AI and you: Is the promise of personalized nutrition apps worth the hype?

Personalized nutrition apps could provide valuable data to people trying to eat healthier, though more research must be done to show effectiveness.
As a type 2 diabetic, Michael Snyder has long been interested in how blood sugar levels vary from one person to another in response to the same food, and whether a more personalized approach to nutrition could help tackle the rapidly cascading levels of diabetes and obesity in much of the western world.
Eight years ago, Snyder, who directs the Center for Genomics and Personalized Medicine at Stanford University, decided to put his theories to the test. In the 2000s continuous glucose monitoring, or CGM, had begun to revolutionize the lives of diabetics, both type 1 and type 2. Using spherical sensors which sit on the upper arm or abdomen – with tiny wires that pierce the skin – the technology allowed patients to gain real-time updates on their blood sugar levels, transmitted directly to their phone.
It gave Snyder an idea for his research at Stanford. Applying the same technology to a group of apparently healthy people, and looking for ‘spikes’ or sudden surges in blood sugar known as hyperglycemia, could provide a means of observing how their bodies reacted to an array of foods.
“We discovered that different foods spike people differently,” he says. “Some people spike to pasta, others to bread, others to bananas, and so on. It’s very personalized and our feeling was that building programs around these devices could be extremely powerful for better managing people’s glucose.”
Unbeknown to Snyder at the time, thousands of miles away, a group of Israeli scientists at the Weizmann Institute of Science were doing exactly the same experiments. In 2015, they published a landmark paper which used CGM to track the blood sugar levels of 800 people over several days, showing that the biological response to identical foods can vary wildly. Like Snyder, they theorized that giving people a greater understanding of their own glucose responses, so they spend more time in the normal range, may reduce the prevalence of type 2 diabetes.
The commercial potential of such apps is clear, but the underlying science continues to generate intriguing findings.
“At the moment 33 percent of the U.S. population is pre-diabetic, and 70 percent of those pre-diabetics will become diabetic,” says Snyder. “Those numbers are going up, so it’s pretty clear we need to do something about it.”
Fast forward to 2022,and both teams have converted their ideas into subscription-based dietary apps which use artificial intelligence to offer data-informed nutritional and lifestyle recommendations. Snyder’s spinoff, January AI, combines CGM information with heart rate, sleep, and activity data to advise on foods to avoid and the best times to exercise. DayTwo–a start-up which utilizes the findings of Weizmann Institute of Science–obtains microbiome information by sequencing stool samples, and combines this with blood glucose data to rate ‘good’ and ‘bad’ foods for a particular person.
“CGMs can be used to devise personalized diets,” says Eran Elinav, an immunology professor and microbiota researcher at the Weizmann Institute of Science in addition to serving as a scientific consultant for DayTwo. “However, this process can be cumbersome. Therefore, in our lab we created an algorithm, based on data acquired from a big cohort of people, which can accurately predict post-meal glucose responses on a personal basis.”
The commercial potential of such apps is clear. DayTwo, who market their product to corporate employers and health insurers rather than individual consumers, recently raised $37 million in funding. But the underlying science continues to generate intriguing findings.
Last year, Elinav and colleagues published a study on 225 individuals with pre-diabetes which found that they achieved better blood sugar control when they followed a personalized diet based on DayTwo’s recommendations, compared to a Mediterranean diet. The journal Cell just released a new paper from Snyder’s group which shows that different types of fibre benefit people in different ways.
“The idea is you hear different fibres are good for you,” says Snyder. “But if you look at fibres they’re all over the map—it’s like saying all animals are the same. The responses are very individual. For a lot of people [a type of fibre called] arabinoxylan clearly reduced cholesterol while the fibre inulin had no effect. But in some people, it was the complete opposite.”

Eight years ago, Stanford's Michael Snyder began studying how continuous glucose monitors could be used by patients to gain real-time updates on their blood sugar levels, transmitted directly to their phone.
The Snyder Lab, Stanford Medicine
Because of studies like these, interest in precision nutrition approaches has exploded in recent years. In January, the National Institutes of Health announced that they are spending $170 million on a five year, multi-center initiative which aims to develop algorithms based on a whole range of data sources from blood sugar to sleep, exercise, stress, microbiome and even genomic information which can help predict which diets are most suitable for a particular individual.
