New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
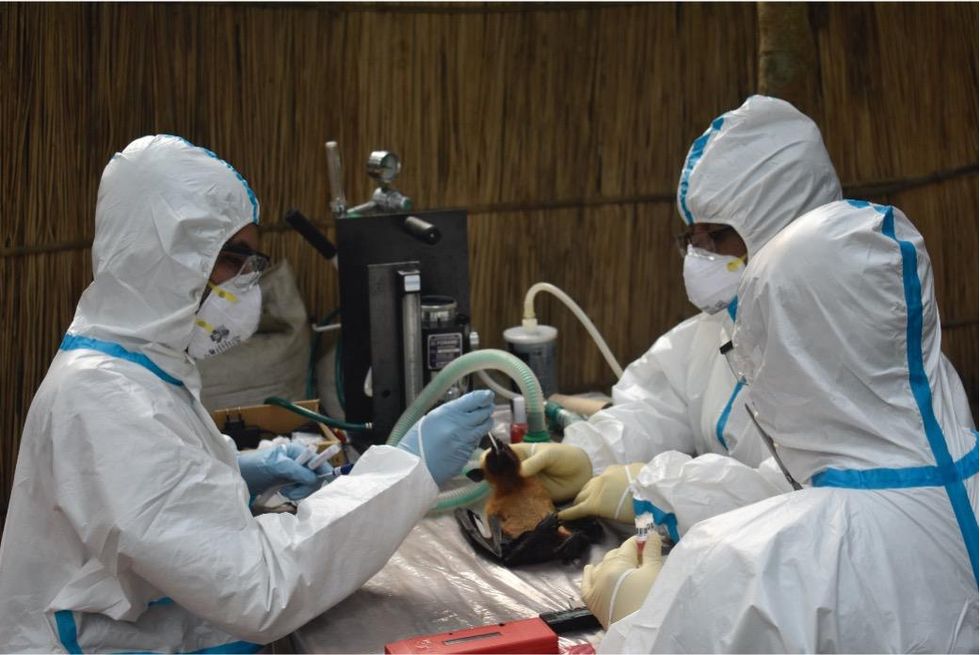
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
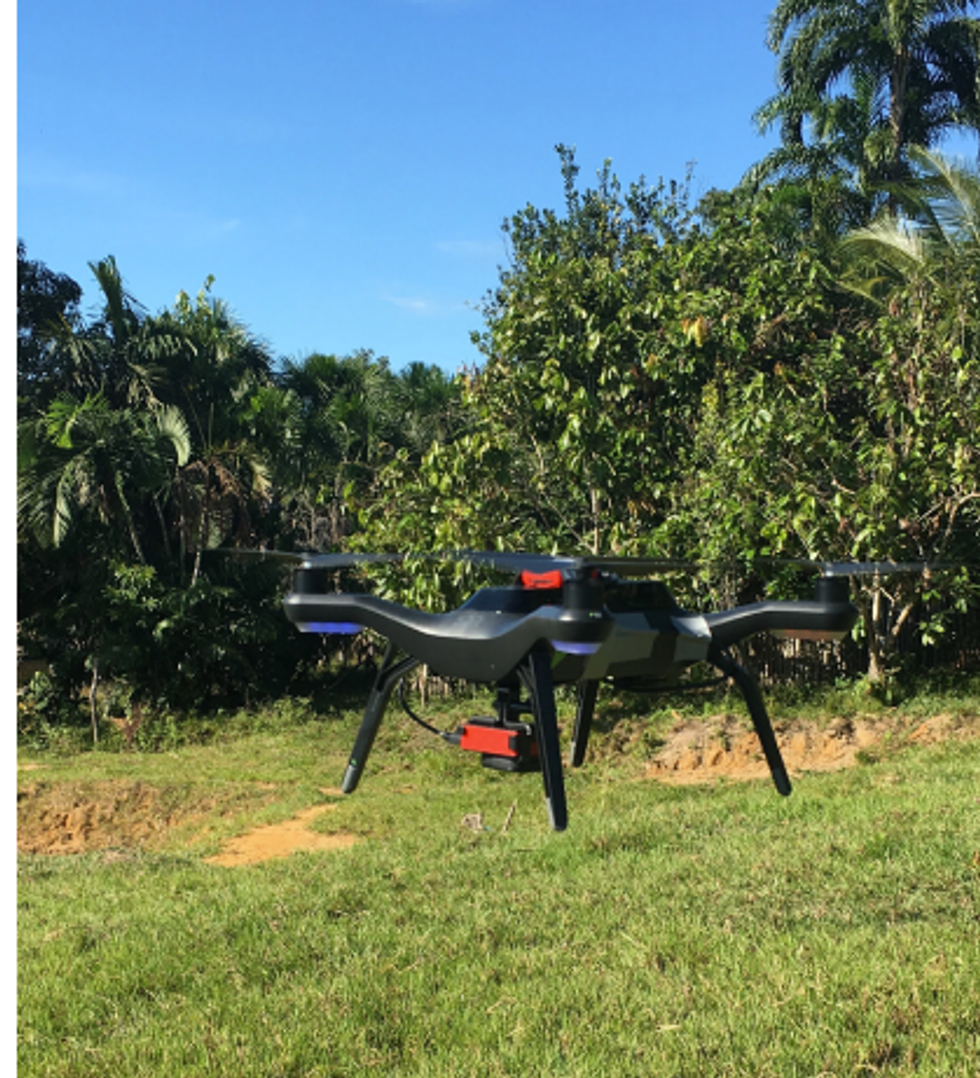

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Researchers at Stanford have found that the virus that causes Covid-19 can infect fat cells, which could help explain why obesity is linked to worse outcomes for those who catch Covid-19.
