New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
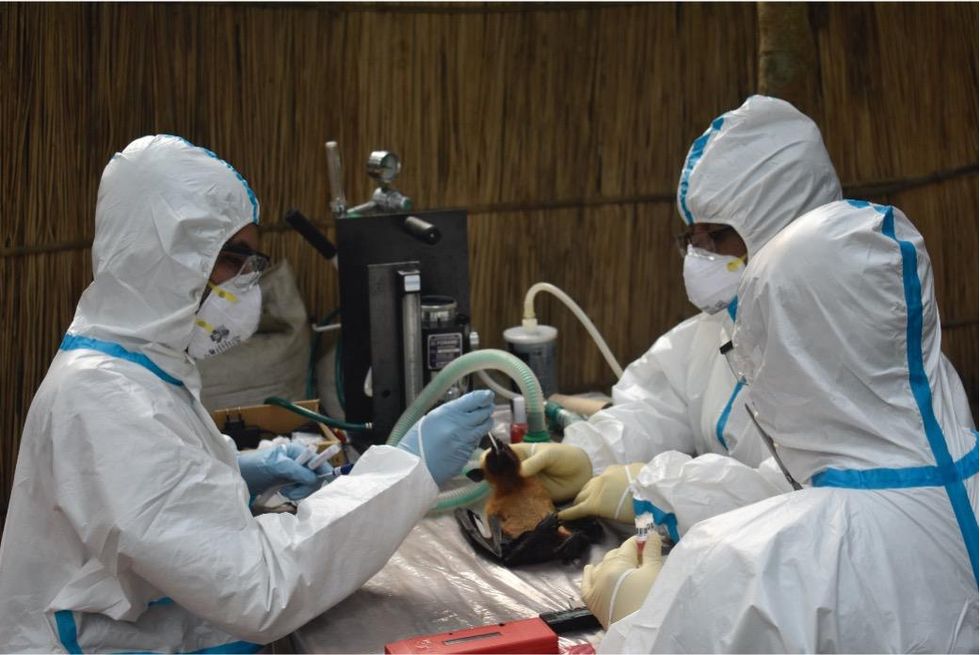
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
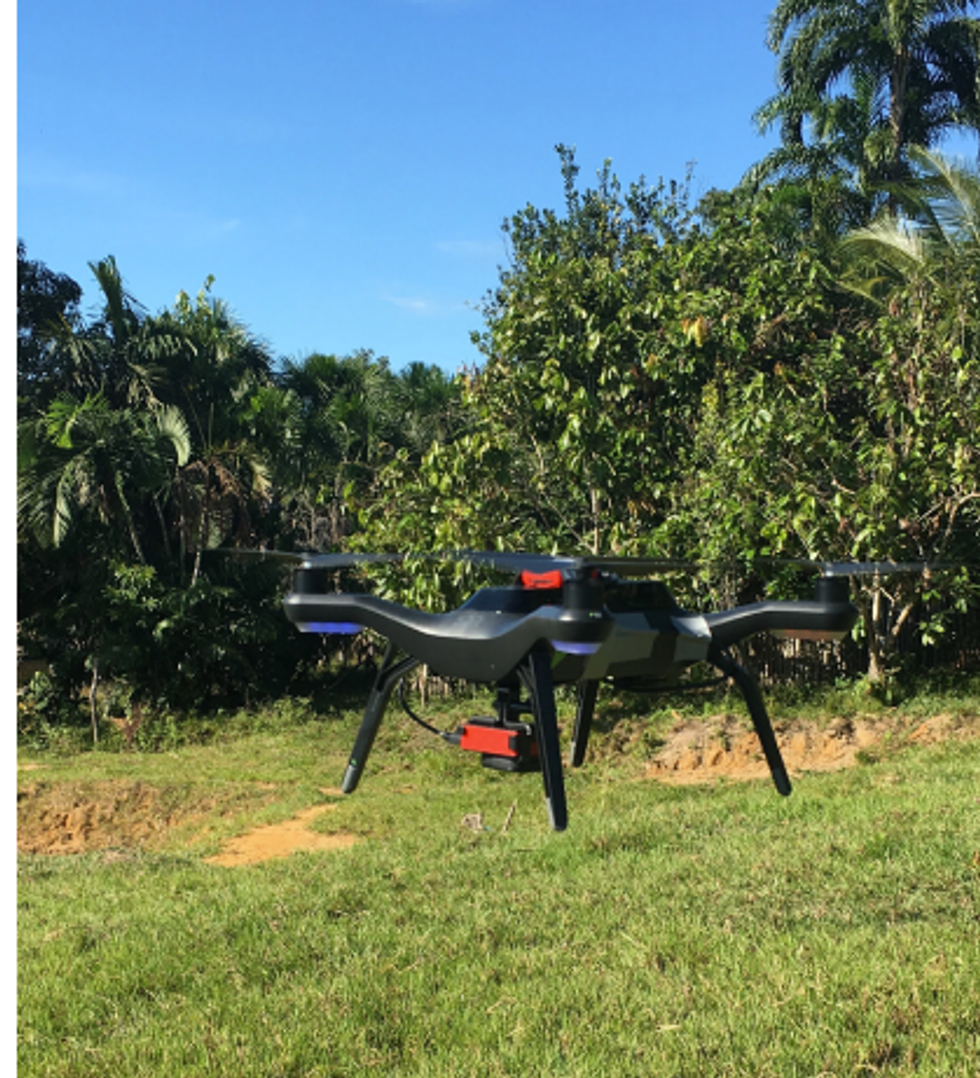

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Waste smothering our oceans is worth billions – here’s what we can do with all that sh$t
In 2015, human poop was valued at $9.5 billion per year, which today would be $11.5 billion. The Ocean Sewage Alliance is uniting experts from key sectors to change how we handle our sewage and, in the process, create all sorts of economic benefits.
There’s hardly a person out there who hasn’t heard of the Great Pacific Garbage Patch. That type of pollution is impossible to miss. It stares you in the face from pictures and videos of sea turtles with drinking straws up their noses and acres of plastic swirling in the sea.
It demands you to solve the problem—and it works. The campaign to raise awareness about plastic pollution in the oceans has resulted in new policies, including bans on microplastics in personal care products, technology to clean up the plastic, and even new plastic-like materials that are better for the environment.
But there’s a different type of pollution smothering the ocean as you read this. Unfortunately, this one is almost invisible, but no less damaging. In fact, it’s even more serious than plastic and most people have no idea it even exists. It is literally under our noses, destroying our oceans, lakes, and rivers – and yet we are missing it completely while contributing to it daily. In fact, we exacerbate it multiple times a day—every time we use the bathroom.
It is the way we do our sewage.
Most of us don’t think much about what happens after we flush the toilet. Most of us probably assume that the substances we flush go “somewhere” and are dealt with safely. But we typically don’t think about it beyond that.