“There's so many different factors which influence what you put into your mouth but also what happens to different types of nutrients and how that ultimately affects your health, which means you can’t have a one-size-fits-all set of nutritional guidelines for everyone,” says Bruce Y. Lee, professor of health policy and management at the City University of New York Graduate School of Public Health.
With the falling costs of genomic sequencing, other precision nutrition clinical trials are choosing to look at whether our genomes alone can yield key information about what our diets should look like, an emerging field of research known as nutrigenomics.
The ASPIRE-DNA clinical trial at Imperial College London is aiming to see whether particular genetic variants can be used to classify individuals into two groups, those who are more glucose sensitive to fat and those who are more sensitive to carbohydrates. By following a tailored diet based on these sensitivities, the trial aims to see whether it can prevent people with pre-diabetes from developing the disease.
But while much hope is riding on these trials, even precision nutrition advocates caution that the field remains in the very earliest of stages. Lars-Oliver Klotz, professor of nutrigenomics at Friedrich-Schiller-University in Jena, Germany, says that while the overall goal is to identify means of avoiding nutrition-related diseases, genomic data alone is unlikely to be sufficient to prevent obesity and type 2 diabetes.
“Genome data is rather simple to acquire these days as sequencing techniques have dramatically advanced in recent years,” he says. “However, the predictive value of just genome sequencing is too low in the case of obesity and prediabetes.”
Others say that while genomic data can yield useful information in terms of how different people metabolize different types of fat and specific nutrients such as B vitamins, there is a need for more research before it can be utilized in an algorithm for making dietary recommendations.
“I think it’s a little early,” says Eileen Gibney, a professor at University College Dublin. “We’ve identified a limited number of gene-nutrient interactions so far, but we need more randomized control trials of people with different genetic profiles on the same diet, to see whether they respond differently, and if that can be explained by their genetic differences.”
Some start-ups have already come unstuck for promising too much, or pushing recommendations which are not based on scientifically rigorous trials. The world of precision nutrition apps was dubbed a ‘Wild West’ by some commentators after the founders of uBiome – a start-up which offered nutritional recommendations based on information obtained from sequencing stool samples –were charged with fraud last year. The weight-loss app Noom, which was valued at $3.7 billion in May 2021, has been criticized on Twitter by a number of users who claimed that its recommendations have led to them developed eating disorders.
With precision nutrition apps marketing their technology at healthy individuals, question marks have also been raised about the value which can be gained through non-diabetics monitoring their blood sugar through CGM. While some small studies have found that wearing a CGM can make overweight or obese individuals more motivated to exercise, there is still a lack of conclusive evidence showing that this translates to improved health.
However, independent researchers remain intrigued by the technology, and say that the wealth of data generated through such apps could be used to help further stratify the different types of people who become at risk of developing type 2 diabetes.
“CGM not only enables a longer sampling time for capturing glucose levels, but will also capture lifestyle factors,” says Robert Wagner, a diabetes researcher at University Hospital Düsseldorf. “It is probable that it can be used to identify many clusters of prediabetic metabolism and predict the risk of diabetes and its complications, but maybe also specific cardiometabolic risk constellations. However, we still don’t know which forms of diabetes can be prevented by such approaches and how feasible and long-lasting such self-feedback dietary modifications are.”
Snyder himself has now been wearing a CGM for eight years, and he credits the insights it provides with helping him to manage his own diabetes. “My CGM still gives me novel insights into what foods and behaviors affect my glucose levels,” he says.
He is now looking to run clinical trials with his group at Stanford to see whether following a precision nutrition approach based on CGM and microbiome data, combined with other health information, can be used to reverse signs of pre-diabetes. If it proves successful, January AI may look to incorporate microbiome data in future.
“Ultimately, what I want to do is be able take people’s poop samples, maybe a blood draw, and say, ‘Alright, based on these parameters, this is what I think is going to spike you,’ and then have a CGM to test that out,” he says. “Getting very predictive about this, so right from the get go, you can have people better manage their health and then use the glucose monitor to help follow that.”
Nobel Prize goes to technology for mRNA vaccines
Katalin Karikó, pictured, and Drew Weissman won the Nobel Prize for advances in mRNA research that led to the first Covid vaccines.
