What will the $100 genome mean?

A company has slashed the cost of assessing a person's genome to just $100. With lower costs - and as other genetic tools mature and evolve - a wave of new therapies could be coming in the near future.
In May 2022, Californian biotech Ultima Genomics announced that its UG 100 platform was capable of sequencing an entire human genome for just $100, a landmark moment in the history of the field. The announcement was particularly remarkable because few had previously heard of the company, a relative unknown in an industry long dominated by global giant Illumina which controls about 80 percent of the world’s sequencing market.
Ultima’s secret was to completely revamp many technical aspects of the way Illumina have traditionally deciphered DNA. The process usually involves first splitting the double helix DNA structure into single strands, then breaking these strands into short fragments which are laid out on a glass surface called a flow cell. When this flow cell is loaded into the sequencing machine, color-coded tags are attached to each individual base letter. A laser scans the bases individually while a camera simultaneously records the color associated with them, a process which is repeated until every single fragment has been sequenced.
Instead, Ultima has found a series of shortcuts to slash the cost and boost efficiency. “Ultima Genomics has developed a fundamentally new sequencing architecture designed to scale beyond conventional approaches,” says Josh Lauer, Ultima’s chief commercial officer.
This ‘new architecture’ is a series of subtle but highly impactful tweaks to the sequencing process ranging from replacing the costly flow cell with a silicon wafer which is both cheaper and allows more DNA to be read at once, to utilizing machine learning to convert optical data into usable information.
To put $100 genome in perspective, back in 2012 the cost of sequencing a single genome was around $10,000, a price tag which dropped to $1,000 a few years later. Before Ultima’s announcement, the cost of sequencing an individual genome was around $600.
Several studies have found that nearly 12 percent of healthy people who have their genome sequenced, then discover they have a variant pointing to a heightened risk of developing a disease that can be monitored, treated or prevented.
While Ultima’s new machine is not widely available yet, Illumina’s response has been rapid. In September 2022, the company unveiled the NovaSeq X series, which it describes as its fastest most cost-efficient sequencing platform yet, capable of sequencing genomes at $200, with further price cuts likely to follow.
But what will the rapidly tumbling cost of sequencing actually mean for medicine? “Well to start with, obviously it’s going to mean more people getting their genome sequenced,” says Michael Snyder, professor of genetics at Stanford University. “It'll be a lot more accessible to people.”
At the moment sequencing is mainly limited to certain cancer patients where it is used to inform treatment options, and individuals with undiagnosed illnesses. In the past, initiatives such as SeqFirst have attempted further widen access to genome sequencing based on growing amounts of research illustrating the potential benefits of the technology in healthcare. Several studies have found that nearly 12 percent of healthy people who have their genome sequenced, then discover they have a variant pointing to a heightened risk of developing a disease that can be monitored, treated or prevented.
“While whole genome sequencing is not yet widely used in the U.S., it has started to come into pediatric critical care settings such as newborn intensive care units,” says Professor Michael Bamshad, who heads the genetic medicine division in the University of Washington’s pediatrics department. “It is also being used more often in outpatient clinical genetics services, particularly when conventional testing fails to identify explanatory variants.”
But the cost of sequencing itself is only one part of the price tag. The subsequent clinical interpretation and genetic counselling services often come to several thousand dollars, a cost which insurers are not always willing to pay.
As a result, while Bamshad and others hope that the arrival of the $100 genome will create new opportunities to use genetic testing in innovative ways, the most immediate benefits are likely to come in the realm of research.
Bigger Data
There are numerous ways in which cheaper sequencing is likely to advance scientific research, for example the ability to collect data on much larger patient groups. This will be a major boon to scientists working on complex heterogeneous diseases such as schizophrenia or depression where there are many genes involved which all exert subtle effects, as well as substantial variance across the patient population. Bigger studies could help scientists identify subgroups of patients where the disease appears to be driven by similar gene variants, who can then be more precisely targeted with specific drugs.
If insurers can figure out the economics, Snyder even foresees a future where at a certain age, all of us can qualify for annual sequencing of our blood cells to search for early signs of cancer or the potential onset of other diseases like type 2 diabetes.
David Curtis, a genetics professor at University College London, says that scientists studying these illnesses have previously been forced to rely on genome-wide association studies which are limited because they only identify common gene variants. “We might see a significant increase in the number of large association studies using sequence data,” he says. “It would be far preferable to use this because it provides information about rare, potentially functional variants.”
