New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
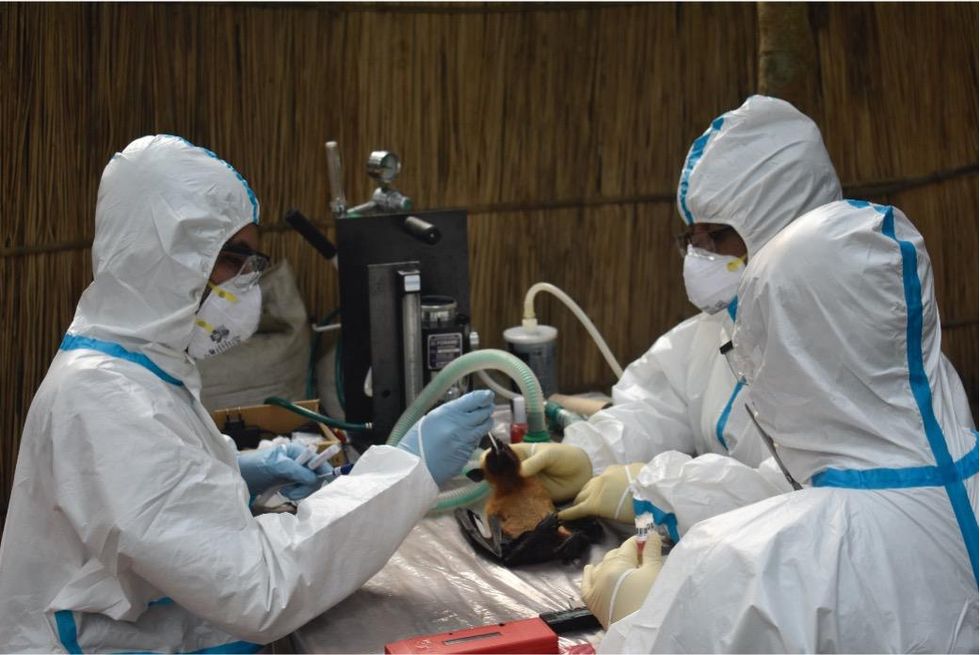
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
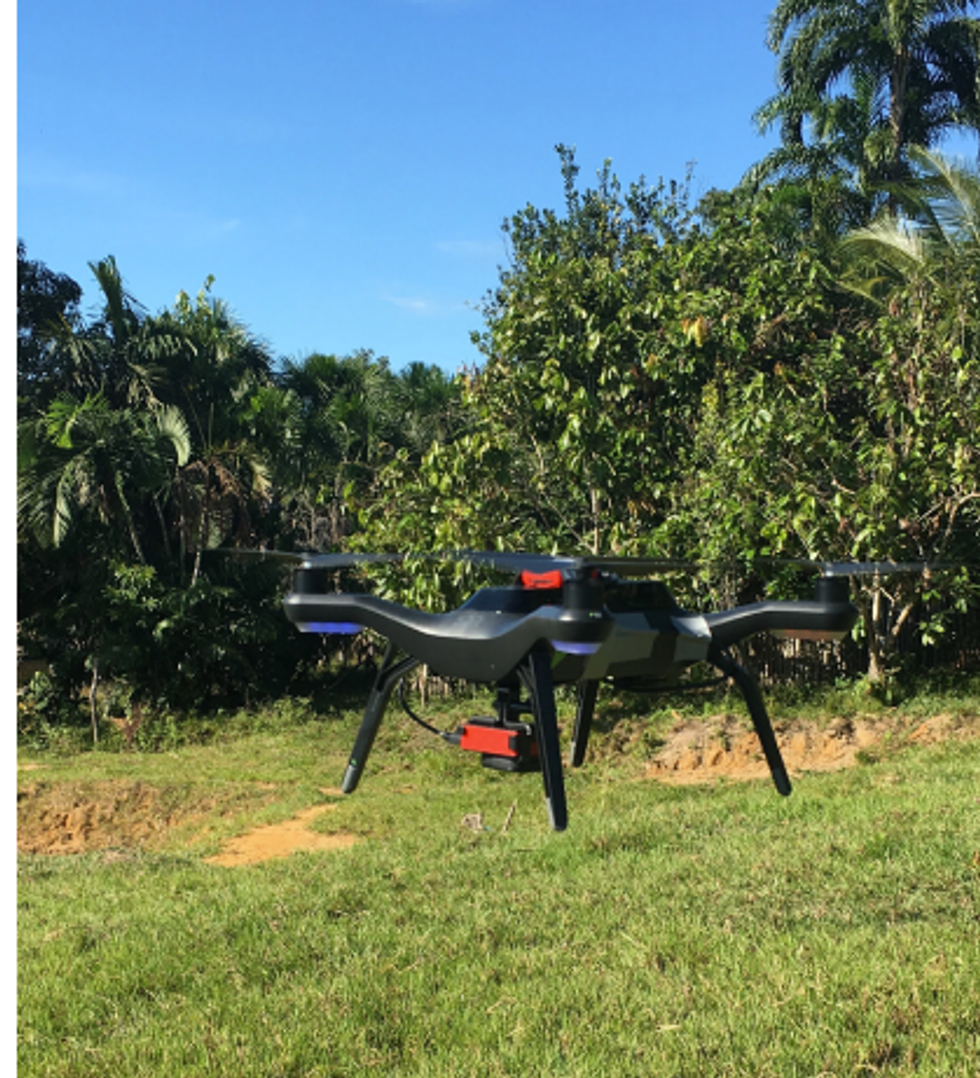

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
The Friday Five: A new blood test to detect Alzheimer's
A new blood test has been developed to detect Alzheimer's. Other promising studies covered in this week's Friday Five include: vets with PTSD can take their psychologist anywhere with a new device, intermittent fasting could improve circadian rhythms, a new year's resolution for living longer, and a discovery in 3-D printing eye tissue.
