we circled back with each adult or infant in order to examine and test them for any possible features of the rare disease in question. When we did this, we were surprised to see that in over a quarter of those carrying such mutations, there were already subtle signs of the disease in question that had not even been suspected! Now our interpretation was different. We now believe that genetic risk may be responsible for subclinical disease in a much higher proportion of people than has ever been suspected!
Meanwhile, colleagues of ours have been demonstrating that detailed analysis of polygenic risk scores can identify individuals at high risk for common conditions like heart disease. So adding up the medically relevant results in any given genome, we start to see that you can learn your risks for a rare monogenic condition, a common polygenic condition, a bad effect from a drug you might take in the future, or for having a child with a devastating recessive condition. Suddenly the information available in the genome of even an apparently healthy individual is looking more robust, and the prospect of preventive genomics is looking feasible.
Preventive Genomics Arrives in Clinical Medicine
There is still considerable evidence to gather before we can recommend genomic screening for the entire population. For example, it is important to make sure that families who learn about such risks do not suffer harms or waste resources from excessive medical attention. And many doctors don't yet have guidance on how to use such information with their patients. But our research is convincing many people that preventive genomics is coming and that it will save lives.
In fact, we recently launched a Preventive Genomics Clinic at Brigham and Women's Hospital where information-seeking adults can obtain predictive genomic testing with the highest quality interpretation and medical context, and be coached over time in light of their disease risks toward a healthier outcome. Insurance doesn't yet cover such testing, so patients must pay out of pocket for now, but they can choose from a menu of genetic screening tests, all of which are more comprehensive than consumer-facing products. Genetic counseling is available but optional. So far, this service is for adults only, but sequencing for children will surely follow soon.
As the costs of sequencing and other Omics technologies continue to decline, we will see both responsible and irresponsible marketing of genetic testing, and we will need to guard against unscientific claims. But at the same time, we must be far more imaginative and fast moving in mainstream medicine than we have been to date in order to claim the emerging benefits of preventive genomics where it is now clear that suffering can be averted, and lives can be saved. The future has arrived if we are bold enough to grasp it.
Funding and Disclosures:
Dr. Green's research is supported by the National Institutes of Health, the Department of Defense and through donations to The Franca Sozzani Fund for Preventive Genomics. Dr. Green receives compensation for advising the following companies: AIA, Applied Therapeutics, Helix, Ohana, OptraHealth, Prudential, Verily and Veritas; and is co-founder and advisor to Genome Medical, Inc, a technology and services company providing genetics expertise to patients, providers, employers and care systems.
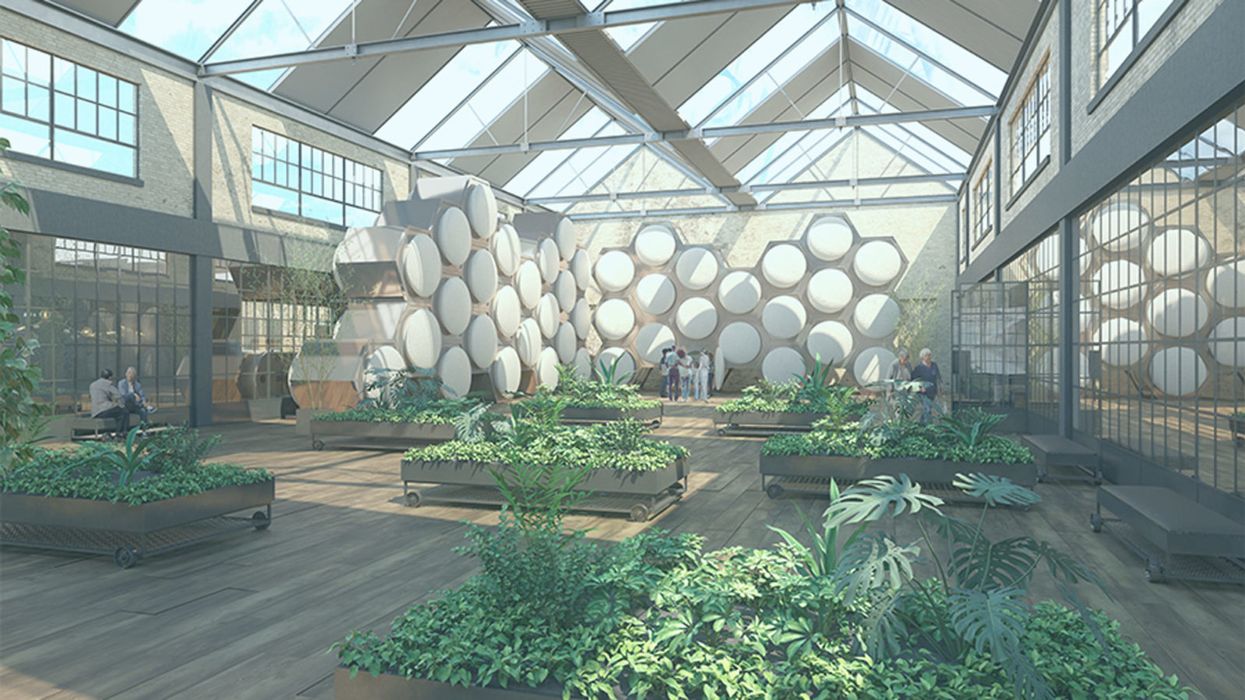
The Grim Reaper Can Now Compost Your Body
An artist's rendering of a future Recompose facility in Washington state, with reusable modular vessels that convert human remains to soil.