When Drew Weissman received a call from Katalin Karikó in the early morning hours this past Monday, he assumed his longtime research partner was calling to share a nascent, nagging idea. Weissman, a professor of medicine at the Perelman School of Medicine at the University of Pennsylvania, and Karikó, a professor at Szeged University and an adjunct professor at UPenn, both struggle with sleep disturbances. Thus, middle-of-the-night discourses between the two, often over email, has been a staple of their friendship. But this time, Karikó had something more pressing and exciting to share: They had won the 2023 Nobel Prize in Physiology or Medicine.
The work for which they garnered the illustrious award and its accompanying $1,000,000 cash windfall was completed about two decades ago, wrought through long hours in the lab over many arduous years. But humanity collectively benefited from its life-saving outcome three years ago, when both Moderna and Pfizer/BioNTech’s mRNA vaccines against COVID were found to be safe and highly effective at preventing severe disease. Billions of doses have since been given out to protect humans from the upstart viral scourge.
“I thought of going somewhere else, or doing something else,” said Katalin Karikó. “I also thought maybe I’m not good enough, not smart enough. I tried to imagine: Everything is here, and I just have to do better experiments.”
Unlocking the power of mRNA
Weissman and Karikó unlocked mRNA vaccines for the world back in the early 2000s when they made a key breakthrough. Messenger RNA molecules are essentially instructions for cells’ ribosomes to make specific proteins, so in the 1980s and 1990s, researchers started wondering if sneaking mRNA into the body could trigger cells to manufacture antibodies, enzymes, or growth agents for protecting against infection, treating disease, or repairing tissues. But there was a big problem: injecting this synthetic mRNA triggered a dangerous, inflammatory immune response resulting in the mRNA’s destruction.
While most other researchers chose not to tackle this perplexing problem to instead pursue more lucrative and publishable exploits, Karikó stuck with it. The choice sent her academic career into depressing doldrums. Nobody would fund her work, publications dried up, and after six years as an assistant professor at the University of Pennsylvania, Karikó got demoted. She was going backward.
“I thought of going somewhere else, or doing something else,” Karikó told Stat in 2020. “I also thought maybe I’m not good enough, not smart enough. I tried to imagine: Everything is here, and I just have to do better experiments.”
A tale of tenacity
Collaborating with Drew Weissman, a new professor at the University of Pennsylvania, in the late 1990s helped provide Karikó with the tenacity to continue. Weissman nurtured a goal of developing a vaccine against HIV-1, and saw mRNA as a potential way to do it.
“For the 20 years that we’ve worked together before anybody knew what RNA is, or cared, it was the two of us literally side by side at a bench working together,” Weissman said in an interview with Adam Smith of the Nobel Foundation.
In 2005, the duo made their 2023 Nobel Prize-winning breakthrough, detailing it in a relatively small journal, Immunity. (Their paper was rejected by larger journals, including Science and Nature.) They figured out that chemically modifying the nucleoside bases that make up mRNA allowed the molecule to slip past the body’s immune defenses. Karikó and Weissman followed up that finding by creating mRNA that’s more efficiently translated within cells, greatly boosting protein production. In 2020, scientists at Moderna and BioNTech (where Karikó worked from 2013 to 2022) rushed to craft vaccines against COVID, putting their methods to life-saving use.
The future of vaccines
Buoyed by the resounding success of mRNA vaccines, scientists are now hurriedly researching ways to use mRNA medicine against other infectious diseases, cancer, and genetic disorders. The now ubiquitous efforts stand in stark contrast to Karikó and Weissman’s previously unheralded struggles years ago as they doggedly worked to realize a shared dream that so many others shied away from. Katalin Karikó and Drew Weissman were brave enough to walk a scientific path that very well could have ended in a dead end, and for that, they absolutely deserve their 2023 Nobel Prize.
This article originally appeared on Big Think, home of the brightest minds and biggest ideas of all time.

Scientists turn pee into power in Uganda
With conventional fuel cells as their model, researchers learned to use similar chemical reactions to make a fuel from microbes in pee.
At the edge of a dirt road flanked by trees and green mountains outside the town of Kisoro, Uganda, sits the concrete building that houses Sesame Girls School, where girls aged 11 to 19 can live, learn and, at least for a while, safely use a toilet. In many developing regions, toileting at night is especially dangerous for children. Without electrical power for lighting, kids may fall into the deep pits of the latrines through broken or unsteady floorboards. Girls are sometimes assaulted by men who hide in the dark.
