Is Finding Out Your Baby’s Genetics A New Responsibility of Parenting?
A doctor pricks the heel of a newborn for a blood test.
Hours after a baby is born, its heel is pricked with a lancet. Drops of the infant's blood are collected on a porous card, which is then mailed to a state laboratory. The dried blood spots are screened for around thirty conditions, including phenylketonuria (PKU), the metabolic disorder that kick-started this kind of newborn screening over 60 years ago. In the U.S., parents are not asked for permission to screen their child. Newborn screening programs are public health programs, and the assumption is that no good parent would refuse a screening test that could identify a serious yet treatable condition in their baby.
Learning as much as you can about your child's health might seem like a natural obligation of parenting. But it's an assumption that I think needs to be much more closely examined.
Today, with the introduction of genome sequencing into clinical medicine, some are asking whether newborn screening goes far enough. As the cost of sequencing falls, should parents take a more expansive look at their children's health, learning not just whether they have a rare but treatable childhood condition, but also whether they are at risk for untreatable conditions or for diseases that, if they occur at all, will strike only in adulthood? Should genome sequencing be a part of every newborn's care?
It's an idea that appeals to Anne Wojcicki, the founder and CEO of the direct-to-consumer genetic testing company 23andMe, who in a 2016 interview with The Guardian newspaper predicted that having newborns tested would soon be considered standard practice—"as critical as testing your cholesterol"—and a new responsibility of parenting. Wojcicki isn't the only one excited to see everyone's genes examined at birth. Francis Collins, director of the National Institutes of Health and perhaps the most prominent advocate of genomics in the United States, has written that he is "almost certain … that whole-genome sequencing will become part of new-born screening in the next few years." Whether that would happen through state-mandated screening programs, or as part of routine pediatric care—or perhaps as a direct-to-consumer service that parents purchase at birth or receive as a baby-shower gift—is not clear.
Learning as much as you can about your child's health might seem like a natural obligation of parenting. But it's an assumption that I think needs to be much more closely examined, both because the results that genome sequencing can return are more complex and more uncertain than one might expect, and because parents are not actually responsible for their child's lifelong health and well-being.
What is a parent supposed to do about such a risk except worry?
Existing newborn screening tests look for the presence of rare conditions that, if identified early in life, before the child shows any symptoms, can be effectively treated. Sequencing could identify many of these same kinds of conditions (and it might be a good tool if it could be targeted to those conditions alone), but it would also identify gene variants that confer an increased risk rather than a certainty of disease. Occasionally that increased risk will be significant. About 12 percent of women in the general population will develop breast cancer during their lives, while those who have a harmful BRCA1 or BRCA2 gene variant have around a 70 percent chance of developing the disease. But for many—perhaps most—conditions, the increased risk associated with a particular gene variant will be very small. Researchers have identified over 600 genes that appear to be associated with schizophrenia, for example, but any one of those confers only a tiny increase in risk for the disorder. What is a parent supposed to do about such a risk except worry?
Sequencing results are uncertain in other important ways as well. While we now have the ability to map the genome—to create a read-out of the pairs of genetic letters that make up a person's DNA—we are still learning what most of it means for a person's health and well-being. Researchers even have a name for gene variants they think might be associated with a disease or disorder, but for which they don't have enough evidence to be sure. They are called "variants of unknown (or uncertain) significance (VUS), and they pop up in most people's sequencing results. In cancer genetics, where much research has been done, about 1 in 5 gene variants are reclassified over time. Most are downgraded, which means that a good number of VUS are eventually designated benign.
While one parent might reasonably decide to learn about their child's risk for a condition about which nothing can be done medically, a different, yet still thoroughly reasonable, parent might prefer to remain ignorant so that they can enjoy the time before their child is afflicted.
Then there's the puzzle of what to do about results that show increased risk or even certainty for a condition that we have no idea how to prevent. Some genomics advocates argue that even if a result is not "medically actionable," it might have "personal utility" because it allows parents to plan for their child's future needs, to enroll them in research, or to connect with other families whose children carry the same genetic marker.
Finding a certain gene variant in one child might inform parents' decisions about whether to have another—and if they do, about whether to use reproductive technologies or prenatal testing to select against that variant in a future child. I have no doubt that for some parents these personal utility arguments are persuasive, but notice how far we've now strayed from the serious yet treatable conditions that motivated governments to set up newborn screening programs, and to mandate such testing for all.
