New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
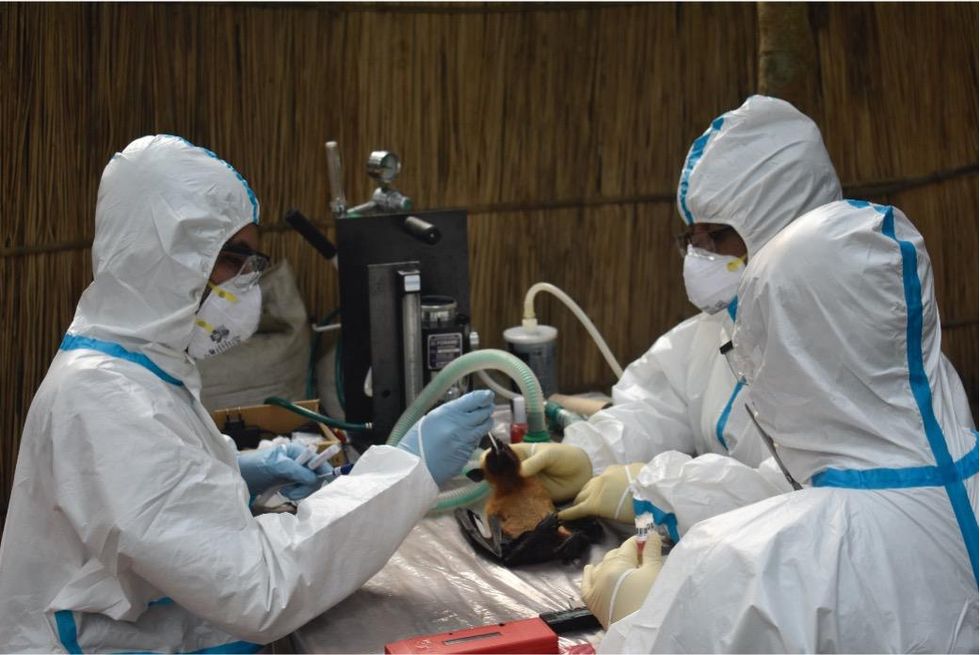
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
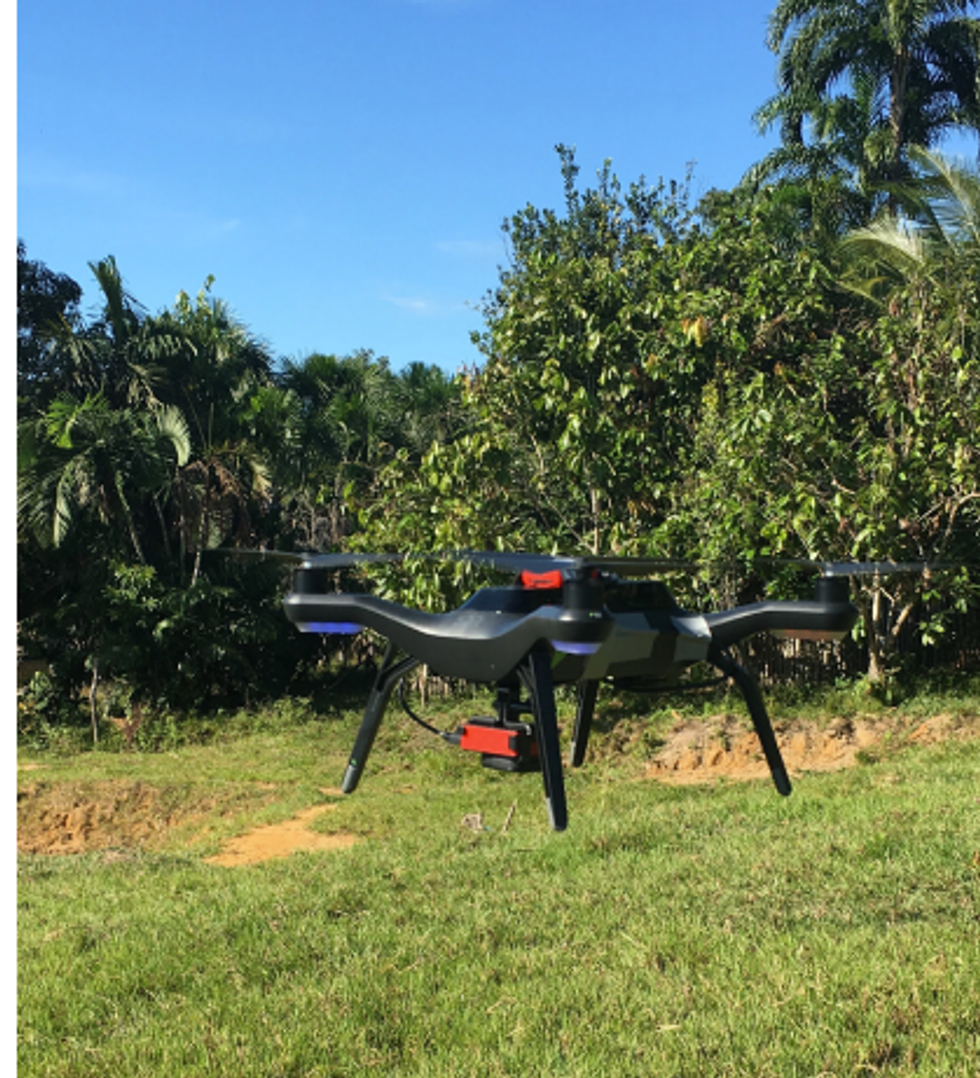

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
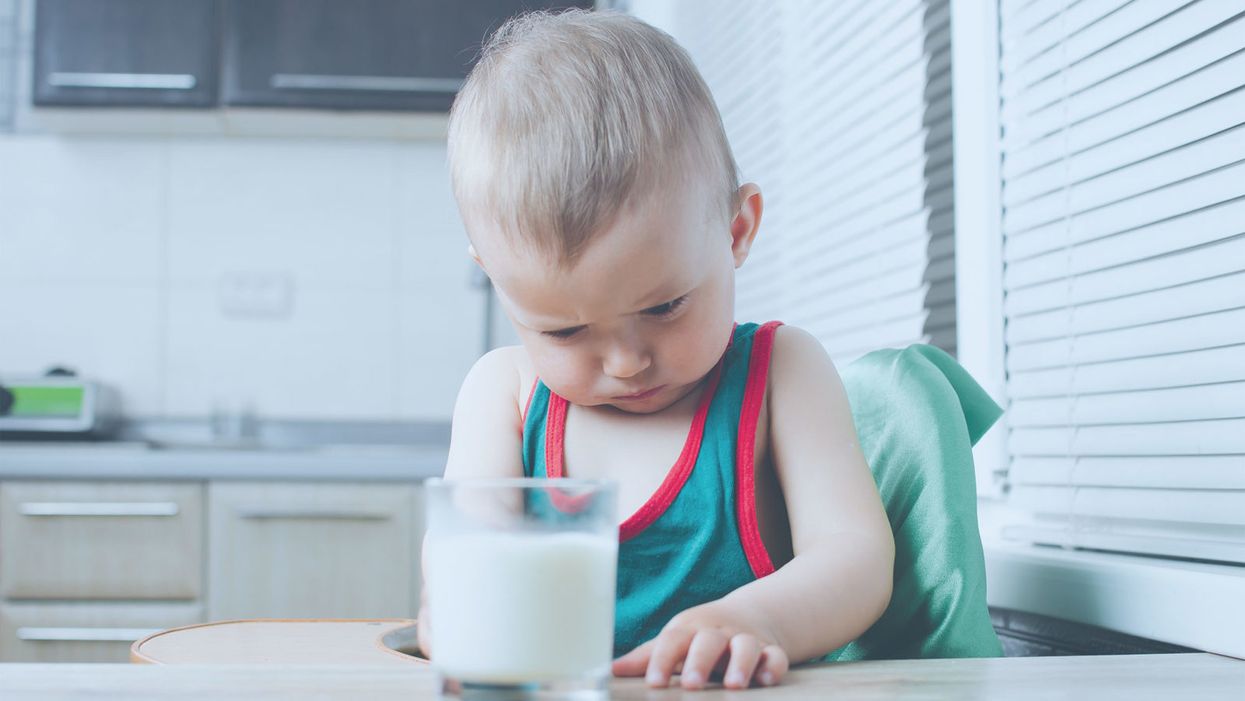
A baby who cannot tolerate milk due to an allergy.
Like any life-threatening medical condition that affects children, food allergies can traumatize more than just the patient. My wife and I learned this one summer afternoon when our daughter was three years old.
Emergency room visits for anaphylaxis in children more than doubled from 2010 to 2016.
At an ice cream parlor, I gave Samantha a lick of my pistachio cone; within seconds, red blotches erupted on her skin, her lips began to swell, and she complained that her throat felt funny. We rushed her to the nearest emergency room, where a doctor injected her with epinephrine. Explaining that the reaction, known as anaphylaxis, could have been fatal if left unchecked, he advised us to have her tested for nut allergies—and to start carrying an injector of our own.