For the Sesame School girls, though, bright LED lights, connected to tiny gadgets, chased the fears away. They got to use new, clean toilets lit by the power of their own pee. Some girls even used the light provided by the latrines to study.
Urine, whether animal or human, is more than waste. It’s a cheap and abundant resource. Each day across the globe, 8.1 billion humans make 4 billion gallons of pee. Cows, pigs, deer, elephants and other animals add more. By spending money to get rid of it, we waste a renewable resource that can serve more than one purpose. Microorganisms that feed on nutrients in urine can be used in a microbial fuel cell that generates electricity – or "pee power," as the Sesame girls called it.
Plus, urine contains water, phosphorus, potassium and nitrogen, the key ingredients plants need to grow and survive. Human urine could replace about 25 percent of current nitrogen and phosphorous fertilizers worldwide and could save water for gardens and crops. The average U.S. resident flushes a toilet bowl containing only pee and paper about six to seven times a day, which adds up to about 3,500 gallons of water down per year. Plus cows in the U.S. produce 231 gallons of the stuff each year.
Pee power
A conventional fuel cell uses chemical reactions to produce energy, as electrons move from one electrode to another to power a lightbulb or phone. Ioannis Ieropoulos, a professor and chair of Environmental Engineering at the University of Southampton in England, realized the same type of reaction could be used to make a fuel from microbes in pee.
Bacterial species like Shewanella oneidensis and Pseudomonas aeruginosa can consume carbon and other nutrients in urine and pop out electrons as a result of their digestion. In a microbial fuel cell, one electrode is covered in microbes, immersed in urine and kept away from oxygen. Another electrode is in contact with oxygen. When the microbes feed on nutrients, they produce the electrons that flow through the circuit from one electrod to another to combine with oxygen on the other side. As long as the microbes have fresh pee to chomp on, electrons keep flowing. And after the microbes are done with the pee, it can be used as fertilizer.
These microbes are easily found in wastewater treatment plants, ponds, lakes, rivers or soil. Keeping them alive is the easy part, says Ieropoulos. Once the cells start producing stable power, his group sequences the microbes and keeps using them.
Like many promising technologies, scaling these devices for mass consumption won’t be easy, says Kevin Orner, a civil engineering professor at West Virginia University. But it’s moving in the right direction. Ieropoulos’s device has shrunk from the size of about three packs of cards to a large glue stick. It looks and works much like a AAA battery and produce about the same power. By itself, the device can barely power a light bulb, but when stacked together, they can do much more—just like photovoltaic cells in solar panels. His lab has produced 1760 fuel cells stacked together, and with manufacturing support, there’s no theoretical ceiling, he says.
Although pure urine produces the most power, Ieropoulos’s devices also work with the mixed liquids of the wastewater treatment plants, so they can be retrofit into urban wastewater utilities.
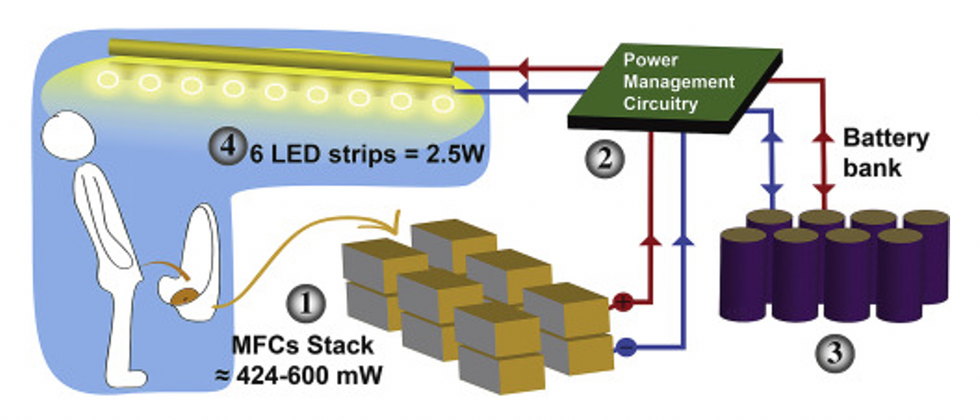
This image shows how the pee-powered system works. Pee feeds bacteria in the stack of fuel cells (1), which give off electrons (2) stored in parallel cylindrical cells (3). These cells are connected to a voltage regulator (4), which smooths out the electrical signal to ensure consistent power to the LED strips lighting the toilet.
