What will the $100 genome mean?

A company has slashed the cost of assessing a person's genome to just $100. With lower costs - and as other genetic tools mature and evolve - a wave of new therapies could be coming in the near future.
In May 2022, Californian biotech Ultima Genomics announced that its UG 100 platform was capable of sequencing an entire human genome for just $100, a landmark moment in the history of the field. The announcement was particularly remarkable because few had previously heard of the company, a relative unknown in an industry long dominated by global giant Illumina which controls about 80 percent of the world’s sequencing market.
Ultima’s secret was to completely revamp many technical aspects of the way Illumina have traditionally deciphered DNA. The process usually involves first splitting the double helix DNA structure into single strands, then breaking these strands into short fragments which are laid out on a glass surface called a flow cell. When this flow cell is loaded into the sequencing machine, color-coded tags are attached to each individual base letter. A laser scans the bases individually while a camera simultaneously records the color associated with them, a process which is repeated until every single fragment has been sequenced.
Instead, Ultima has found a series of shortcuts to slash the cost and boost efficiency. “Ultima Genomics has developed a fundamentally new sequencing architecture designed to scale beyond conventional approaches,” says Josh Lauer, Ultima’s chief commercial officer.
This ‘new architecture’ is a series of subtle but highly impactful tweaks to the sequencing process ranging from replacing the costly flow cell with a silicon wafer which is both cheaper and allows more DNA to be read at once, to utilizing machine learning to convert optical data into usable information.
To put $100 genome in perspective, back in 2012 the cost of sequencing a single genome was around $10,000, a price tag which dropped to $1,000 a few years later. Before Ultima’s announcement, the cost of sequencing an individual genome was around $600.
Several studies have found that nearly 12 percent of healthy people who have their genome sequenced, then discover they have a variant pointing to a heightened risk of developing a disease that can be monitored, treated or prevented.
While Ultima’s new machine is not widely available yet, Illumina’s response has been rapid. In September 2022, the company unveiled the NovaSeq X series, which it describes as its fastest most cost-efficient sequencing platform yet, capable of sequencing genomes at $200, with further price cuts likely to follow.
But what will the rapidly tumbling cost of sequencing actually mean for medicine? “Well to start with, obviously it’s going to mean more people getting their genome sequenced,” says Michael Snyder, professor of genetics at Stanford University. “It'll be a lot more accessible to people.”
At the moment sequencing is mainly limited to certain cancer patients where it is used to inform treatment options, and individuals with undiagnosed illnesses. In the past, initiatives such as SeqFirst have attempted further widen access to genome sequencing based on growing amounts of research illustrating the potential benefits of the technology in healthcare. Several studies have found that nearly 12 percent of healthy people who have their genome sequenced, then discover they have a variant pointing to a heightened risk of developing a disease that can be monitored, treated or prevented.
“While whole genome sequencing is not yet widely used in the U.S., it has started to come into pediatric critical care settings such as newborn intensive care units,” says Professor Michael Bamshad, who heads the genetic medicine division in the University of Washington’s pediatrics department. “It is also being used more often in outpatient clinical genetics services, particularly when conventional testing fails to identify explanatory variants.”
But the cost of sequencing itself is only one part of the price tag. The subsequent clinical interpretation and genetic counselling services often come to several thousand dollars, a cost which insurers are not always willing to pay.
As a result, while Bamshad and others hope that the arrival of the $100 genome will create new opportunities to use genetic testing in innovative ways, the most immediate benefits are likely to come in the realm of research.
Bigger Data
There are numerous ways in which cheaper sequencing is likely to advance scientific research, for example the ability to collect data on much larger patient groups. This will be a major boon to scientists working on complex heterogeneous diseases such as schizophrenia or depression where there are many genes involved which all exert subtle effects, as well as substantial variance across the patient population. Bigger studies could help scientists identify subgroups of patients where the disease appears to be driven by similar gene variants, who can then be more precisely targeted with specific drugs.