Cheaper sequencing will also aid researchers working on diseases which have traditionally been underfunded. Bamshad cites cystic fibrosis, a condition which affects around 40,000 children and adults in the U.S., as one particularly pertinent example.
“Funds for gene discovery for rare diseases are very limited,” he says. “We’re one of three sites that did whole genome sequencing on 5,500 people with cystic fibrosis, but our statistical power is limited. A $100 genome would make it much more feasible to sequence everyone in the U.S. with cystic fibrosis and make it more likely that we discover novel risk factors and pathways influencing clinical outcomes.”
For progressive diseases that are more common like cancer and type 2 diabetes, as well as neurodegenerative conditions like multiple sclerosis and ALS, geneticists will be able to go even further and afford to sequence individual tumor cells or neurons at different time points. This will enable them to analyze how individual DNA modifications like methylation, change as the disease develops.
In the case of cancer, this could help scientists understand how tumors evolve to evade treatments. Within in a clinical setting, the ability to sequence not just one, but many different cells across a patient’s tumor could point to the combination of treatments which offer the best chance of eradicating the entire cancer.
“What happens at the moment with a solid tumor is you treat with one drug, and maybe 80 percent of that tumor is susceptible to that drug,” says Neil Ward, vice president and general manager in the EMEA region for genomics company PacBio. “But the other 20 percent of the tumor has already got mutations that make it resistant, which is probably why a lot of modern therapies extend life for sadly only a matter of months rather than curing, because they treat a big percentage of the tumor, but not the whole thing. So going forwards, I think that we will see genomics play a huge role in cancer treatments, through using multiple modalities to treat someone's cancer.”
If insurers can figure out the economics, Snyder even foresees a future where at a certain age, all of us can qualify for annual sequencing of our blood cells to search for early signs of cancer or the potential onset of other diseases like type 2 diabetes.
“There are companies already working on looking for cancer signatures in methylated DNA,” he says. “If it was determined that you had early stage cancer, pre-symptomatically, that could then be validated with targeted MRI, followed by surgery or chemotherapy. It makes a big difference catching cancer early. If there were signs of type 2 diabetes, you could start taking steps to mitigate your glucose rise, and possibly prevent it or at least delay the onset.”
This would already revolutionize the way we seek to prevent a whole range of illnesses, but others feel that the $100 genome could also usher in even more powerful and controversial preventative medicine schemes.
Newborn screening
In the eyes of Kári Stefánsson, the Icelandic neurologist who been a visionary for so many advances in the field of human genetics over the last 25 years, the falling cost of sequencing means it will be feasible to sequence the genomes of every baby born.
“We have recently done an analysis of genomes in Iceland and the UK Biobank, and in 4 percent of people you find mutations that lead to serious disease, that can be prevented or dealt with,” says Stefansson, CEO of deCODE genetics, a subsidiary of the pharmaceutical company Amgen. “This could transform our healthcare systems.”
As well as identifying newborns with rare diseases, this kind of genomic information could be used to compute a person’s risk score for developing chronic illnesses later in life. If for example, they have a higher than average risk of colon or breast cancer, they could be pre-emptively scheduled for annual colonoscopies or mammograms as soon as they hit adulthood.
To a limited extent, this is already happening. In the UK, Genomics England has launched the Newborn Genomes Programme, which plans to undertake whole-genome sequencing of up to 200,000 newborn babies, with the aim of enabling the early identification of rare genetic diseases.
"I have not had my own genome sequenced and I would not have wanted my parents to have agreed to this," Curtis says. "I don’t see that sequencing children for the sake of some vague, ill-defined benefits could ever be justifiable.”
However, some scientists feel that it is tricky to justify sequencing the genomes of apparently healthy babies, given the data privacy issues involved. They point out that we still know too little about the links which can be drawn between genetic information at birth, and risk of chronic illness later in life.
“I think there are very difficult ethical issues involved in sequencing children if there are no clear and immediate clinical benefits,” says Curtis. “They cannot consent to this process. I have not had my own genome sequenced and I would not have wanted my parents to have agreed to this. I don’t see that sequencing children for the sake of some vague, ill-defined benefits could ever be justifiable.”
