New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
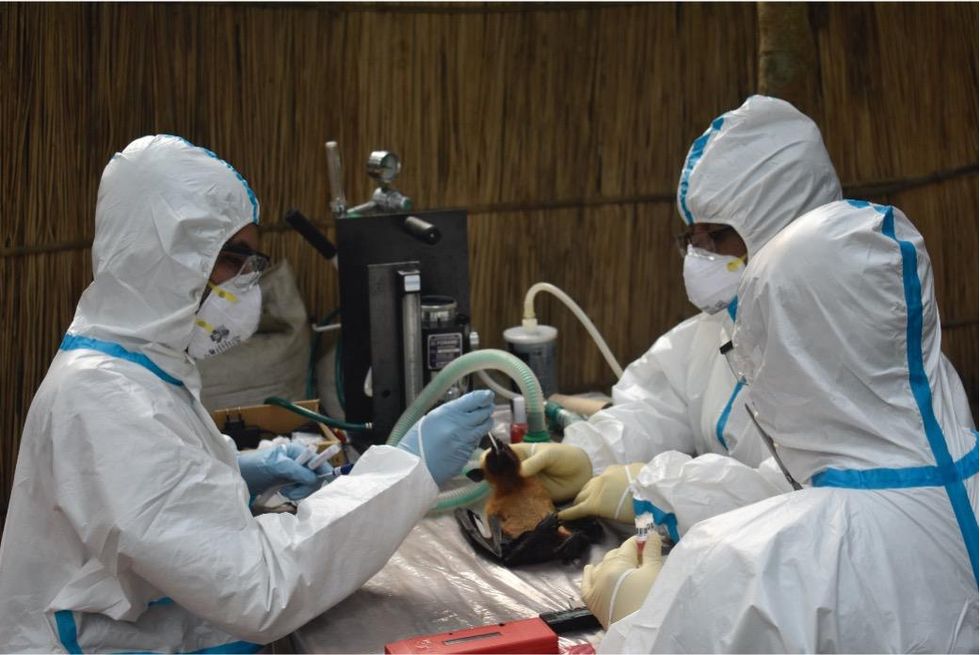
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
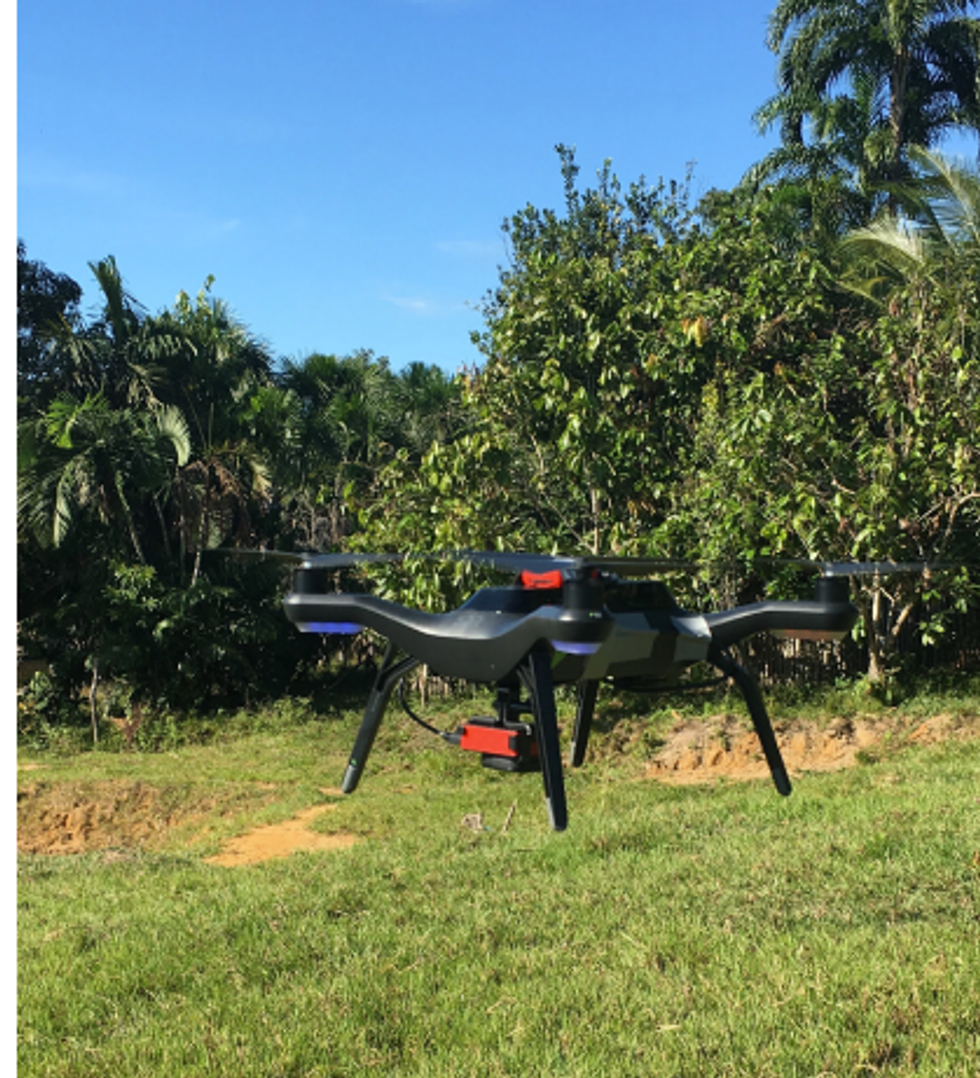

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Did researchers finally find a way to lick COVID?