The Friday Five covers five stories in research that you may have missed this week. There are plenty of controversies and troubling ethical issues in science – and we get into many of them in our online magazine – but this news roundup focuses on scientific creativity and progress to give you a therapeutic dose of inspiration headed into the weekend.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
Here are the promising studies covered in this week's Friday Five:
- A blood test to detect Alzheimer's
- War vets can take their psychologist wherever they go
- Does intermittent fasting affect circadian rhythms?
- A new year's resolution for living longer
- 3-D printed eyes?
Staying well in the 21st century is like playing a game of chess
The control of infectious diseases was considered to be one of the “10 Great Public Health Achievements.” What we didn’t take into account was the very concept of evolution: as we built better protections, our enemies eventually boosted their attacking prowess, so soon enough we found ourselves on the defensive once again.
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
On July 30, 1999, the Centers for Disease Control and Prevention published a report comparing data on the control of infectious disease from the beginning of the 20th century to the end. The data showed that deaths from infectious diseases declined markedly. In the early 1900s, pneumonia, tuberculosis and diarrheal diseases were the three leading killers, accounting for one-third of total deaths in the U.S.—with 40 percent being children under five.
Mass vaccinations, the discovery of antibiotics and overall sanitation and hygiene measures eventually eradicated smallpox, beat down polio, cured cholera, nearly rid the world of tuberculosis and extended the U.S. life expectancy by 25 years. By 1997, there was a shift in population health in the U.S. such that cancer, diabetes and heart disease were now the leading causes of death.
The control of infectious diseases is considered to be one of the “10 Great Public Health Achievements.” Yet on the brink of the 21st century, new trouble was already brewing. Hospitals were seeing periodic cases of antibiotic-resistant infections. Novel viruses, or those that previously didn’t afflict humans, began to emerge, causing outbreaks of West Nile, SARS, MERS or swine flu.In the years that followed, tuberculosis made a comeback, at least in certain parts of the world. What we didn’t take into account was the very concept of evolution: as we built better protections, our enemies eventually boosted their attacking prowess, so soon enough we found ourselves on the defensive once again.
At the same time, new, previously unknown or extremely rare disorders began to rise, such as autoimmune or genetic conditions. Two decades later, scientists began thinking about health differently—not as a static achievement guaranteed to last, but as something dynamic and constantly changing—and sometimes, for the worse.
What emerged since then is a different paradigm that makes our interactions with the microbial world more like a biological chess match, says Victoria McGovern, a biochemist and program officer for the Burroughs Wellcome Fund’s Infectious Disease and Population Sciences Program. In this chess game, humans may make a clever strategic move, which could involve creating a new vaccine or a potent antibiotic, but that advantage is fleeting. At some point, the organisms we are up against could respond with a move of their own—such as developing resistance to medication or genetic mutations that attack our bodies. Simply eradicating the “opponent,” or the pathogenic microbes, as efficiently as possible isn’t enough to keep humans healthy long-term.
Instead, scientists should focus on studying the complexity of interactions between humans and their pathogens. “We need to better understand the lifestyles of things that afflict us,” McGovern says. “The solutions are going to be in understanding various parts of their biology so we can influence how they behave around our systems.”
Genetics and cell biology, combined with imaging techniques that allow one to see tissues and individual cells in actions, will enable scientists to define and quantify what it means to be healthy at the molecular level.
What is being proposed will require a pivot to basic biology and other disciplines that have suffered from lack of research funding in recent years. Yet, according to McGovern, the research teams of funded proposals are answering bigger questions. “We look for people exploring questions about hosts and pathogens, and what happens when they touch, but we’re also looking for people with big ideas,” she says. For example, if one specific infection causes a chain of pathological events in the body, can other infections cause them too? And if we find a way to break that chain for one pathogen, can we play the same trick on another? “We really want to see people thinking of not just one experiment but about big implications of their work,” McGovern says.