If insurers can figure out the economics, Snyder even foresees a future where at a certain age, all of us can qualify for annual sequencing of our blood cells to search for early signs of cancer or the potential onset of other diseases like type 2 diabetes.
David Curtis, a genetics professor at University College London, says that scientists studying these illnesses have previously been forced to rely on genome-wide association studies which are limited because they only identify common gene variants. “We might see a significant increase in the number of large association studies using sequence data,” he says. “It would be far preferable to use this because it provides information about rare, potentially functional variants.”
Cheaper sequencing will also aid researchers working on diseases which have traditionally been underfunded. Bamshad cites cystic fibrosis, a condition which affects around 40,000 children and adults in the U.S., as one particularly pertinent example.
“Funds for gene discovery for rare diseases are very limited,” he says. “We’re one of three sites that did whole genome sequencing on 5,500 people with cystic fibrosis, but our statistical power is limited. A $100 genome would make it much more feasible to sequence everyone in the U.S. with cystic fibrosis and make it more likely that we discover novel risk factors and pathways influencing clinical outcomes.”
For progressive diseases that are more common like cancer and type 2 diabetes, as well as neurodegenerative conditions like multiple sclerosis and ALS, geneticists will be able to go even further and afford to sequence individual tumor cells or neurons at different time points. This will enable them to analyze how individual DNA modifications like methylation, change as the disease develops.
In the case of cancer, this could help scientists understand how tumors evolve to evade treatments. Within in a clinical setting, the ability to sequence not just one, but many different cells across a patient’s tumor could point to the combination of treatments which offer the best chance of eradicating the entire cancer.
“What happens at the moment with a solid tumor is you treat with one drug, and maybe 80 percent of that tumor is susceptible to that drug,” says Neil Ward, vice president and general manager in the EMEA region for genomics company PacBio. “But the other 20 percent of the tumor has already got mutations that make it resistant, which is probably why a lot of modern therapies extend life for sadly only a matter of months rather than curing, because they treat a big percentage of the tumor, but not the whole thing. So going forwards, I think that we will see genomics play a huge role in cancer treatments, through using multiple modalities to treat someone's cancer.”
If insurers can figure out the economics, Snyder even foresees a future where at a certain age, all of us can qualify for annual sequencing of our blood cells to search for early signs of cancer or the potential onset of other diseases like type 2 diabetes.
“There are companies already working on looking for cancer signatures in methylated DNA,” he says. “If it was determined that you had early stage cancer, pre-symptomatically, that could then be validated with targeted MRI, followed by surgery or chemotherapy. It makes a big difference catching cancer early. If there were signs of type 2 diabetes, you could start taking steps to mitigate your glucose rise, and possibly prevent it or at least delay the onset.”
This would already revolutionize the way we seek to prevent a whole range of illnesses, but others feel that the $100 genome could also usher in even more powerful and controversial preventative medicine schemes.
Newborn screening
In the eyes of Kári Stefánsson, the Icelandic neurologist who been a visionary for so many advances in the field of human genetics over the last 25 years, the falling cost of sequencing means it will be feasible to sequence the genomes of every baby born.
“We have recently done an analysis of genomes in Iceland and the UK Biobank, and in 4 percent of people you find mutations that lead to serious disease, that can be prevented or dealt with,” says Stefansson, CEO of deCODE genetics, a subsidiary of the pharmaceutical company Amgen. “This could transform our healthcare systems.”
As well as identifying newborns with rare diseases, this kind of genomic information could be used to compute a person’s risk score for developing chronic illnesses later in life. If for example, they have a higher than average risk of colon or breast cancer, they could be pre-emptively scheduled for annual colonoscopies or mammograms as soon as they hit adulthood.
To a limited extent, this is already happening. In the UK, Genomics England has launched the Newborn Genomes Programme, which plans to undertake whole-genome sequencing of up to 200,000 newborn babies, with the aim of enabling the early identification of rare genetic diseases.
"I have not had my own genome sequenced and I would not have wanted my parents to have agreed to this," Curtis says. "I don’t see that sequencing children for the sake of some vague, ill-defined benefits could ever be justifiable.”