After an allergist confirmed Sam's vulnerability to tree nuts and peanuts, we figured that keeping her safe would be relatively simple. But food allergies often come in bunches. Over the next year, she wound up back in the ER after eating bread with sesame seeds at an Italian restaurant, and again after slurping buckwheat noodles at our neighborhood Japanese. She hated eggs, so we discovered that (less severe) allergy only when she vomited after eating a variety of products containing them.
In recent years, a growing number of families have had to grapple with such challenges. An estimated 32 million Americans have food allergies, or nearly 10 percent of the population—10 times the prevalence reported 35 years ago. The severity of symptoms seems to be increasing, too. According to a study released in January by Food Allergy Research & Education (FARE), a Virginia-based nonprofit, insurance claims for anaphylactic food reactions rose 377 percent in the U.S. from 2007 to 2016.
Because food allergies most commonly emerge in childhood, these trends are largely driven by the young. An insurance-industry study found that emergency room visits for anaphylaxis in children more than doubled from 2010 to 2016. Peanut allergies, once rare, tripled in kids between 1997 and 2008. "The first year, it was 1 in 250," says Scott Sicherer, chief of pediatric allergy and immunology at New York City's Mount Sinai Hospital, who led that study. "When we did the next round of research, in 2002, it was 1 in 125. I thought there must be a mistake. But by 2008, it was 1 in 70."
The forces behind these dire statistics—as well as similar numbers throughout the developed world—have yet to be positively identified. But the leading suspects are elements of our modern lifestyle that can throw the immune system out of whack, prompting potentially deadly overreactions to harmless proteins. Although parents can take a few steps that might lessen their children's risk, societal changes may be needed to brighten the larger epidemiological picture.
Meanwhile, scientists are racing to develop therapies that can induce patients' hyped-up immune defenses to chill. And lately, they've made some big strides toward that goal.
A Variety of Culprits
In the United States, about 90 percent of allergic reactions come from eight foods: milk, eggs, peanuts, tree nuts, soy, wheat, fish, and shellfish. The list varies from country to country, depending on dietary customs, but what the trigger foods all have in common is proteins that can survive breakdown in the stomach and enter the bloodstream more or less intact.
"When we were kids, we played in the dirt. Today, children tend to be on their screens, inside sealed buildings."
A food allergy results from a chain of biochemical misunderstandings. The first time the immune system encounters an allergen (as a protein that triggers an allergy is known), it mistakes the substance for a hostile invader—perhaps a parasite with a similar molecular profile. In response, it produces an antibody called immunoglobin E (IgE), which is designed to bind to a specific protein and flag it for attack. These antibodies circulate through the bloodstream and attach to immune-system foot soldiers known as mast cells and basophils, which congregate in the nose, throat, lungs, skin, and gastrointestinal tract.
The next time the person is exposed to the allergen, the IgE antibodies signal the warrior cells to blast the intruder with histamines and other chemical weapons. Tissues in the affected areas swell and leak fluid; blood pressure may fall. Depending on the strength of the reaction, collateral damage to the patient can range from unpleasant—itching, runny nose, nausea—to catastrophic.
This kind of immunological glitchiness runs in families. Genome-wide association studies have identified a dozen genes linked to allergies of all types, and twin studies suggest that about 80 percent of the risk of food allergies is heritable. But why one family member shows symptoms while another doesn't remains unknown. Nor can genetics explain why food allergy rates have skyrocketed in such a brief period. For that, we must turn to the environment.
First, it's important to note that rates of all allergies are rising—including skin and respiratory afflictions—though none as rapidly or with as much risk of anaphylaxis as those involving food. The takeoff was already underway in the late 1980s, when British epidemiologist David P. Strachan found that children in larger households had fewer instances of hay fever. The reason, he suggested, was that their immune systems were strengthened by exposure to their siblings' germs. Since then, other researchers have discerned more evidence for Strachan's "hygiene hypothesis": higher rates of allergy (as well as autoimmune disorders) in cities versus rural areas, in industrialized countries versus developing ones, in lab animals raised under sterile conditions versus those exposed to germs.
Fending off a variety of pathogens, experts theorize, helps train the immune system to better distinguish friend from foe, and to respond to threats in a more nuanced manner. In an era of increasing urbanization, shrinking family sizes, and more sheltered lifestyles, such conditioning may be harder to come by. "When we were kids, we played in the dirt," observes Cathryn R. Nagler, a professor and food allergy researcher at the University of Chicago. "Today, children tend to be on their screens, inside sealed buildings."