Jonah Cool, a cell biologist, geneticist and science officer at the Chan Zuckerberg Initiative, says that it’s necessary to define what constitutes a healthy organism and how it overcomes infections or environmental assaults, such as pollution from forest fires or toxins from industrial smokestacks. An organism that catches a disease isn’t necessarily an unhealthy one, as long as it fights it off successfully—an ability that arises from the complex interplay of its genes, the immune system, age, stress levels and other factors. Modern science allows many of these factors to be measured, recorded and compared. “We need a data-driven, deep-phenotyping approach to defining healthy biological systems and their responses to insults—which can be infectious disease or environmental exposures—and their ability to navigate their way through that space,” Cool says.
Genetics and cell biology, combined with imaging techniques that allow one to see tissues and individual cells in actions, will enable scientists to define and quantify what it means to be healthy at the molecular level. “As a geneticist and cell biologist, I believe in all these molecular underpinnings and how they arise in phenotypic differences in cells, genes, proteins—and how their combinations form complex cellular states,” Cool says.
Julie Graves, a physician, public health consultant, former adjunct professor of management, policy and community health at the University of Texas Health Science Center in Houston, stresses the necessity of nutritious diets. According to the Rockefeller Food Initiative, “poor diet is the leading risk factor for disease, disability and premature death in the majority of countries around the world.” Adequate nutrition is critical for maintaining human health and life. Yet, Western diets are often low in essential nutrients, high in calories and heavy on processed foods. Overconsumption of these foods has contributed to high rates of obesity and chronic disease in the U.S. In fact, more than half of American adults have at least one chronic disease, and 27 percent have more than one—which increases vulnerability to COVID-19 infections, according to the 2018 National Health Interview Survey.
Further, the contamination of our food supply with various agricultural and industrial toxins—petrochemicals, pesticides, PFAS and others—has implications for morbidity, mortality, and overall quality of life. “These chemicals are insidiously in everything, including our bodies,” Graves says—and they are interfering with our normal biological functions. “We need to stop how we manufacture food,” she adds, and rid our sustenance of these contaminants.
According to the Humane Society of the United States, factory farms result in nearly 40 percent of emissions of methane. Concentrated animal feeding operations or CAFOs may serve as breeding grounds for pandemics, scientists warn, so humans should research better ways to raise and treat livestock. Diego Rose, a professor of food and nutrition policy at Tulane University School of Public Health & Tropical Medicine, and his colleagues found that “20 percent of Americans’ diets account for about 45 percent of the environmental impacts [that come from food].” A subsequent study explored the impacts of specific foods and found that substituting beef for chicken lowers an individual’s carbon footprint by nearly 50 percent, with water usage decreased by 30 percent. Notably, however, eating too much red meat has been associated with a variety of illnesses.
In some communities, the option to swap food types is limited or impossible. For example, “many populations live in relative food deserts where there’s not a local grocery store that has any fresh produce,” says Louis Muglia, the president and CEO of Burroughs Wellcome. Individuals in these communities suffer from an insufficient intake of beneficial macronutrients, and they’re “probably being exposed to phenols and other toxins that are in the packaging.” An equitable, sustainable and nutritious food supply will be vital to humanity’s wellbeing in the era of climate change, unpredictable weather and spillover events.
A recent report by See Change Institute and the Climate Mental Health Network showed that people who are experiencing socioeconomic inequalities, including many people of color, contribute the least to climate change, yet they are impacted the most. For example, people in low-income communities are disproportionately exposed to vehicle emissions, Muglia says. Through its Climate Change and Human Health Seed Grants program, Burroughs Wellcome funds research that aims to understand how various factors related to climate change and environmental chemicals contribute to premature births, associated with health vulnerabilities over the course of a person’s life—and map such hot spots.
“It’s very complex, the combinations of socio-economic environment, race, ethnicity and environmental exposure, whether that’s heat or toxic chemicals,” Muglia explains. “Disentangling those things really requires a very sophisticated, multidisciplinary team. That’s what we’ve put together to describe where these hotspots are and see how they correlate with different toxin exposure levels.”
In addition to mapping the risks, researchers are developing novel therapeutics that will be crucial to our armor arsenal, but we will have to be smarter at designing and using them. We will need more potent, better-working monoclonal antibodies. Instead of directly attacking a pathogen, we may have to learn to stimulate the immune system—training it to fight the disease-causing microbes on its own. And rather than indiscriminately killing all bacteria with broad-scope drugs, we would need more targeted medications. “Instead of wiping out the entire gut flora, we will need to come up with ways that kill harmful bacteria but not healthy ones,” Graves says. Training our immune systems to recognize and react to pathogens by way of vaccination will keep us ahead of our biological opponents, too. “Continued development of vaccines against infectious diseases is critical,” says Graves.
With all of the unpredictable events that lie ahead, it is difficult to foresee what achievements in public health will be reported at the end of the 21st century. Yet, technological advances, better modeling and pursuing bigger questions in science, along with education and working closely with communities will help overcome the challenges. The Chan Zuckerberg Initiative displays an optimistic message on its website: “Is it possible to cure, prevent, or manage all diseases by the end of this century? We think so.” Cool shares the view of his employer—and believes that science can get us there. Just give it some time and a chance. “It’s a big, bold statement,” he says, “but the end of the century is a long way away.”Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.