However, some scientists feel that it is tricky to justify sequencing the genomes of apparently healthy babies, given the data privacy issues involved. They point out that we still know too little about the links which can be drawn between genetic information at birth, and risk of chronic illness later in life.
“I think there are very difficult ethical issues involved in sequencing children if there are no clear and immediate clinical benefits,” says Curtis. “They cannot consent to this process. I have not had my own genome sequenced and I would not have wanted my parents to have agreed to this. I don’t see that sequencing children for the sake of some vague, ill-defined benefits could ever be justifiable.”
Curtis points out that there are many inherent risks about this data being available. It may fall into the hands of insurance companies, and it could even be used by governments for surveillance purposes.
“Genetic sequence data is very useful indeed for forensic purposes. Its full potential has yet to be realized but identifying rare variants could provide a quick and easy way to find relatives of a perpetrator,” he says. “If large numbers of people had been sequenced in a healthcare system then it could be difficult for a future government to resist the temptation to use this as a resource to investigate serious crimes.”
While sequencing becoming more widely available will present difficult ethical and moral challenges, it will offer many benefits for society as a whole. Cheaper sequencing will help boost the diversity of genomic datasets which have traditionally been skewed towards individuals of white, European descent, meaning that much of the actionable medical information which has come out of these studies is not relevant to people of other ethnicities.
Ward predicts that in the coming years, the growing amount of genetic information will ultimately change the outcomes for many with rare, previously incurable illnesses.
“If you're the parent of a child that has a susceptible or a suspected rare genetic disease, their genome will get sequenced, and while sadly that doesn’t always lead to treatments, it’s building up a knowledge base so companies can spring up and target that niche of a disease,” he says. “As a result there’s a whole tidal wave of new therapies that are going to come to market over the next five years, as the genetic tools we have, mature and evolve.”
This article was first published by Leaps.org in October 2022.
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
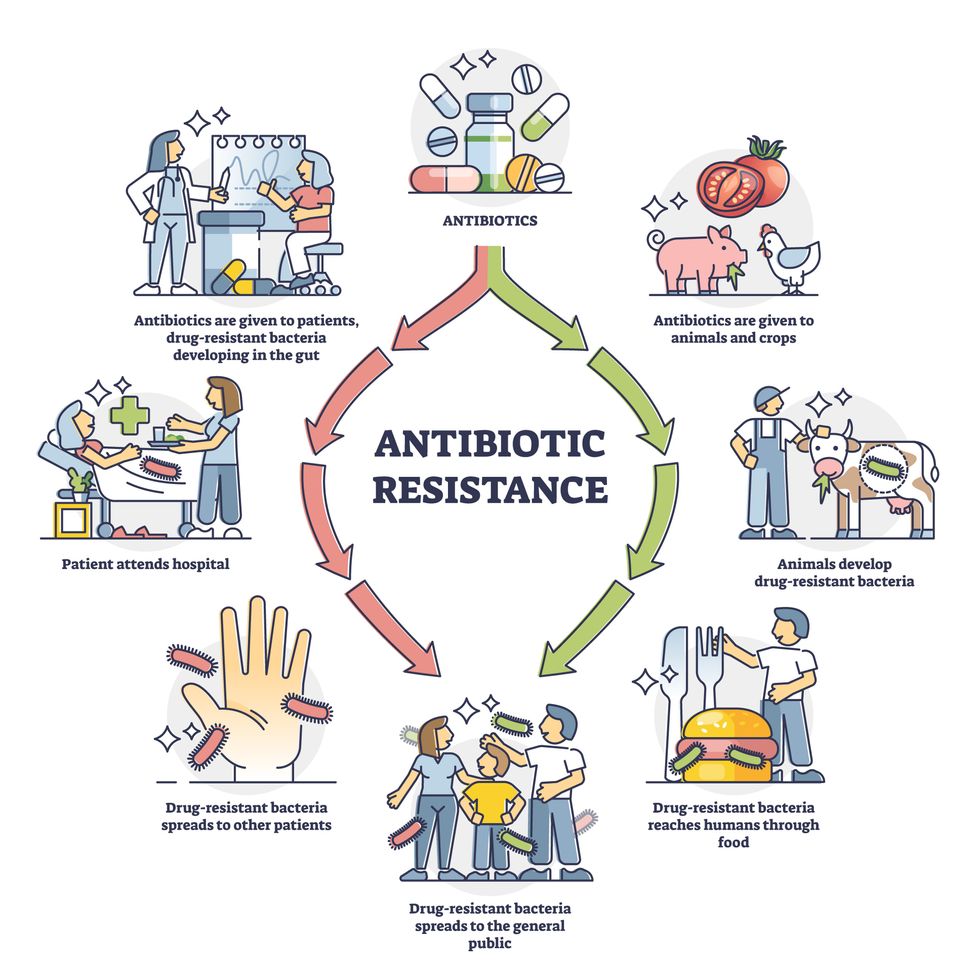
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
Meet Dr. Renee Wegrzyn, the first Director of President Biden's new health agency, ARPA-H
Today's podcast guest, Dr. Renee Wegrzyn, directs ARPA-H, a new agency formed last year to spearhead health innovations. Time will tell if ARPA-H will produce advances on the level of its fellow agency, DARPA.