Which brings me to the other problem with the call for sequencing newborn babies: the idea that even if it's not what the law requires, it's what good parents should do. That idea is very compelling when we're talking about sequencing results that show a serious threat to the child's health, especially when interventions are available to prevent or treat that condition. But as I have shown, many sequencing results are not of this type.
While one parent might reasonably decide to learn about their child's risk for a condition about which nothing can be done medically, a different, yet still thoroughly reasonable, parent might prefer to remain ignorant so that they can enjoy the time before their child is afflicted. This parent might decide that the worry—and the hypervigilence it could inspire in them—is not in their child's best interest, or indeed in their own. This parent might also think that it should be up to the child, when he or she is older, to decide whether to learn about his or her risk for adult-onset conditions, especially given that many adults at high familial risk for conditions like Alzheimer's or Huntington's disease choose never to be tested. This parent will value the child's future autonomy and right not to know more than they value the chance to prepare for a health risk that won't strike the child until 40 or 50 years in the future.
Parents are not obligated to learn about their children's risk for a condition that cannot be prevented, has a small risk of occurring, or that would appear only in adulthood.
Contemporary understandings of parenting are famously demanding. We are asked to do everything within our power to advance our children's health and well-being—to act always in our children's best interests. Against that backdrop, the need to sequence every newborn baby's genome might seem obvious. But we should be skeptical. Many sequencing results are complex and uncertain. Parents are not obligated to learn about their children's risk for a condition that cannot be prevented, has a small risk of occurring, or that would appear only in adulthood. To suggest otherwise is to stretch parental responsibilities beyond the realm of childhood and beyond factors that parents can control.
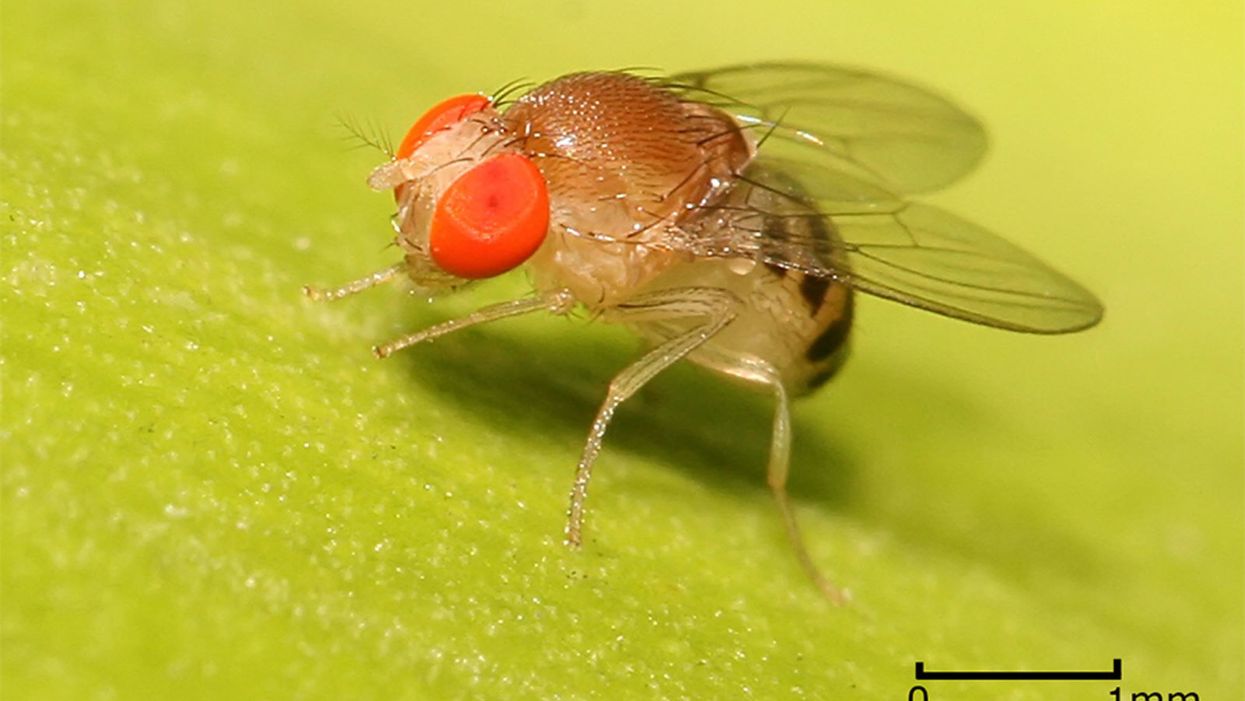
Scientists Used Fruit Flies to Quickly Develop a Personalized Cancer Treatment for a Dying Man
Genetically engineered fruit flies are being used in early research to screen novel combinations of cancer drugs for individual patients.