A professor of medicine at the University of Michigan is researching whether lactoferrin, which is found in dairy products such as ice cream, can help to prevent COVID-19 infections.
Already vaccinated and want more protection from COVID-19? A protein found in ice cream could help, some research suggests, though there are a bunch of caveats.
The protein, called lactoferrin, is found in the milk of mammals and thus in dairy products, including ice cream. It has astounding antiviral properties that have been taken for granted and remain largely unexplored because it is a natural product, meaning that it cannot be patented and exploited by pharmaceutical companies.
Still, a few researchers in Europe and elsewhere have sought to better understand the compound.
Jonathan Sexton runs a drug screening program at the University of Michigan where cells are infected with a pathogen and then exposed to a library of the thousands of small molecule drug compounds – which can enter the body more easily than drugs with heavier molecules – approved by the FDA. In addition, the library includes compounds that passed phase 1 safety studies but later proved ineffective against the targeted disease. Each drug is dissolved in a solvent for exposure to the cells in the laborious testing process made feasible by robotic automation.
When COVID hit, researchers scrambled to identify any approved drug that might help fight the infection. Sexton decided to screen the drug library as well as some dietary supplements against SARS-CoV-2, the virus that causes the disease. Sexton says that the grunt work fell to Jesse Wotring, “a very talented PhD student,” who pulled lactoferrin off the shelf. But the regular solvent used in the testing process would destroy the protein, so he had to take another approach and do all the work by hand.
“We were agnostic,” says Sexton, who didn't have a strong interest in lactoferrin or any of the other compounds in the library, but the data was quite clear; lactoferrin “consistently produced the best efficacy...it was the absolute home run.” The findings were published in separate papers last year and in February.
It turns out that lactoferrin has several different mechanisms of action against SARS-CoV-2, inhibiting the virus from entering cells, moving around within them and replicating. Lactoferrin also modulates the overall immune response, which makes it difficult for the virus to simultaneously mutate resistance to the protein at every step of replication. “It has broad efficacy against every [SARS-CoV-2] variant that we've tested,” he says.
From bench to bedside
Sexton's initial interest was to develop a drug for the acute phase of COVID infection, to treat a hospitalized patient or prevent that hospitalization. But with the quick approval of vaccines and drugs to treat the disease, he increasingly focused on ways to better prevent infection and inhibit spread of the virus.
“If you can get lactoferrin to persist in your upper GI tract, then it may very well prevent the primary infection, and that's what we're really interested in.” He reasoned that a chewing gum formula might release enough lactoferrin into the mucosal tissue of the mouth and upper airways to inhibit replication and give the immune system a chance to knock out the virus before it can establish a foothold. It could also reduce the amount of virus spread through talking.
To get enough lactoferrin to have a possible beneficial effect, one would have to drink gallons of milk a day, “and that would have other undesirable consequences, like getting extremely obese,” says Sexton. Obesity is one of the leading risk factors for severe COVID disease.
Testing that theory has been difficult. The easiest way would be a “challenge trial,” where volunteers take the drug, or in this case gum, are exposed to the pathogen, and protection is measured. Some COVID challenge studies have been conducted in Europe but the FDA remains hesitant to allow such a study in the U.S. A traditional prevention study would be like a vaccine trial, involving thousands, perhaps tens of thousands of volunteers over a period of months or years, and it would be very expensive. No one has stepped forward to foot the bill.
So the next step for Sexton is a clinical trial of newly diagnosed COVID patients who will be given standard of care treatment, and layered on top of that they will receive either lactoferrin, probably in pill form, or a placebo. He has identified initial funding. “We would study their viral load over time as well as their symptoms.”