Most of us also probably don’t think about what’s in the ocean or lakes we swim in. Since others are swimming, jumping in is just fine. But our waterways are far from clean. In fact, at times they are incredibly filthy. In the US, we are dumping 1.2 trillion of gallons of untreated sewage into the environment every year. Just New York City alone discharges 27 billion gallons into the Hudson River basin annually.
How does this happen? Part of it is the unfortunate side effect of our sewage system design that dates back to over a century ago when cities were smaller and fewer people were living so close together.
Back then, engineers designed the so-called “combine sewer overflow systems,” or CSOs, in which the storm water pipes are connected to the sanitary sewer pipes. In normal conditions, the sewage effluent from homes flows to the treatment plants where it gets cleaned and released into the waterways. But when it rains, the pipe system becomes so overwhelmed with water that the treatment plant can’t process it fast enough. So the treatment plant has to release the excess water through its discharge pipes—directly, without treatment, into streams, rivers and the ocean.
The 1.2 trillion gallons of CSO releases isn’t even the full picture. There are also discharges from poorly maintained septic systems, cesspools and busted pipes of the aging wastewater infrastructure. The state of Hawaii alone has 88,000 cesspools that need replacing and are currently leaking 53 million gallons of raw sewage daily into their coastal waters. You may think twice about swimming on your Hawaii vacations.
Overall, the US is facing a $271 billion backlog in wastewater infrastructure projects to update these aging systems. Across the Western world, countries are facing similar challenges with their aging sewage systems, especially the UK and European Union.
That’s not to say that other parts of the planet are in better shape. Out of the 7+ billion people populating our earth, 4.2 billion don’t have access to safe sanitation. Included in this insane number are roughly 2 billion people who have no toilet at all. Whether washed by rains or dumped directly into the waterways, a lot of this sludge pollutes the environment, the drinking water, and ultimately the ocean.