Obesity is a risk factor for worse outcomes for a variety of medical conditions ranging from cancer to Covid-19. Most experts attribute it simply to underlying low-grade inflammation and added weight that make breathing more difficult.
Now researchers have found a more direct reason: SARS-CoV-2, the virus that causes Covid-19, can infect adipocytes, more commonly known as fat cells, and macrophages, immune cells that are part of the broader matrix of cells that support fat tissue. Stanford University researchers Catherine Blish and Tracey McLaughlin are senior authors of the study.
Most of us think of fat as the spare tire that can accumulate around the middle as we age, but fat also is present closer to most internal organs. McLaughlin's research has focused on epicardial fat, “which sits right on top of the heart with no physical barrier at all,” she says. So if that fat got infected and inflamed, it might directly affect the heart.” That could help explain cardiovascular problems associated with Covid-19 infections.
Looking at tissue taken from autopsy, there was evidence of SARS-CoV-2 virus inside the fat cells as well as surrounding inflammation. In fat cells and immune cells harvested from health humans, infection in the laboratory drove "an inflammatory response, particularly in the macrophages…They secreted proteins that are typically seen in a cytokine storm” where the immune response runs amok with potential life-threatening consequences. This suggests to McLaughlin “that there could be a regional and even a systemic inflammatory response following infection in fat.”
It is easy to see how the airborne SARS-CoV-2 virus infects the nose and lungs, but how does it get into fat tissue? That is a mystery and the source of ample speculation.
The macrophages studied by McLaughlin and Blish were spewing out inflammatory proteins, While the the virus within them was replicating, the new viral particles were not able to replicate within those cells. It was a different story in the fat cells. “When [the virus] gets into the fat cells, it not only replicates, it's a productive infection, which means the resulting viral particles can infect another cell,” including microphages, McLaughlin explains. It seems to be a symbiotic tango of the virus between the two cell types that keeps the cycle going.
It is easy to see how the airborne SARS-CoV-2 virus infects the nose and lungs, but how does it get into fat tissue? That is a mystery and the source of ample speculation.
Macrophages are mobile; they engulf and carry invading pathogens to lymphoid tissue in the lymph nodes, tonsils and elsewhere in the body to alert T cells of the immune system to the pathogen. Perhaps some of them also carry the virus through the bloodstream to more distant tissue.
ACE2 receptors are the means by which SARS-CoV-2 latches on to and enters most cells. They are not thought to be common on fat cells, so initially most researchers thought it unlikely they would become infected.
However, while some cell receptors always sit on the surface of the cell, other receptors are expressed on the surface only under certain conditions. Philipp Scherer, a professor of internal medicine and director of the Touchstone Diabetes Center at the University of Texas Southwestern Medical Center, suggests that, in people who have obesity, “There might be higher levels of dysfunctional [fat cells] that facilitate entry of the virus,” either through transiently expressed ACE2 or other receptors. Inflammatory proteins generated by macrophages might contribute to this process.
Another hypothesis is that viral RNA might be smuggled into fat cells as cargo in small bits of material called extracellular vesicles, or EVs, that can travel between cells. Other researchers have shown that when EVs express ACE2 receptors, they can act as decoys for SARS-CoV-2, where the virus binds to them rather than a cell. These scientists are working to create drugs that mimic this decoy effect as an approach to therapy.
Do fat cells play a role in Long Covid? “Fat cells are a great place to hide. You have all the energy you need and fat cells turn over very slowly; they have a half-life of ten years,” says Scherer. Observational studies suggest that acute Covid-19 can trigger the onset of diabetes especially in people who are overweight, and that patients taking medicines to regulate their diabetes “were actually quite protective” against acute Covid-19. Scherer has funding to study the risks and benefits of those drugs in animal models of Long Covid.