Imagine a man with colorectal cancer that has spread throughout his body. His tumor is not responding to traditional chemotherapy. He needs a radically effective treatment as soon as possible and there's no time to wait for a new drug or a new clinical trial.
A plethora of novel combinations of treatments can be screened quickly on as many as 400,000 flies at once.
This was the very real, and terrifying, situation of a recent patient at Mount Sinai Medical Center in New York City. So his doctors turned to a new tactic to speed up the search for a treatment that would save him: Fruit flies.
Yes, fruit flies. Those annoying little buggers that descend on opened food containers are actually leading scientists to fully personalized cancer treatments. Oncology advances often are more about about utilizing old drugs in new combinations than about adding new drugs. But classically, the development of each new chemotherapy drug combination has required studies involving numerous patients spread over many years or decades.
With the fruit fly method, however, a novel treatment -- in the sense that a particular combination of drugs and the timing of their administration has never been used before -- is developed for each patient, almost like on Star Trek, when, faced suddenly with an unknown disease, a futuristic physician researches it and develops a cure quickly enough to save the patient's life.
How It Works
Using genetic engineering techniques, researchers produce a population of fruit fly embryos, each of which is programmed to develop a replica of the patient's cancer.
Since a lot of genetically identical fly embryos can be created, and since they hatch from eggs within 30 hours and then mature within days, a plethora of novel combinations of treatments can be screened quickly on as many as 400,000 flies at once. Then, only the regimens that are effective are administered to the patient.
Biotech entrepreneur Laura Towart, CEO of the UK- and Ireland-based company, My Personal Therapeutics, is partnering with Mount Sinai to develop and test the fruit fly tactic. The researchers recently published a paper demonstrating that the tumor of the man with metastatic colorectal cancer had shrunk considerably following the treatment, and remained stable for 11 months, although he eventually succumbed to his illness.
Open Questions
Cancer is in fact many different diseases, even if it strikes two people in the same place, and both cancers look the same under a microscope. At the level of DNA, RNA, proteins, and other molecular factors, each cancer is unique – and may require a unique treatment approach.
Determining the true impact on cancer mortality will require clinical trials involving many more patients.
"Anatomy of a cancer still plays a major role, if you're a surgeon or radiation oncologist, but the medical approach to cancer therapy is moving toward treatments that are personalized based on other factors," notes Dr. Howard McLeod, an internationally recognized expert on cancer genetics at the Moffitt Cancer Center, in Tampa, Florida. "We are also headed into an era when even the methods for monitoring patients are individualized."
One big unresolved question about the fruit fly screening approach is how effective it will be in terms of actually extending life. Determining the true impact on cancer mortality will require clinical trials involving many more patients.
Next Up
Using machine learning and artificial intelligence, Towart is now working to build a service called TuMatch that will offer rapid and affordable personalized treatment recommendations for all genetically driven cancers. "We hope to have TuMatch available to patients with colorectal/GI cancers by January 2020," she says. "We are also offering [the fruit fly approach] for patients with rare genetic diseases and for patients who are diabetic."
Are Towart's fruit flies the answer to why the man's tumor shrunk? To be sure, the definitive answer will come from further research that is expected soon, but it's also clear that, prior to the treatment, there was nothing left to do for that particular patient. Thus, although it's early in the game, there's a pretty good rationale for optimism.