Curtis points out that there are many inherent risks about this data being available. It may fall into the hands of insurance companies, and it could even be used by governments for surveillance purposes.
“Genetic sequence data is very useful indeed for forensic purposes. Its full potential has yet to be realized but identifying rare variants could provide a quick and easy way to find relatives of a perpetrator,” he says. “If large numbers of people had been sequenced in a healthcare system then it could be difficult for a future government to resist the temptation to use this as a resource to investigate serious crimes.”
While sequencing becoming more widely available will present difficult ethical and moral challenges, it will offer many benefits for society as a whole. Cheaper sequencing will help boost the diversity of genomic datasets which have traditionally been skewed towards individuals of white, European descent, meaning that much of the actionable medical information which has come out of these studies is not relevant to people of other ethnicities.
Ward predicts that in the coming years, the growing amount of genetic information will ultimately change the outcomes for many with rare, previously incurable illnesses.
“If you're the parent of a child that has a susceptible or a suspected rare genetic disease, their genome will get sequenced, and while sadly that doesn’t always lead to treatments, it’s building up a knowledge base so companies can spring up and target that niche of a disease,” he says. “As a result there’s a whole tidal wave of new therapies that are going to come to market over the next five years, as the genetic tools we have, mature and evolve.”
This article was first published by Leaps.org in October 2022.
Thanks to safety cautions from the COVID-19 pandemic, a strain of influenza has been completely eliminated.
If you were one of the millions who masked up, washed your hands thoroughly and socially distanced, pat yourself on the back—you may have helped change the course of human history.
Scientists say that thanks to these safety precautions, which were introduced in early 2020 as a way to stop transmission of the novel COVID-19 virus, a strain of influenza has been completely eliminated. This marks the first time in human history that a virus has been wiped out through non-pharmaceutical interventions, such as vaccines.
The flu shot, explained
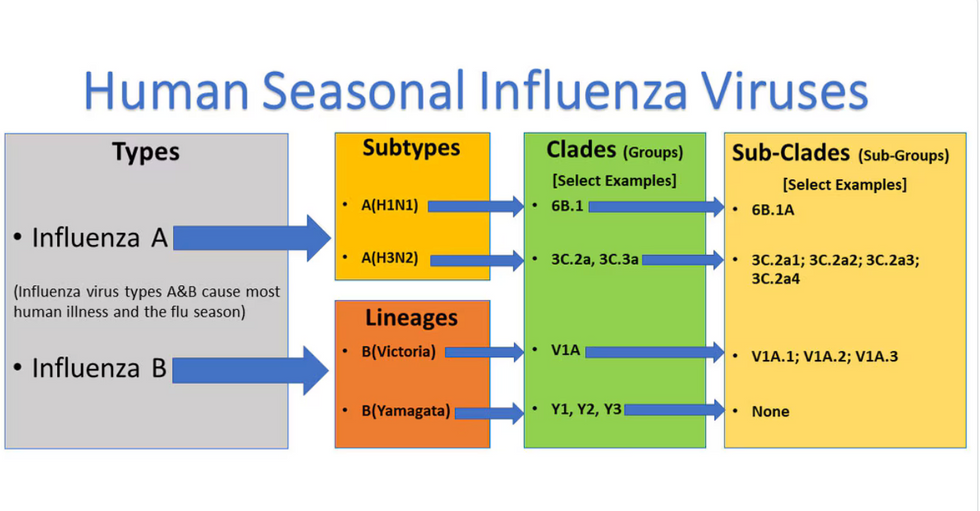
Influenza viruses type A and B are responsible for the majority of human illnesses and the flu season.
Centers for Disease Control
For more than a decade, flu shots have protected against two types of the influenza virus–type A and type B. While there are four different strains of influenza in existence (A, B, C, and D), only strains A, B, and C are capable of infecting humans, and only A and B cause pandemics. In other words, if you catch the flu during flu season, you’re most likely sick with flu type A or B.
Flu vaccines contain inactivated—or dead—influenza virus. These inactivated viruses can’t cause sickness in humans, but when administered as part of a vaccine, they teach a person’s immune system to recognize and kill those viruses when they’re encountered in the wild.