McLaughlin says there are two areas of potential concern with fat tissue and Long Covid. One is that this tissue might serve as a “big reservoir where the virus continues to replicate and is sent out” to other parts of the body. The second is that inflammation due to infected fat cells and macrophages can result in fibrosis or scar tissue forming around organs, inhibiting their function. Once scar tissue forms, the tissue damage becomes more difficult to repair.
Current Covid-19 treatments work by stopping the virus from entering cells through the ACE2 receptor, so they likely would have no effect on virus that uses a different mechanism. That means another approach will have to be developed to complement the treatments we already have. So the best advice McLaughlin can offer today is to keep current on vaccinations and boosters and lose weight to reduce the risk associated with obesity.
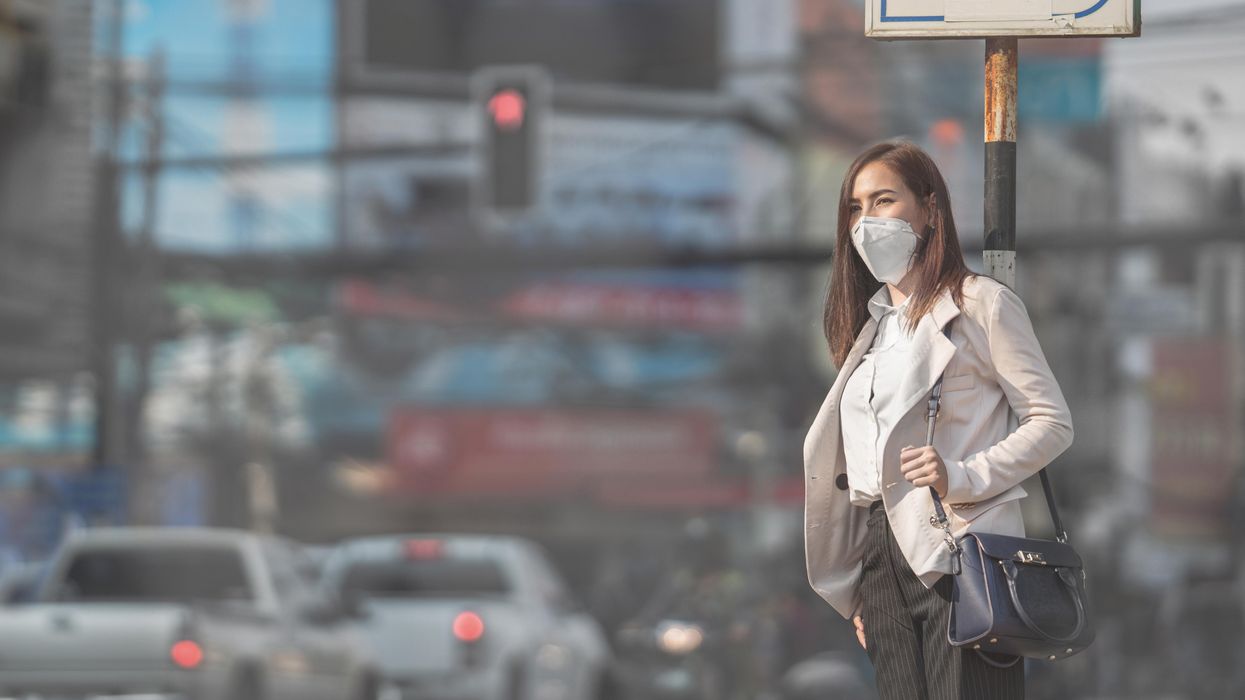
Air pollution can lead to lung cancer. The connection suggests new ways to stop cancer in its tracks.
Researchers at Francis Crick Institute found that air pollution can "wake up" existing mutations, triggering them to turn into lung cancer. The scientists used their new understanding about this connection to prevent cancer in mice.
Forget taking a deep breath. Around the world, 99 percent of people breathe air polluted to unsafe levels, according to data from the World Health Organization. Activities such as burning fossil fuels release greenhouse gases that contribute to air pollution, which could lead to heart disease, stroke, asthma, emphysema, and some types of cancer.
“The burden of disease attributable to air pollution is now estimated to be on a par with other major global health risks such as unhealthy diet and tobacco smoking, and air pollution is now recognized as the single biggest environmental threat to human health,” wrote the authors of a 2021 WHO report.
The majority of lung cancer is attributed to smoking. But as pollution levels have increased, and anti-smoking campaigns have discouraged smoking, the proportion of lung cancers diagnosed in non-smokers has grown. The CDC estimates that 10 to 20 percent of lung cancers in the U.S. currently occur in non-smokers.