In today’s podcast episode, I talk with Renee Wegrzyn, appointed by President Biden as the first director of a health agency created last year, the Advanced Research Projects Agency for Health, or ARPA-H. It’s inspired by DARPA, the agency that develops innovations for the Defense department and has been credited with hatching world-changing technologies such as ARPANET, which became the internet.
Time will tell if ARPA-H will lead to similar achievements in the realm of health. That’s what President Biden and Congress expect in return for funding ARPA-H at 2.5 billion dollars over three years.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
How will the agency figure out which projects to take on, especially with so many patient advocates for different diseases demanding moonshot funding for rapid progress?
I talked with Dr. Wegrzyn about the opportunities and challenges, what lessons ARPA-H is borrowing from Operation Warp Speed, how she decided on the first ARPA-H project that was announced recently, why a separate agency was needed instead of reforming HHS and the National Institutes of Health to be better at innovation, and how ARPA-H will make progress on disease prevention in addition to treatments for cancer, Alzheimer’s and diabetes, among many other health priorities.
Dr. Wegrzyn’s resume leaves no doubt of her suitability for this role. She was a program manager at DARPA where she focused on applying gene editing and synthetic biology to the goal of improving biosecurity. For her work there, she received the Superior Public Service Medal and, in case that wasn’t enough ARPA experience, she also worked at another ARPA that leads advanced projects in intelligence, called I-ARPA. Before that, she ran technical teams in the private sector working on gene therapies and disease diagnostics, among other areas. She has been a vice president of business development at Gingko Bioworks and headed innovation at Concentric by Gingko. Her training and education includes a PhD and undergraduate degree in applied biology from the Georgia Institute of Technology and she did her postdoc as an Alexander von Humboldt Fellow in Heidelberg, Germany.
Dr. Wegrzyn told me that she’s “in the hot seat.” The pressure is on for ARPA-H especially after the need and potential for health innovation was spot lit by the pandemic and the unprecedented speed of vaccine development. We'll soon find out if ARPA-H can produce gamechangers in health that are equivalent to DARPA’s creation of the internet.
Show links:
ARPA-H - https://arpa-h.gov/
Dr. Wegrzyn profile - https://arpa-h.gov/people/renee-wegrzyn/
Dr. Wegrzyn Twitter - https://twitter.com/rwegrzyn?lang=en
President Biden Announces Dr. Wegrzyn's appointment - https://www.whitehouse.gov/briefing-room/statement...
Leaps.org coverage of ARPA-H - https://leaps.org/arpa/
ARPA-H program for joints to heal themselves - https://arpa-h.gov/news/nitro/ -
ARPA-H virtual talent search - https://arpa-h.gov/news/aco-talent-search/

Dr. Renee Wegrzyn was appointed director of ARPA-H last October.