One issue the FDA is grappling with in considering the proposed trial is that it typically decides whether to approve drugs from a factory by applying a rigorous standard, called good manufacturing practices, while food products, which are the source of lactoferrin, are produced under somewhat different standards. The agency still has not finalized rules on how to deal with natural products used as drugs, such as fecal transplants, convalescent plasma, or medical marijuana.
Sexton is frustrated by the delay because lactoferrin derived from bovine milk whey has been used for many decades as a protein supplement by athletes, it is a large component of most infant formula, and the largest number of clinical studies of lactoferrin involve premature infants. There is no question of its safety, he says.
Do it yourself
So what can you do while waiting for regulatory wheels to spin and clinical trial data to be generated?
Could a dose of Ben & Jerry's provide some protection against SARS-CoV-2?
Sexton chuckles at the suggestion. He supposes it couldn't hurt. But to get enough lactoferrin to have a possible beneficial effect, one would have to drink gallons of milk a day, “and that would have other undesirable consequences, like getting extremely obese.” Obesity is one of the leading risk factors for severe COVID disease.
Pseudo-milk products made from soy, almonds, oats, or other plant products do not contain lactoferrin; it has to come from a teat. So that rules them out.
Whey-based protein shakes might be a useful way to add lactoferrin to the diet.
Probably the best option is to take conventional gelatin capsules of lactoferrin that are widely available wherever supplements are sold. Sexton calculates that about a gram a day, four 250 milligram capsules, should do it. He advises two in the morning and two a night. “You really want to take them on an empty stomach...your stomach treats [the lactoferrin protein] like it would a steak” and chops it for absorption in the intestine, which you do not want. About 70 percent of lactoferrin can get through an empty stomach, but eating food cranks up digestive gastric acids and the amount of intact lactoferrin that gets through to the gut plummets.
Sexton cautions, “We have not determined clinical efficacy yet,” and he is not offering advice as a physician, but in the spirit of harm reduction, he realizes that some people are going to try things that might help them. Lactoferrin “is remarkably safe. And so people have to make their own decisions about what they are willing to take and what they are not,” he says.
In today's episode, Leaps.org interviews Camila dos Santos, a molecular biologist at Cold Spring Harbor Lab, about her research on breasts and what makes them unique compared to any other part of the body.
My guest today for the Making Sense of Science podcast is Camila dos Santos, associate professor at Cold Spring Harbor Lab, who is a leading researcher of the inner lives of human mammary glands, more commonly known as breasts. These organs are unlike any other because throughout life they undergo numerous changes, first in puberty, then during pregnancies and lactation periods, and finally at the end of the cycle, when babies are weaned. A complex interplay of hormones governs these processes, in some cases increasing the risk of breast cancer and sometimes lowering it. Witnessing the molecular mechanics behind these processes in humans is not possible, so instead Dos Santos studies organoids—the clumps of breast cells donated by patients who undergo breast reduction surgeries or biopsies.
Show notes:
2:52 In response to hormones that arise during puberty, the breast cells grow and become more specialized, preparing the tissue for making milk.
7:53 How do breast cells know when to produce milk? It’s all governed by chemical messaging in the body. When the baby is born, the brain will release the hormone called oxytocin, which will make the breast cells contract and release the milk.
12:40 Breast resident immune cells are including T-cells and B-cells, but because they live inside the breast tissue their functions differ from the immune cells in other parts of the body,
17:00 With organoids—dimensional clumps of cells that are cultured in a dish—it is possible to visualize and study how these cells produce milk.
21:50 Women who are pregnant later in life are more likely to require medical intervention to breastfeed. Scientists are trying to understand the fundamental reasons why it happens.
26:10 Breast cancer has many risks factors. Generic mutations play a big role. All of us have the BRCA genes, but it is the alternation in the DNA sequence of the BRCA gene that can increase the predisposition to breast cancer. Aging and menopause are the risk factors for breast cancer, and so are pregnancies.
29:22 Women that are pregnant before the age of 20 to 25, have a decreased risk of breast cancer. And the hypothesis here is that during pregnancy breast cells more specialized, as specialized cells, they have a limited lifespan. It's more likely that they die before they turn into cancer.
33:08 Organoids are giving scientists an opportunity to practice personalized medicine. Scientists can test drugs on organoids taken from a patient to identify the most efficient treatment protocol.
Links:
Camila dos Santos’s Lab Page.
Editor's note: In addition to being a regular writer for Leaps.org, Lina Zeldovich is the guest host for today's episode of the Making Sense of Science podcast.
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.