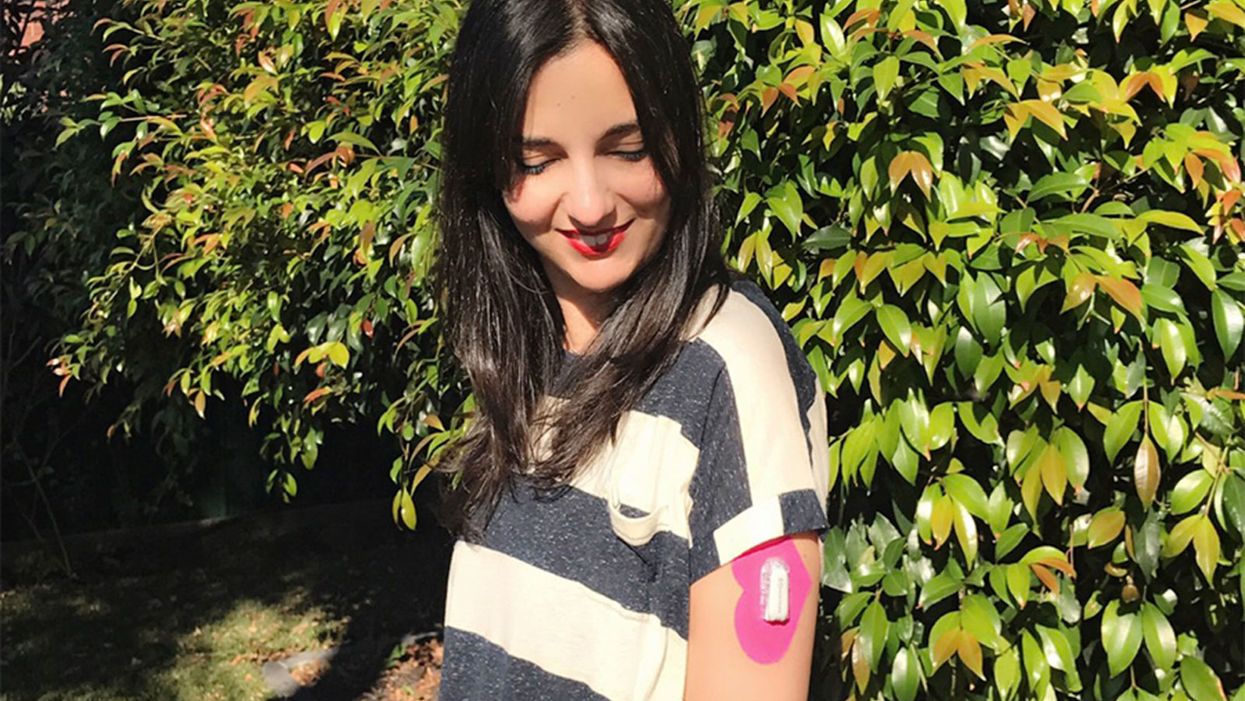
A Million Patients Have Innovated Their Own Medical Solutions, And Doctors Are Terrified
Diabetes patient advocate Renza Scibilia with her continuous glucose monitor, used in innovative DIY health technology.
In the fall of 2017, patient advocate Renza Scibilia told a conference of endocrinologists in Australia about new, patient-developed artificial pancreas technology that helped her manage her Type 1 diabetes.
"Because it's not a regulated product, some [doctors] were worried and said 'What if it goes wrong?'"
"They were in equal measure really interested and really scared," recalled Scibilia. "Because it's not a regulated product, some were worried and said 'What if it goes wrong? What is my liability going to be?'"
That was two years ago. Asked if physicians have been more receptive to the same "looping" technology now that its benefits have been supported by considerable data (as Leapsmag pointed out in May), Scibilia said, "No. Clinicians are still really insecure. They're always going to be reluctant to accept consumer-driven technology."
This exemplifies a major challenge to the growing Do-It-Yourself (DIY) biohealth movement: physicians are unnerved and worried about innovations developed by patients and other consumers that haven't been tested in elaborate clinical trials or sanctioned by regulatory authorities.
"It's difficult for patients who develop new health technology to demonstrate the advantage in a way that physicians would accept." said Howard DeMonaco, visiting scientist at MIT's Sloan School of Management. "New approaches to the treatment of diseases are by definition suspect to clinicians. Most are risk averse unless there is a substantial advantage to the new approach and the risks in doing so appear to be minimized."
Nevertheless, the DIY biohealth movement is booming. About a million people reported that they created medical innovations to address their own medical needs in surveys conducted from 2010-2015 in the U.S., U.K., Finland, Canada and South Korea.
Add in other DIY health innovations created in homes, community biolabs and "Maker" health fairs, and it's clear that health care providers are increasingly confronted with medical devices, information technology, and even medications that were developed in unconventional settings and lack the blessing of regulatory authorities.
