New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
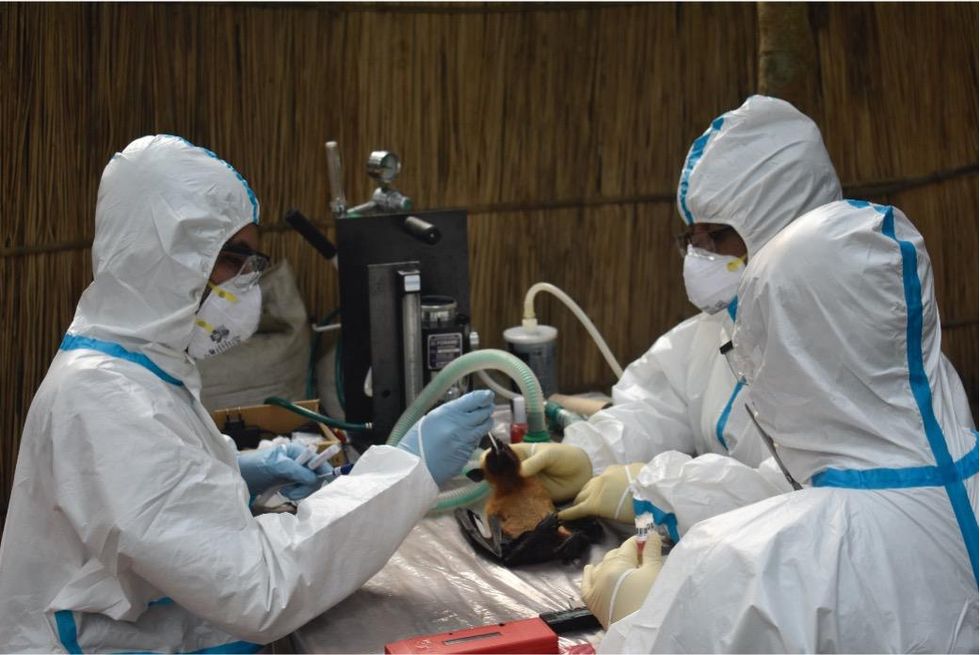
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
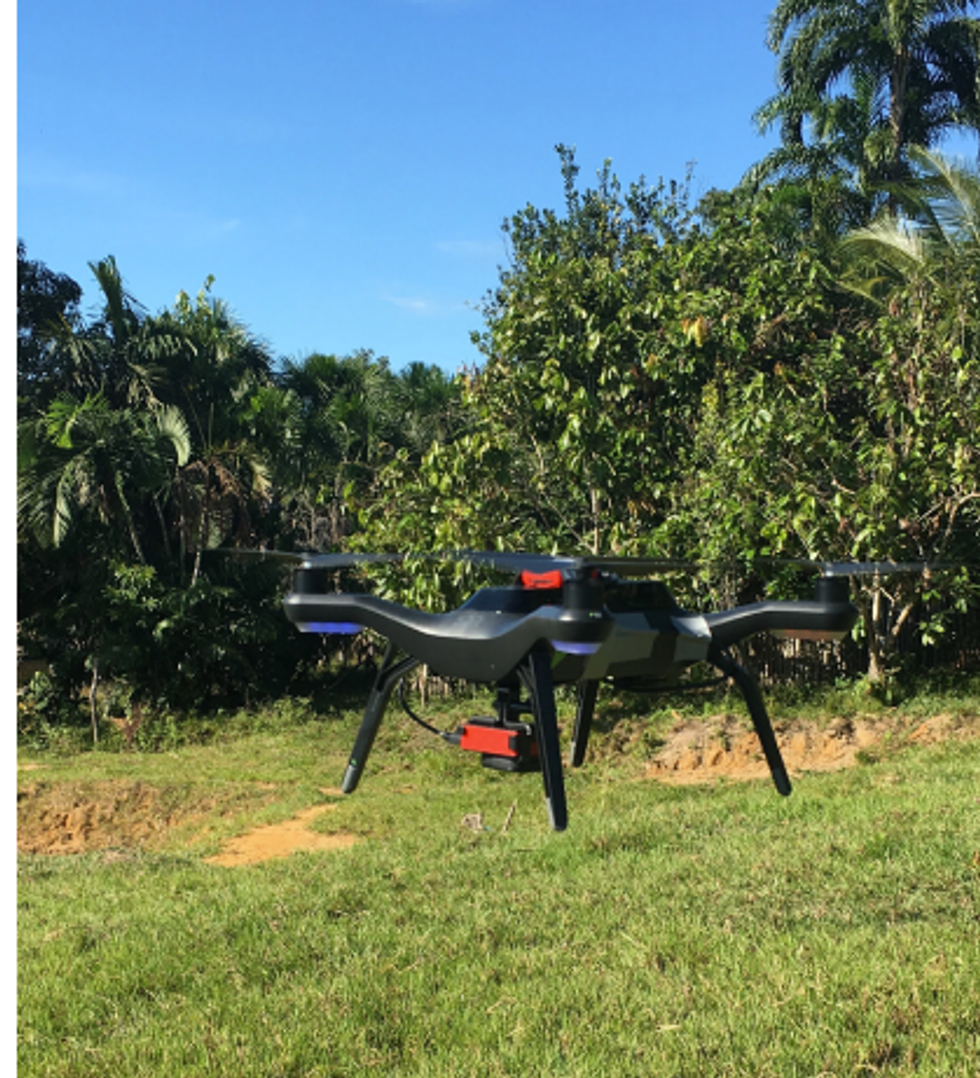