Each spring, a panel of experts gives a recommendation to the US Food and Drug Administration on which strains of each flu type to include in that year’s flu vaccine, depending on what surveillance data says is circulating and what they believe is likely to cause the most illness during the upcoming flu season. For the past decade, Americans have had access to vaccines that provide protection against two strains of influenza A and two lineages of influenza B, known as the Victoria lineage and the Yamagata lineage. But this year, the seasonal flu shot won’t include the Yamagata strain, because the Yamagata strain is no longer circulating among humans.
How Yamagata Disappeared

Flu surveillance data from the Global Initiative on Sharing All Influenza Data (GISAID) shows that the Yamagata lineage of flu type B has not been sequenced since April 2020.
Nature
Experts believe that the Yamagata lineage had already been in decline before the pandemic hit, likely because the strain was naturally less capable of infecting large numbers of people compared to the other strains. When the COVID-19 pandemic hit, the resulting safety precautions such as social distancing, isolating, hand-washing, and masking were enough to drive the virus into extinction completely.
Because the strain hasn’t been circulating since 2020, the FDA elected to remove the Yamagata strain from the seasonal flu vaccine. This will mark the first time since 2012 that the annual flu shot will be trivalent (three-component) rather than quadrivalent (four-component).
Should I still get the flu shot?
The flu shot will protect against fewer strains this year—but that doesn’t mean we should skip it. Influenza places a substantial health burden on the United States every year, responsible for hundreds of thousands of hospitalizations and tens of thousands of deaths. The flu shot has been shown to prevent millions of illnesses each year (more than six million during the 2022-2023 season). And while it’s still possible to catch the flu after getting the flu shot, studies show that people are far less likely to be hospitalized or die when they’re vaccinated.
Another unexpected benefit of dropping the Yamagata strain from the seasonal vaccine? This will possibly make production of the flu vaccine faster, and enable manufacturers to make more vaccines, helping countries who have a flu vaccine shortage and potentially saving millions more lives.
After his grandmother’s dementia diagnosis, one man invented a snack to keep her healthy and hydrated.
Founder Lewis Hornby and his grandmother Pat, sampling Jelly Drops—an edible gummy containing water and life-saving electrolytes.
On a visit to his grandmother’s nursing home in 2016, college student Lewis Hornby made a shocking discovery: Dehydration is a common (and dangerous) problem among seniors—especially those that are diagnosed with dementia.
Hornby’s grandmother, Pat, had always had difficulty keeping up her water intake as she got older, a common issue with seniors. As we age, our body composition changes, and we naturally hold less water than younger adults or children, so it’s easier to become dehydrated quickly if those fluids aren’t replenished. What’s more, our thirst signals diminish naturally as we age as well—meaning our body is not as good as it once was in letting us know that we need to rehydrate. This often creates a perfect storm that commonly leads to dehydration. In Pat’s case, her dehydration was so severe she nearly died.
When Lewis Hornby visited his grandmother at her nursing home afterward, he learned that dehydration especially affects people with dementia, as they often don’t feel thirst cues at all, or may not recognize how to use cups correctly. But while dementia patients often don’t remember to drink water, it seemed to Hornby that they had less problem remembering to eat, particularly candy.

Hornby wanted to create a solution for elderly people who struggled keeping their fluid intake up. He spent the next eighteen months researching and designing a solution and securing funding for his project. In 2019, Hornby won a sizable grant from the Alzheimer’s Society, a UK-based care and research charity for people with dementia and their caregivers. Together, through the charity’s Accelerator Program, they created a bite-sized, sugar-free, edible jelly drop that looked and tasted like candy. The candy, called Jelly Drops, contained 95% water and electrolytes—important minerals that are often lost during dehydration. The final product launched in 2020—and was an immediate success. The drops were able to provide extra hydration to the elderly, as well as help keep dementia patients safe, since dehydration commonly leads to confusion, hospitalization, and sometimes even death.
Not only did Jelly Drops quickly become a favorite snack among dementia patients in the UK, but they were able to provide an additional boost of hydration to hospital workers during the pandemic. In NHS coronavirus hospital wards, patients infected with the virus were regularly given Jelly Drops to keep their fluid levels normal—and staff members snacked on them as well, since long shifts and personal protective equipment (PPE) they were required to wear often left them feeling parched.
In April 2022, Jelly Drops launched in the United States. The company continues to donate 1% of its profits to help fund Alzheimer’s research.