Researchers in Portugal have tried to spread the word about many of these solutions on the Patent Innovations website, which has more than 500 examples, ranging from a 3-D printed arm and hand to a sensor device that warns someone when an osteomy bag is full.
When Reddit asked medical professionals, "What is the craziest DIY health treatment you've seen a patient attempt?" thousands shared horror stories.
But even in this era of patient empowerment, more widespread use of DIY health solutions still depends upon the approval and cooperation of physicians, nurses and other caregivers. And health care providers still lack awareness of promising patient-developed innovations, according to Dr. Joyce Lee, a pediatric endocrinologist at the University of Michigan who advocates involving patients in the design of healthcare technology. "Most physicians are scared of what they don't know," she said.
They're also understandably worried about patients who don't know what they're doing and make irresponsible decisions. When Reddit asked medical professionals, "What is the craziest DIY health treatment you've seen a patient attempt?" thousands shared horror stories, including a man who poked a hole in his belly button with a knitting needle to relieve gas.
Yet DeMonaco and Lee think it's possible to start bridging the gaps between responsible patient innovators and skeptical doctors as well as unprepared regulatory systems.
One obstacle to consumer-driven health innovations is that clinical trials to prove their safety and effectiveness are expensive and time-consuming, as De Monaco points out in a recent article. He and his colleagues suggested that low-cost clinical trials by and for patients could help address this challenge. They urged patients to publish their own research and detail the impact of innovations on their own health, and create databases that incorporate the findings of other patients.
For example, Adam Brown, who has Type 1 diabetes, compared the effects of low and high carbohydrate diets on his blood sugar management, and conveyed the results in an online journal. "Sharing the information allowed others to copy the experiment," the article noted, suggesting that this could be a model to create multi-patient trials that could be "analyzed by expert patients and/or by professionals."
Asked how to convince health care providers to consider such research, DeMonaco cited the example of doctors prescribing "off label" drugs for purposes that aren't approved by the FDA. "The secret to off label use, like any other user innovation, is dissemination," he said. Sharing case reports and other low-cost research serves to disseminate the information "in a way that is comfortable for physicians," he said, and urged patient innovators to take the same approach.
The FDA regulates commercial products and has no authority if consumers want to use medical devices, medications, or information systems that they find on their own.
Physicians should also be encouraged to engage in patient-driven research, said Dr. Lee. She suggests forming "maker spaces in which patients and physicians are involved in designing personalized technology for chronic diseases. In my vision, patient peers would build, iterate, and learn from each other and the doctor would be part of the team, constantly assessing and evaluating the technology and facilitating the process."
Some kind of regulatory oversight of DIY health technology is also necessary, said Todd Kuiken, senior research scholar at NC State and former principal investigator at the Woodrow Wilson Center's Synthetic Biology Project.
The FDA regulates commercial products and has no authority if consumers want to use medical devices, medications, or information systems that they find on their own. But that doesn't stop regulators from worrying about patients who use them. For example, the FDA issued a warning about diabetes looping technology earlier this year after one diabetic was hospitalized with hypoglycemia.
Kuiken, for one, believes that citizen-driven innovation requires oversight "to move forward." He suggested that Internal Review Boards, with experts on medical technology, safety and ethics, could play a helpful role in validating the work of patient innovators and others engaged in DIY health research. "As people are developing health products, there would be experts available to take a look and check in," he said.
Kuiken pointed out that in native American territories, tribally based IRBs working with the national Indian Health Services help to oversee new health science research. The model could be applied more broadly.
He also offered hope to those who want to integrate the current health regulatory structure into the ecosystem of DIY health innovations. "I didn't expect people from the FDA or NIH to show up" he said about a workshop on citizen-driven biomedical research that he helped organize at the Wilson Center last year. But senior officials from both agencies attended.
He indicated they "were open to new ideas." While he wouldn't disclose contributions made by individual participants in the workshop, he said the government staffers were "very interested in figuring out how to engage with citizen health innovators, to build bridges with the DIY community."
"Why should we wait for regulatory bodies? Why wait for trials that take too long?"
Time will tell whether those bridges will be built quickly enough to increase the comfort of physicians with health innovations developed by patients and other consumers. In the meantime, DIY health innovators like patient advocate Scibilia are undeterred.
"Why should we wait for regulatory bodies?" she asked. "Why wait for trials that take too long? There are plenty of data out there indicating the [diabetes looping] technology works. So we're just going to do it. We're not waiting."