Ultra-green Seattle isn't just getting serious about living eco-friendly, but dying that way, too. As of this week, Washington is officially the first state to allow citizens to compost their own dead bodies.
Their bodies, including bones, were converted into clean, odorless soil free of harmful pathogens.
The Lowdown
Keep in mind this doesn't mean dumping your relative in a nearby river. Scientists and organizations have ways to help Mother Nature process the remains. For instance, the late actor Luke Perry reportedly was buried in a mushroom suit. Perry's garment is completely biodegradable and the attached microorganisms help the decomposition process cleanly and efficiently.
A biodegradable burial requires only a fraction of the energy used for cremation and can save a metric ton of CO2. The body decomposes in about a month. Besides a mushroom suit, another option coming down the pike in Washington state is to have your body converted directly into soil in a special facility.
A pilot study last summer by a public benefit corporation called Recompose signed up six terminally ill people who donated their remains for such research. Their bodies, including bones, were converted into clean, odorless soil free of harmful pathogens. That soil—about a cubic yard per person--could then be returned after 30 days to the subjects' families.
Green burials open the door to creative memorials. A tree or garden could be planted with your soil. This method provides a climate-friendly alternative to traditional funerals, circumventing toxic embalming fluid, expensive casket materials and other ecological overhead. The fertile soil could also be given to conservationist organizations.
Next Up
The new legislation in Washington will take effect May 1, 2020. The Pacific Northwest state has one of the highest cremation rates in the nation at 78 percent, only second to Nevada. Rising climate change and increased interest in death management will only speed this discussion to the forefront in other states.
A biodegradable burial requires only a fraction of the energy used for cremation and can save a metric ton of CO2.
It's also worth noting Perry wasn't buried in Washington State, but in Tennessee. It is unknown where exactly he was laid to rest, nor if it was done under a legal precedent or special exception.
According to the Green Burial Council, each state varies on how and where you can bury someone. Home burials are usually legal, but to do so requires establishing an official cemetery area on the property. How someone is buried has even more dynamic legislation. There will be new discussions about how neighbors contend with nearby decomposing bodies, legal limitations to private burial techniques, and other issues never addressed before in modern mainstream America.
Open Questions
It's unclear if green burials will be commonplace for those with less financial means or access. Mushroom suits average a couple thousand dollars, making them more expensive than a low-end casket. There are also the less obvious expenses, including designating the place of burial, and getting proper burial support and guidance. In short, you likely won't go to the local funeral home and be taken care of properly. It is still experimental.
As for "natural organic reduction" (converting human remains to soil in reusable modular vessels), Recompose is still figuring out its pricing for Washington residents, but expects the service to cost more than cremation and less than a conventional burial.
For now, environmentally sustainable death care may be comparable to vegetarianism in the 1970s or solar paneling in the 1980s: A discussion among urbanites and upwardly-mobile financial classes, but not yet an accessible option for the average American. It's not a coincidence that the new Washington law received support in Seattle, one of the top 10 wealthiest cities in America. A similar push may take off in less affluent areas if ecological concerns drive a demand for affordable green burial options.