But other factors may be driving the allergy epidemic as well. More time indoors, for example, means less exposure to sunlight, which can lead to a deficiency in vitamin D—a nutrient crucial to immune system regulation. The growing popularity of processed foods filled with refined fats and sugars may play a role, along with rising rates of obesity, by promoting tissue inflammation that could increase some people's risk of immunological mayhem. And the surge in allergies also correlates with several trends that may be altering the human microbiome, the community of microbes (including bacteria, viruses, and fungi, among others) that inhabits our guts, skin, and bodily orifices.
The microbiome connection may be particularly relevant to food allergies. In 2014, a team led by Nagler published a landmark study showing that Clostridia, a common class of gut bacteria, protects against these allergies. When the researchers fed peanut allergens to germ-free mice (born and raised in sterile conditions) and to mice treated with antibiotics as newborns (reducing their gut bacteria), the animals showed a strong immunological response. This sensitization could be reversed, however, by reintroducing Clostridia—but not another class of bacteria, Bacteroides—into the mice. Further experiments revealed that Clostridia caused immune cells to produce high levels of interleukin-22 (IL-22), a signaling molecule known to decrease the permeability of the intestinal lining.
"In simple terms," Nagler says, "what we found is that these bacteria prevent food allergens from gaining access to the blood in an intact form that elicits an allergic reaction."
A growing body of evidence suggests that our eating habits are throwing our gut microbiota off-balance, in part by depriving helpful species of the dietary fiber they feed on. Our increasing exposure to antibiotics and antimicrobial compounds may be harming our beneficial bugs as well. These depletions could affect kids from the moment they enter the world: Because babies are seeded with their mothers' microbiota as they pass through the birth canal, they may be inheriting a less diverse microbiome than did previous generations. And the rising rate of caesarian deliveries may be further depriving our children of the bugs they need.
On expert suggests two measures worth a try: increasing consumption of fiber, and reducing use of antimicrobial agents, from antibacterial cleaners to antibiotics.
So which culprit is most responsible for the food allergy upsurge? "The illnesses that we're measuring are complex," says Sicherer. "There are multiple genetic inputs, which interact with one another, and there are multiple environmental inputs, which interact with each other and with the genes. There's not one single thing that's causing this. It's a conglomeration."
What Parents Can Do
For anyone hoping to reduce their child's or their own odds of developing a food allergy (rates of adult onset are also increasing), the current state of science offers few guideposts. As with many other areas of health research, it's hard to know when the data is solid enough to warrant a particular course of action. A case in point: the American Academy of Pediatrics once recommended that children at risk of allergy to peanuts (as evidenced by family history, other food allergies, or eczema) wait to eat them until age three; now, the AAP advises those parents to start their babies at four months, citing epidemiological evidence that early exposure may prevent peanut allergies.
And it's all too easy for a layperson to draw mistaken conclusions from media coverage of such research—inferring, for instance, that taking commercially available probiotics might have a protective effect. Unfortunately, says Nagler, none of those products even contain the relevant kind of bacteria.
Although, as a research scientist, she refrains from giving medical advice, Nagler does suggest (based on a large body of academic literature) that two measures are worth a try: increasing consumption of fiber, and reducing use of antimicrobial agents, from antibacterial cleaners to antibiotics. Yet she acknowledges that it's not always possible to avoid the suspected risk factors for food allergies. Sometimes an antibiotic is a lifesaving necessity, for example—and it's tough to avoid exposure to such drugs altogether, due to their use in animal feed and their consequent presence in many foods and in the water supply. If these chemicals are contributing to the food allergy epidemic, protecting ourselves will require action from farmers, doctors, manufacturers, and policymakers.
My family's experience illustrates the limits of healthy lifestyle choices in mitigating allergy risk. My daughter and son were born without C-sections; both were breastfed as well, receiving maximum microbial seeding from their mother. As a family, we eat exemplary diets, and no one could describe our home as excessively clean. Yet one child can't taste nuts, sesame, or buckwheat without becoming dangerously ill. "You can do everything right and still have allergies," says Ian A. Myles, a staff clinician at the National Institute of Allergy and Infectious Diseases. "You can do everything wrong and not have allergies. The two groups overlap."
The Latest Science Shows Promise
But while preventing all food allergies is clearly unrealistic, researchers are making remarkable progress in developing better treatments—therapies that, instead of combating symptoms after they've started (like epinephrine or antihistamines), aim to make patients less sensitive to allergens in the first place. One promising approach is oral immunotherapy (OIT), in which patients consume small but slowly increasing amounts of an allergen, gradually reducing their sensitivity. A study published last year in the New England Journal of Medicine showed that an experimental OIT called AR101, consisting of a standardized peanut powder mixed into food, enabled 67 percent of participants to tolerate a dose equivalent to two peanut kernels—a potential lifesaver if they were accidentally exposed to the real thing.
