New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
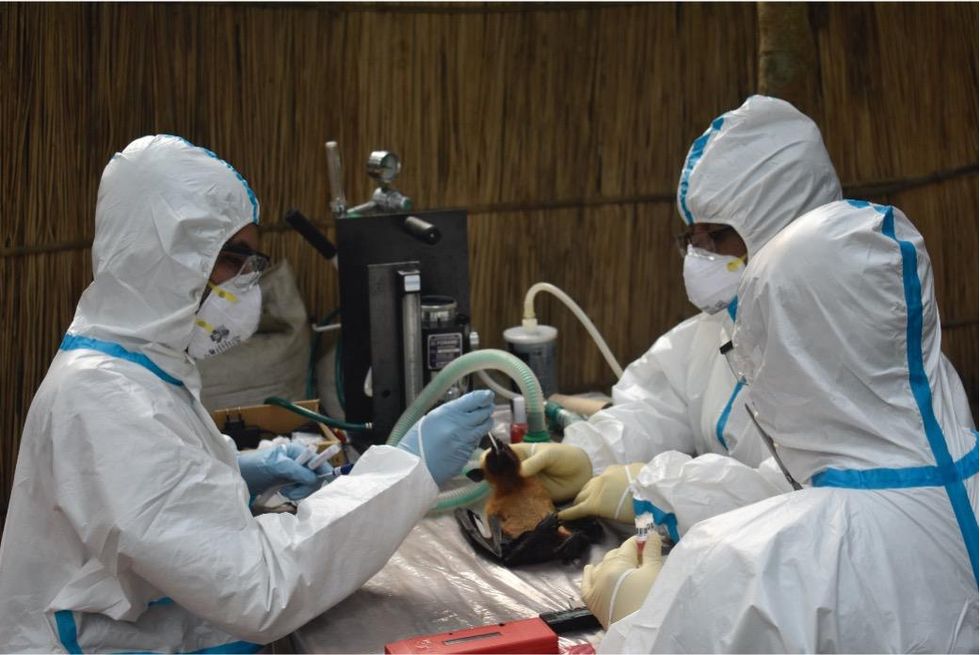
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
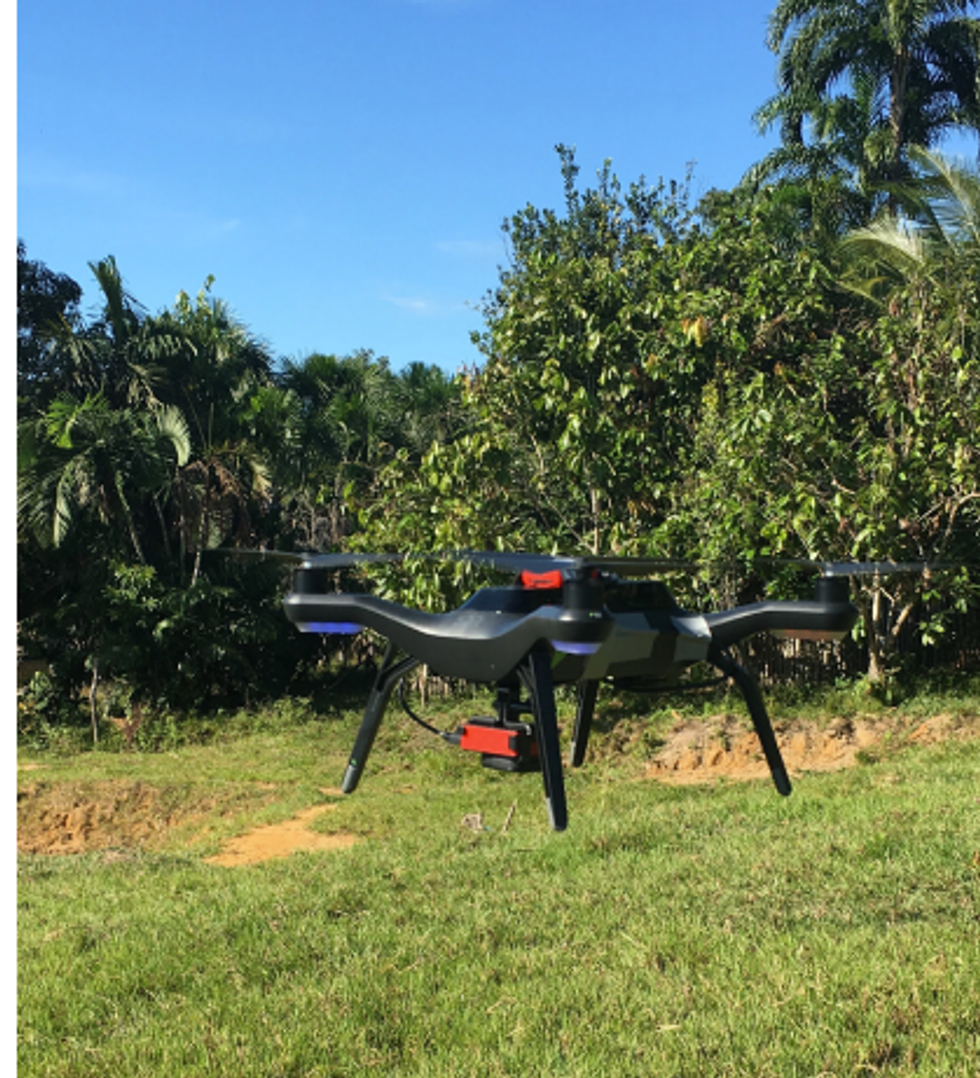

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
When Willem Johan Kolff invented dialysis, the "father" of artificial organs was just getting started.