Courtesy Ioannis Ieropoulos
Key to the long-term success of any urine reclamation effort, says Orner, is avoiding what he calls “parachute engineering”—when well-meaning scientists solve a problem with novel tech and then abandon it. “The way around that is to have either the need come from the community or to have an organization in a community that is committed to seeing a project operate and maintained,” he says.
Success with urine reclamation also depends on the economy. “If energy prices are low, it may not make sense to recover energy,” says Orner. “But right now, fertilizer prices worldwide are generally pretty high, so it may make sense to recover fertilizer and nutrients.” There are obstacles, too, such as few incentives for builders to incorporate urine recycling into new construction. And any hiccups like leaks or waste seepage will cost builders money and reputation. Right now, Orner says, the risks are just too high.
Despite the challenges, Ieropoulos envisions a future in which urine is passed through microbial fuel cells at wastewater treatment plants, retrofitted septic tanks, and building basements, and is then delivered to businesses to use as agricultural fertilizers. Although pure urine produces the most power, Ieropoulos’s devices also work with the mixed liquids of the wastewater treatment plants, so they can be retrofitted into urban wastewater utilities where they can make electricity from the effluent. And unlike solar cells, which are a common target of theft in some areas, nobody wants to steal a bunch of pee.
When Ieropoulos’s team returned to wrap up their pilot project 18 months later, the school’s director begged them to leave the fuel cells in place—because they made a major difference in students’ lives. “We replaced it with a substantial photovoltaic panel,” says Ieropoulos, They couldn’t leave the units forever, he explained, because of intellectual property reasons—their funders worried about theft of both the technology and the idea. But the photovoltaic replacement could be stolen, too, leaving the girls in the dark.
The story repeated itself at another school, in Nairobi, Kenya, as well as in an informal settlement in Durban, South Africa. Each time, Ieropoulos vowed to return. Though the pandemic has delayed his promise, he is resolute about continuing his work—it is a moral and legal obligation. “We've made a commitment to ourselves and to the pupils,” he says. “That's why we need to go back.”
Urine as fertilizer
Modern day industrial systems perpetuate the broken cycle of nutrients. When plants grow, they use up nutrients the soil. We eat the plans and excrete some of the nutrients we pass them into rivers and oceans. As a result, farmers must keep fertilizing the fields while our waste keeps fertilizing the waterways, where the algae, overfertilized with nitrogen, phosphorous and other nutrients grows out of control, sucking up oxygen that other marine species need to live. Few global communities remain untouched by the related challenges this broken chain create: insufficient clean water, food, and energy, and too much human and animal waste.
The Rich Earth Institute in Vermont runs a community-wide urine nutrient recovery program, which collects urine from homes and businesses, transports it for processing, and then supplies it as fertilizer to local farms.
One solution to this broken cycle is reclaiming urine and returning it back to the land. The Rich Earth Institute in Vermont is one of several organizations around the world working to divert and save urine for agricultural use. “The urine produced by an adult in one day contains enough fertilizer to grow all the wheat in one loaf of bread,” states their website.
Notably, while urine is not entirely sterile, it tends to harbor fewer pathogens than feces. That’s largely because urine has less organic matter and therefore less food for pathogens to feed on, but also because the urinary tract and the bladder have built-in antimicrobial defenses that kill many germs. In fact, the Rich Earth Institute says it’s safe to put your own urine onto crops grown for home consumption. Nonetheless, you’ll want to dilute it first because pee usually has too much nitrogen and can cause “fertilizer burn” if applied straight without dilution. Other projects to turn urine into fertilizer are in progress in Niger, South Africa, Kenya, Ethiopia, Sweden, Switzerland, The Netherlands, Australia, and France.
Eleven years ago, the Institute started a program that collects urine from homes and businesses, transports it for processing, and then supplies it as fertilizer to local farms. By 2021, the program included 180 donors producing over 12,000 gallons of urine each year. This urine is helping to fertilize hay fields at four partnering farms. Orner, the West Virginia professor, sees it as a success story. “They've shown how you can do this right--implementing it at a community level scale."