Because OIT itself can trigger troublesome reactions in some patients, however, it's not for everyone. Another experimental treatment, sublingual immunotherapy (SLIT) uses an allergen solution or dissolving tablet placed beneath the tongue; although its results are less robust than OIT's, it seems to generate milder side effects. Epicutaneous immunotherapy (EPIT) avoids the mouth entirely, using a technology similar to a nicotine patch to deliver allergens through the skin. Researchers are also exploring the use of medications known as biologics, aiming to speed up the action of immunotherapies by suppressing IgE or targeting other immune-system molecules.
These findings suggest that drugs based on microbial metabolites could help protect vulnerable individuals against a wide range of allergies.
One downside of the immunotherapy approach is that in most cases the allergen must be taken indefinitely to maintain desensitization. To provide a potentially permanent fix, scientists are working on vaccines that use DNA or peptides (protein fragments) from allergens to reset patients' immune systems.
Nagler is attacking the problem from a different angle—one that starts with the microbiome. In a recent study, a follow-up to her peanut-allergy investigation, she and her colleagues found that Clostridia bacteria protect mice against milk allergy as well; they also identified a particular species responsible, known as Anaerostipes caccae. The bugs, the team determined, produce a short-chain fatty acid called butyrate, which modulates many immune activities crucial to maintaining a well-sealed gut.
These findings suggest that drugs based on microbial metabolites could help protect vulnerable individuals against a wide range of allergies. Nagler has launched a company, ClostraBio, to develop biotherapeutics based on this notion; she expects its first product, using synthetic butyrate, to be ready for clinical trials within the next two years.
My daughter could well be a candidate for such a medication. Sam, now 15, is a vibrant, resilient kid who handles her allergies with confidence and humor. Thanks to vigilance and luck (on her part as well as her parents'), she hasn't had another food-related ER visit in more than a decade; she's never had to use her Epi-Pen. Still, she says, she would welcome the arrival of a pill that could reduce the danger. "I've learned how to watch out for myself," she says. "But it would be nice not to have to be so careful."
A brownie cake and whipped cream beckon on a plate.
Imagine eating a slice of cake for breakfast. It's deliciously indulgent, but instead of your blood sugar spiking, your body processes all that sweetness as a healthy high-protein meal. It may sound like sci-fi, but this scenario is not necessarily far off.
"People with diabetes could especially benefit because sweet proteins don't trigger a need for insulin."
The Lowdown
An award-winning agtech startup called Amai is developing "sweet proteins," based on the molecular structure of naturally occuring exotic fruits. These new sugar substitutes could potentially replace artificial sweeteners and help people who are trying to curb their sugar intake. People with diabetes could especially benefit because sweet proteins don't trigger a need for insulin.
While there is a sweet protein currently on the market today called thaumatin, it's expensive, has a short shelf life, and is lacking in the taste department. But Amai's proteins taste 70 to 100 percent identical to the sweet ones found in nature. Once their molecular structure is designed through a sophisticated computing platform, they are made through fermentation, which is akin to brewing beer. These non-GMO proteins are over 10,000 times sweeter than sugar, which means much less needs to be produced and used.
Diseases like diabetes and heart disease, which are often linked to sugar overconsumption, have been on a major upswing over the last few decades, especially in the United States. According to the CDC, 100 million adults in the United States are now living with diabetes or prediabetes, which if not treated, often leads to type 2 diabetes within five years. By 2030, scientists predict cases of diabetes in the U.S. will increase by 54 percent. If sugar proteins like the type Amai is creating become widely available, these numbers could begin to decrease.
Next Up
Amai's sweet proteins are still in the research and development stage, but the Israeli startup is raising significant funding that should help expedite the process. They're also substantially upping their production ability by expanding their facilities.
Will consumers be comfortable ingesting a lab-designed food product?
And in March, the USDA and FDA announced plans to regulate cell-cultured foods, the category in which these sugar proteins would fall, so Amai researchers are hopeful they'll have an easier path to approval once their product is market ready.
Open Questions
All this progress may sound promising, but Amai still has a long way to go before the reality of healthy cake becomes tangible. Some questions to consider: Will consumers be comfortable ingesting a lab-designed food product? Will it taste enough like real sugar?
And if some products and brands begin to adopt it, will it ever overtake the real thing in popularity and make a dent in diseases like diabetes and obesity? Only time, more research, and a lot more money will tell, but in the meantime, feel free to daydream about eating entire pints of ice cream without needing to hit the gym.