Until then, your neighborhood mortician still has the death business on lock.
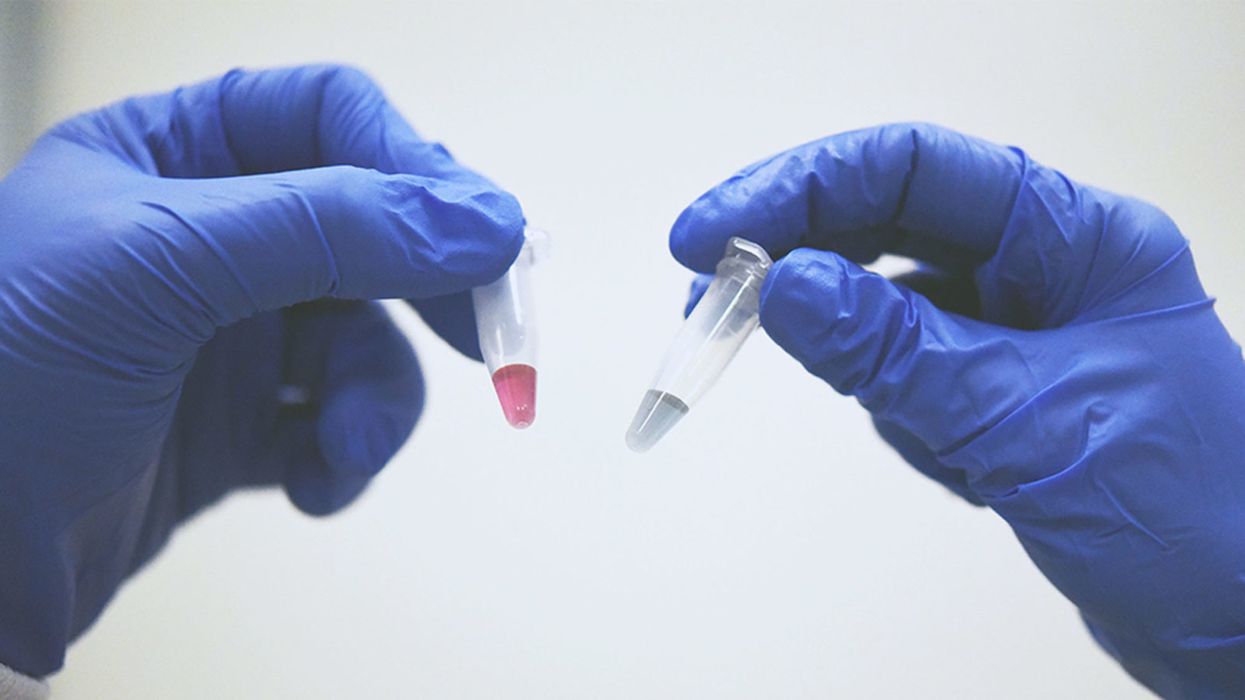
A simple ten-minute universal cancer test that can be detected by the human eye or an electronic device - published in Nature Communications (Dec 2018) by the Trau lab at the University of Queensland. Red indicates the presence of cancerous cells and blue doesn't.
Matt Trau, a professor of chemistry at the University of Queensland, stunned the science world back in December when the prestigious journal Nature Communications published his lab's discovery about a unique property of cancer DNA that could lead to a simple, cheap, and accurate test to detect any type of cancer in under 10 minutes.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke."
Trau granted very few interviews in the wake of the news, but he recently opened up to leapsmag about the significance of this promising early research. Here is his story in his own words, as told to Editor-in-Chief Kira Peikoff.
There's been an incredible explosion of knowledge over the past 20 years, particularly since the genome was sequenced. The area of diagnostics has a tremendous amount of promise and has caught our lab's interest. If you catch cancer early, you can improve survival rates to as high as 98 percent, sometimes even now surpassing that.
My lab is interested in devices to improve the trajectory of cancer patients. So, once people get diagnosed, can we get really sophisticated information about the molecular origins of the disease, and can we measure it in real time? And then can we match that with the best treatment and monitor it in real time, too?
I think those approaches, also coupled with immunotherapy, where one dreams of monitoring the immune system simultaneously with the disease progress, will be the future.