The mechanism between air pollution and the development of lung cancer has been unclear, but researchers at London’s Francis Crick Institute recently made an important breakthrough in understanding the connection. Lead investigator Charles Swanton presented this research last month at a conference in Paris.
Pollution awakens mutations
The Crick Institute scientists were able to identify a new link between common air pollutants and non-small cell lung cancer (NSCLC). They focused on pollutants called particulate matter, or PM, that are 2.5 microns wide, narrower than human cells.
Most cancer diagnosed in non-smoking people is NSCLC, but this type of cancer hasn’t received the same research attention as more common lung cancers found in smokers, according to Clare Weeden, a cancer researcher at the Crick Institute and a co-author of the study.
“This is a really underserved and under-researched population that we really need to tackle, as well as lung cancers that occur in smokers,” she says. “Lung cancer is the number one cancer killer worldwide.”
In the past, some researchers believed air pollution caused mutations that led to cancer. Others believed these mutations could remain dormant without any detriment to health until pollutants or other stressors triggered them to become cancerous. Reviving the latter hypothesis that carcinogens may activate pre-existing mutations, instead of directly causing them, the Crick researchers analyzed samples from 463,679 people in the UK and parts of Asia, noting mutations and comparing changes in gene expression in mice and human cells.
“The mutation can exist in a nascent clone without causing cancer,” says Emilia Lim, a bioinformatics expert and a co-first author of the Crick study. “It is the carcinogen that promotes a conducive environment for this one little clone to grow and expand into cancer. Through our work, we were able to revive excitement for this hypothesis and bring it to light.”
The study explains a confusing pattern of lung cancer developing, particularly in women, despite a lack of environmental risk factors like smoking, secondhand smoke, or radon exposure. The culprit in these cases may have been too much PM 2.5 exposure.
Other researchers had previously identified a link between mutations in certain genes that control epidermal growth factor receptors, or EGFR mutations, and the development of NSCLC. In a 2019 study of 250 people with this type of cancer, about 32 percent had the mutation. Women are more likely to have EGFR mutations than men.
Not everyone who has the EGFR mutation will develop lung cancer. Respirologist Stephen Lam studies lung cancer at the BC Cancer Research Centre in Vancouver, Canada, but was not involved in the Crick Institute research. He says the study explains a confusing pattern of lung cancer developing, particularly in women, despite a lack of environmental risk factors like smoking, secondhand smoke, or radon exposure. The culprit in these cases may have been too much PM 2.5 exposure.
More exposure leads to inflammation and lesions
The Crick researchers found that an excess of PM 2.5 in the air sparked an inflammatory process in cells within the lung. This inflammation set the stage for NSCLC to develop in people and mice with existing EGFR mutations.
The researchers also exposed mice without EGFR mutations to PM 2.5 pollution—an experiment that couldn’t be ethically conducted in humans—to link pollution exposure to NSCLC. The mice experiments also showed that NSCLC is dose-dependent; higher levels of exposure were associated with higher number of cancerous lesions forming.
Ultimately, the study “fundamentally changed how we view lung cancer in people who have never smoked,” said Swanton in a Crick Institute press release. “Cells with cancer-causing mutations accumulate naturally as we age, but they are normally inactive. We’ve demonstrated that air pollution wakes these cells up in the lungs, encouraging them to grow and potentially form tumors.”
Preventing cancer before it begins
Targeted therapies already exist for people with EGFR mutations who’ve developed NSCLC, but they have many side effects, according to Weeden. Researchers hope that making more definitive links between pollutants and cancer could help prevent people with EGFR or other mutations from developing lung cancer in the first place.
Along those lines, as an additional component of their study presented last month, the Crick researchers were able to prevent cancer in mice that had the EGFR mutations by blocking inflammation. They used an antibody to inhibit a protein called interleukin 1 beta, which plays a key role in inflammation. Scientists could eventually use such antibodies or other therapies to make a drug treatment that people can take to stop cancer in its tracks, even if they live in highly polluted areas.
Such potential could reach beyond lung cancer; in the past, Crick and other researchers have also found associations between exposure to air pollution and mesothelioma, as well as cancers of the small intestine, lip, mouth, and throat. These links could be meaningful to a growing number of people as climate change intensifies, and with increases in air pollution from fossil fuel combustion and natural disasters like forest fires.
Plus, air pollution is just one external condition that can flip the switch of these inflammatory pathways. Identifying a link between pollution and cancer “has wide ramifications for many other environmental factors that may [play] similar roles,” Weedon says. She hopes that the Crick study and future research in this area will offer some hope for non-smokers frustrated by cancer diagnoses.