One of the Netherlands’ most famous pieces of pop culture is “Soldier of Orange.” It’s the title of the country’s most celebrated war memoir, movie and epic stage musical, all of which detail the exploits of the nation’s resistance fighters during World War II.
Willem Johan Kolff was a member of the Dutch resistance, but he doesn’t rate a mention in the “Solider of Orange” canon. Yet his wartime toils in a rural backwater not only changed medicine, but the world.
Kolff had been a physician less than two years before Germany invaded the Netherlands in May 1940. He had been engaged in post-graduate studies at the University of Gronigen but withdrew because he refused to accommodate the demands of the Nazi occupiers. Kolff’s Jewish supervisor made an even starker choice: He committed suicide.
After his departure from the university, Kolff took a job managing a small hospital in Kampen. Located 50 miles from the heavily populated coastal region, the facility was far enough away from the prying eyes of Germans that not only could Kolff care for patients, he could hide fellow resistance fighters and even Jewish refugees in relative safety. Kolff coached many of them to feign convincing terminal illnesses so the Nazis would allow them to remain in the hospital.
Despite the demands of practicing medicine and resistance work, Kolff still found time to conduct research. He had been haunted and inspired when, not long before the Nazi invasion, one of his patients died in agony from kidney disease. Kolff wanted to find a way to save future patients.
He broke his problem down to a simple task: If he could remove 20 grams of urea from a patient’s blood in 24 hours, they would survive. He began experimenting with ways to filter blood and return it to a patient’s body. Since the war had ground all non-military manufacturing to a halt, he was mostly forced to make do with material he could find at the hospital and around Kampen. Kolff eventually built a device from a washing machine parts, juice cans, sausage casings, a valve from an old Ford automobile radiator, and even scrap from a downed German aircraft.
The world’s first dialysis machine was hardly imposing; it resembled a rotating drum for a bingo game or raffle. Yet it carried on the highly sophisticated task of moving a patient’s blood through a semi-permeable membrane (about a 50-foot length of sausage casings) into a saline solution that drew out urea while leaving the blood cells untouched.
In emigrating to the U.S. to practice medicine, Kolff's intent was twofold: Advocate for a wider adoption of dialysis, and work on new projects. He wildly succeeded at both.
Kolff began using the machine to treat patients in 1943, most of whom had lapsed into comas due to their kidney failure. But like most groundbreaking medical devices, it was not an immediate success. By the end of the war, Kolff had dialyzed more than a dozen patients, but all had died. He briefly suspended use of the device after the Allied invasion of Europe, but he continued to refine its operation and the administration of blood thinners to patients.
In September 1945, Kolff dialyzed another comatose patient, 67-year-old Sofia Maria Schafstadt. She regained consciousness after 11 hours, and would live well into the 1950s with Kolff’s assistance. Yet this triumph contained a dark irony: At the time of her treatment, Schafstadt had been imprisoned for collaborating with the Germans.
With a tattered Europe struggling to overcome the destruction of the war, Kolff and his family emigrated to the U.S. in 1950, where he began working for the Cleveland Clinic while undergoing the naturalization process so he could practice medicine in the U.S. His intent was twofold: Advocate for a wider adoption of dialysis, and work on new projects. He wildly succeeded at both.
By the mid-1950s, dialysis machines had become reliable and life-saving medical devices, and Kolff had become a U.S. citizen. About that time he invented a membrane oxygenator that could be used in heart bypass surgeries. This was a critical component of the heart-lung machine, which would make heart transplants possible and bypass surgeries routine. He also invented among the very first practical artificial hearts, which in 1957 kept a dog alive for 90 minutes.