But currently, the methodologies for cancer are still pretty old. So, for example, let's talk about biopsies in general. Liquid biopsy just means using a blood test or a urine test, rather than extracting out a piece of solid tissue. Now consider breast cancer. Still, the cutting-edge screening method is mammography or the physical interrogation for lumps. This has had a big impact in terms of early detection and awareness, but it's still primitive compared to interrogating, forensically, blood samples to look at traces of DNA.
Large machines like CAT scans, PET scans, MRIs, are very expensive and very subjective in terms of the operator. They don't look at the root causes of the cancer. Cancer is caused by changes in DNA. These can be changes in the hard drive of the DNA (the genomic changes) or changes in the apps that the DNA are running (the epigenetics and the transcriptomics).
We don't look at that now, even though we have, emerging, all of these technologies to do it, and those technologies are getting so much cheaper. I saw some statistics at a conference just a few months ago that, in the United States, less than 1 percent of cancer patients have their DNA interrogated. That's the current state-of-the-art in the modern medical system.

Professor Matt Trau, a cancer researcher at the University of Queensland in Australia.
(Courtesy)
Blood, as the highway of the body, is carrying all of this information. Cancer cells, if they are present in the body, are constantly getting turned over. When they die, they release their contents into the blood. Many of these cells end up in the urine and saliva. Having technologies that can forensically scan the highways looking for evidence of cancer is little bit like looking for explosives at the airport. That's very valuable as a security tool.
The trouble is that there are thousands of different types of cancer. Going back to breast cancer, there's at least a dozen different types, probably more, and each of them change the DNA (the hard drive of the disease) and the epigenetics (or the RAM memory). So one of the problems for diagnostics in cancer is to find something that is a signature of all cancers. That's been a really, really, really difficult problem.
Ours was a completely serendipitous discovery. What we found in the lab was this one marker that just kept coming up in all of the types of breast cancers we were studying.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke, maybe it works just for breast cancer." So we went on to test it in prostate cancer, which is also many different types of diseases, and it seemed to be working in all of those. We then tested it further in lymphoma. Again, many different types of lymphoma. It worked across all of those. We tested it in gastrointestinal cancer. Again, many different types, and still, it worked, but we were skeptical.
Then we looked at cell lines, which are cells that have come from previous cancer patients, that we grow in the lab, but are used as model experimental systems. We have many of those cell lines, both ones that are cancerous, and ones that are healthy. It was quite remarkable that the marker worked in all of the cancer cell lines and didn't work in the healthy cell lines.
What could possibly be going on?
Well, imagine DNA as a piece of string, that's your hard drive. Epigenetics is like the beads that you put on that string. Those beads you can take on and off as you wish and they control which apps are run, meaning which genetic programs the cell runs. We hypothesized that for cancer, those beads cluster together, rather than being randomly distributed across the string.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office.
The implications of this are profound. It means that DNA from cancer folds in water into three-dimensional structures that are very different from healthy cells' DNA. It's quite literally the needle in a haystack. Because when you do a liquid biopsy for early detection of cancer, most of the DNA from blood contains a vast abundance of healthy DNA. And that's not of interest. What's of interest is to find the cancerous DNA. That's there only in trace.
Once we figured out what was going on, we could easily set up a system to detect the trace cancerous DNA. It binds to gold nanoparticles in water and changes color. The test takes 10 minutes, and you can detect it by eye. Red indicates cancer and blue doesn't.
We're very, very excited about where we go from here. We're starting to test the test on a greater number of cancers, in thousands of patient samples. We're looking to the scientific community to engage with us, and we're getting a really good response from groups around the world who are supplying more samples to us so we can test this more broadly.
We also are very interested in testing how early can we go with this test. Can we detect cancer through a simple blood test even before there are any symptoms whatsoever? If so, we might be able to convert a cancer diagnosis to something almost as good as a vaccine.
Of course, we have to watch what are called false positives. We don't want to be detecting people as positives when they don't have cancer, and so the technology needs to improve there. We see this version as the iPhone 1. We're interested in the iPhone 2, 3, 4, getting better and better.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office. If it came back positive, your doctor could say, "Look, there's some news here, but actually, it's not bad news, it's good news. We've caught this so early that we will be able to manage this, and this won't be a problem for you."
If this were to be in routine use in the medical system, countless lives could be saved. Cancer is now becoming one of the biggest killers in the world. We're talking millions upon millions upon millions of people who are affected. This really motivates our work. We might make a difference there.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.