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Donna Shirley pictured at her home in Tulsa, with a model of the Sojourner rover she was in charge of that explored Mars.
When NASA's Perseverance rover landed successfully on Mars on February 18, 2021, calling it "one giant leap for mankind" – as Neil Armstrong said when he set foot on the moon in 1969 – would have been inaccurate. This year actually marked the fifth time the U.S. space agency has put a remote-controlled robotic exploration vehicle on the Red Planet. And it was a female engineer named Donna Shirley who broke new ground for women in science as the manager of both the Mars Exploration Program and the 30-person team that built Sojourner, the first rover to land on Mars on July 4, 1997.
For Shirley, the Mars Pathfinder mission was the climax of her 32-year career at NASA's Jet Propulsion Laboratory (JPL) in Pasadena, California. The Oklahoma-born scientist, who earned her Master's degree in aerospace engineering from the University of Southern California, saw her profile skyrocket with media appearances from CNN to the New York Times, and her autobiography Managing Martians came out in 1998. Now 79 and living in a Tulsa retirement community, she still embraces her status as a female pioneer.
"Periodically, I'll hear somebody say they got into the space program because of me, and that makes me feel really good," Shirley told Leaps.org. "I look at the mission control area, and there are a lot of women in there. I'm quite pleased I was able to break the glass ceiling."
Her $25-million, 25-pound microrover – powered by solar energy and designed to get rock samples and test soil chemistry for evidence of life – was named after Sojourner Truth, a 19th-century Black abolitionist and women's rights activist. Unlike Mars Pathfinder, Shirley didn't have to travel more than 131 million miles to reach her goal, but her path to scientific fame as a woman sometimes resembled an asteroid field.
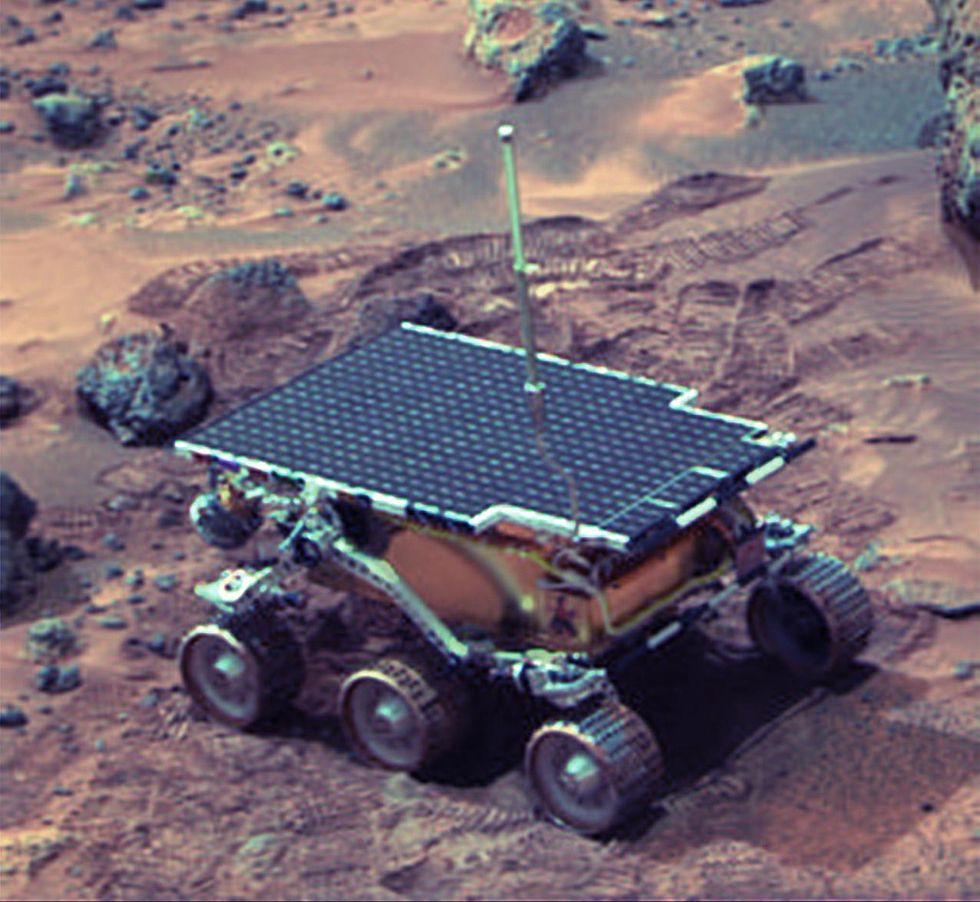 The Sojourner Rover in 1997 on Mars.NASA/JPL
The Sojourner Rover in 1997 on Mars.NASA/JPLAs a high-IQ tomboy growing up in Wynnewood, Oklahoma (pop. 2,300), Shirley yearned to escape. She decided to become an engineer at age 10 and took flying lessons at 15. Her extraterrestrial aspirations were fueled by Ray Bradbury's The Martian Chronicles and Arthur C. Clarke's The Sands of Mars. Yet when she entered the University of Oklahoma (OU) in 1958, her freshman academic advisor initially told her: "Girls can't be engineers." She ignored him.
Years later, Shirley would combat such archaic thinking, succeeding at JPL with her creative, collaborative management style. "If you look at the literature, you'll find that teams that are either led by or heavily involved with women do better than strictly male teams," she noted.
However, her career trajectory stalled at OU. Burned out by her course load and distracted by a broken engagement to marry a fellow student, she switched her major to professional writing. After graduation, she applied her aeronautical background as a McDonnell Aircraft technical writer, but her boss, she says, harassed her and she faced gender-based hostility from male co-workers.
Returning to OU, Shirley finished off her engineering degree and became a JPL aerodynamist in 1966 after answering an ad in the St. Louis Post-Dispatch. At first, she was the only female engineer among the research center's 2,000-odd engineers. She wore many hats, from designing planetary atmospheric entry vehicles to picking the launch date of November 4, 1973 for Mariner 10's mission to Venus and Mercury.
By the mid-1980's, she was managing teams that focused on robotics and Mars, delivering creative solutions when NASA budget cuts loomed. In 1989, the same year the Sojourner microrover concept was born, President George H.W. Bush announced his Space Exploration Initiative, including plans for a human mission to Mars by 2019.
That target, of course, wasn't attained, despite huge advances in technology and our understanding of the Martian environment. Today, Shirley believes humans could land on Mars by 2030. She became the founding director of the Science Fiction Museum and Hall of Fame in Seattle in 2004 after leaving NASA, and to this day, she enjoys checking out pop culture portrayals of Mars landings – even if they're not always accurate.
After the novel The Martian was published in 2011, which later was adapted into the hit film starring Matt Damon, Shirley phoned author Andy Weir: "You've got a major mistake in here. It says there's a storm that tries to blow the rocket over. But actually, the Mars atmosphere is so thin, it would never blow a rocket over!"
Fearlessly speaking her mind and seeking the stars helped Donna Shirley make history. However, a 2019 Washington Post story noted: "Women make up only about a third of NASA's workforce. They comprise just 28 percent of senior executive leadership positions and are only 16 percent of senior scientific employees." Whether it's traveling to Mars or trending toward gender equality, we've still got a long way to go.
Announcing March Event: "COVID Vaccines and the Return to Life: Part 1"
Leading medical and scientific experts will discuss the latest developments around the COVID-19 vaccines at our March 11th event.
EVENT INFORMATION
DATE:
Thursday, March 11th, 2021 at 12:30pm - 1:45pm EST
On the one-year anniversary of the global declaration of the pandemic, this virtual event will convene leading scientific and medical experts to discuss the most pressing questions around the COVID-19 vaccines. Planned topics include the effect of the new circulating variants on the vaccines, what we know so far about transmission dynamics post-vaccination, how individuals can behave post-vaccination, the myths of "good" and "bad" vaccines as more alternatives come on board, and more. A public Q&A will follow the expert discussion.
CONTACT:
kira@goodinc.com
LOCATION:
Zoom webinar
SPEAKERS:

Dr. Paul Offit speaking at Communicating Vaccine Science.
commons.wikimedia.orgDr. Paul Offit, M.D., is the director of the Vaccine Education Center and an attending physician in infectious diseases at the Children's Hospital of Philadelphia. He is a co-inventor of the rotavirus vaccine for infants, and he has lent his expertise to the advisory committees that review data on new vaccines for the CDC and FDA.

Dr. Monica Gandhi
UCSF Health
Dr. Monica Gandhi, M.D., MPH, is Professor of Medicine and Associate Division Chief (Clinical Operations/ Education) of the Division of HIV, Infectious Diseases, and Global Medicine at UCSF/ San Francisco General Hospital.

Dr. Onyema Ogbuagu, MBBCh, FACP, FIDSA
Yale Medicine
Dr. Onyema Ogbuagu, MBBCh, is an infectious disease physician at Yale Medicine who treats COVID-19 patients and leads Yale's clinical studies around COVID-19. He ran Yale's trial of the Pfizer/BioNTech vaccine.

Dr. Eric Topol
Dr. Topol's Twitter
Dr. Eric Topol, M.D., is a cardiologist, scientist, professor of molecular medicine, and the director and founder of Scripps Research Translational Institute. He has led clinical trials in over 40 countries with over 200,000 patients and pioneered the development of many routinely used medications.
REGISTER NOW
This event is the first of a four-part series co-hosted by LeapsMag, the Aspen Institute Science & Society Program, and the Sabin–Aspen Vaccine Science & Policy Group, with generous support from the Gordon and Betty Moore Foundation and the Howard Hughes Medical Institute.

Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.