Kolff moved to the University of Utah in 1967 to become director of its Institute for Biomedical Engineering. It was a promising time for such a move, as the first successful transplant of a donor heart to a human occurred that year. But he was interested in going a step further and creating an artificial heart for human use.
It took more than a decade of tinkering and research, but in 1982, a team of physicians and engineers led by Kolff succeeded in implanting the first artificial heart in dentist Barney Clark, whose failing health disqualified him from a heart transplant. Although Clark died in March 1983 after 112 days tethered to the device, that it kept him alive generated international headlines. While graduate student Robert Jarvik received the named credit for the heart, he was directly supervised by Kolff, whose various endeavors into artificial organ research at the University of Utah were segmented into numerous teams.
Forty years later, several artificial hearts have been approved for use by the Food and Drug Administration, although all are a “bridge” that allow patients to wait for a transplant.
Kolff continued researching and tinkering with biomedical devices – including artificial eyes and ears – until he retired in 1997 at the age of 86. When he died in 2009, the medical community acknowledged that he was not only a pioneer in biotechnology, but the “father” of artificial organs.
An image from one of TRIPP's virtual reality experiences for mental wellness. Nanea Reeves is TRIPP's CEO.
The "Making Sense of Science" podcast features interviews with leading experts about health innovations and the ethical questions they raise. The podcast is hosted by Matt Fuchs, editor of Leaps.org, the award-winning science outlet.
My guest today is Nanea Reeves, the CEO of TRIPP, a wellness platform with some big differences from meditation apps you may have tried like Calm and Headspace. TRIPP's experiences happen in virtual reality, and its realms are designed based on scientific findings about states of mindfulness. Users report feelings of awe and wonder and even mystical experiences. Nanea brings over 15 years of leadership in digital distribution, apps and video game technologies. Before co-founding TRIPP, she had several other leadership roles in tech with successful companies like textPlus and Machinima. Read her full bio below in the links section.

Nanea Reeves, CEO of TRIPP.
TRIPP
Listen to the Episode
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
This conversation coincided with National Brain Awareness Week. The topic is a little different from the Making Sense of Science podcast’s usual focus on breakthroughs in treating and preventing disease, but there’s a big overlap when it comes to breakthroughs in optimal health. Nanea’s work is at the leading edge of health, technology and the science of wellness.
With TRIPP, you might find yourself deep underwater, looking up at the sunlight shimmering on the ocean surface, or in the cosmos staring down at a planet glowing with an arresting diversity of colors. Using TRIPP for the past six months has been a window for me into the future of science-informed wellness and an overall fascinating experience, as was my conversation with Nanea.
Show notes:
Nanea and I discuss her close family members' substance addictions and her own struggle with mental illness as a teen, which led to her first meditation experiences, and much more:
- The common perception that technology is an obstacle for mental well-being, a narrative that overlooks how tech can also increase wellness when it’s designed right.
- Emerging ways of measuring meditation experiences by recording brain waves - and the shortcomings of the ‘measured self’ movement.
- Why TRIPP’s users multiplied during the stress and anxiety of the pandemic, and how TRIPP can can be used to enhance emotional states.
- Ways in which TRIPP’s visuals and targeted sound frequencies have been informed by innovative research from psychologists like Johns Hopkins’ Matthew Johnson.
- Ways to design apps and other technologies to better fulfill the true purpose of mindfulness meditation. (Hint: not simply relaxation.)
- And of course, because the topic is mental wellness and tech, I had to get Nanea's thoughts on Elon Musk, Neuralink and brain machine interfaces.
Here are links for learning more about TRIPP:
- TRIPP website: https://www.tripp.com/about/
- Nanea Reeves bio: https://www.tripp.com/team/nanea-reeves/
- Study of data collected by UK's Office for National Statistics on behavior during the pandemic, which suggests that TRIPP enhanced users' psychological and emotional mindsets: https://link.springer.com/chapter/10.1007/978-3-03...
- Research that's informed TRIPP: https://www.tripp.com/research/
- Washington Post Top Pick at CES: https://www.washingtonpost.com/technology/2019/01/...
- TRIPP's new offering, PsyAssist, to provide support for ketamine-assisted therapy: https://www.mobihealthnews.com/news/tripp-acquires...
- Randomized pilot trial involving TRIPP: https://bmjopen.bmj.com/content/bmjopen/11/4/e0441...