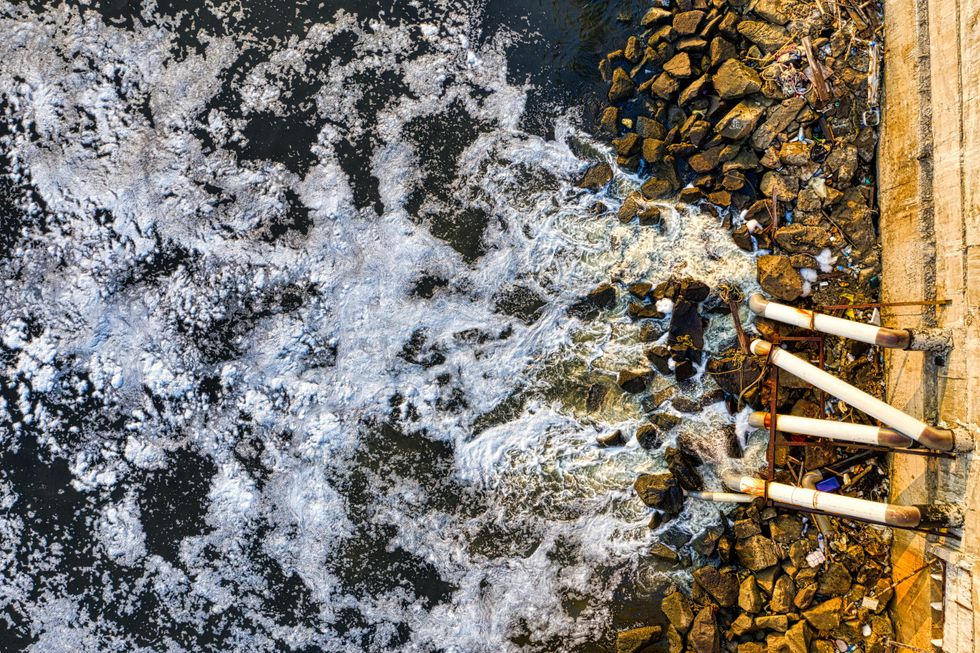

Pipes pour water onto a rocky shore in Jakarta, Indonesia.
Tom Fisk
What complicates this from an ocean health perspective is that it’s not just poop and pee that gets dumped into nearby waterways. It is all the things we put in and on our bodies and flush down our drains. That vicious mix of chemicals includes caffeine, antibiotics, antidepressants, painkillers, hormones, microplastics, cocaine, cooking oils, paint thinners, and PFAS—the forever chemicals present in everything from breathable clothing to fire retardant fabrics of our living room couches. Recent reports have found all of the above substances in fish—and then some.
Why do we allow so much untreated sewage spill into the sea? Frankly speaking, for decades scientists and engineers thought that the ocean could handle it. The mantra back then was “dilution is the solution to pollution,” which might’ve worked when there were much fewer people living on earth—but not now. Today science is telling us that this old approach doesn’t hold. That marine habitats are much more sensitive than we had expected and can’t handle the amount of wastewater we are discharging into them.
The excess nitrogen and phosphorus that the sewage (and agricultural runoff) dumps into the water causes harmful algal blooms, more commonly known as red or brown tides. The water column is overtaken by tiny algae that sucks up all the oxygen from the water, creating dead zones like the big fish kills in the Gulf of Mexico. These algae also cause public health issues by releasing gases toxic to people and animals, including dementia, neurological damage, and respiratory illness. Marshes and mangroves end up with weakened root systems and start dying off. In a wastewater modeling study I published last year, we found that 31 percent of salt marshes globally were heavily polluted with human sewage. Coral reefs get riddled with disease and overgrown by seaweed.
We could convert sewage into high-value goods. It can be used to generate electricity, fertilizer, and drinking water. The technologies not only exist but are getting better and more efficient all the time.
Moreover, by way of our sewage, we managed to transmit a human pathogen—Serratia marcescens, which causes urinary, respiratory and other infections in people—to corals! Recent reports from the Florida Keys are showing white pox disease popping up in elk horn corals caused by S.marcescens, which somehow managed to jump species. Many recent studies have documented just how common this type of pollution is across the globe.
Yet, there is some good news in that abysmal sewage flow. Just like with plastic pollution, realizing that there’s a problem is the first step, so awareness is key. That’s exactly why I co-founded Ocean Sewage Alliance last year—a nonprofit that aims to “re-potty train the world” by breaking taboos in talking about the poop and pee problem, as well as uniting experts from various key sectors to work together to end sewage pollution in coastal areas.
To end this pollution, we have to change the ways we handle our sewage. Even more exciting is that by solving the sewage problem we can create all sorts of economic benefits. In 2015, human poop was valued at $9.5 billion a year globally, which today would be $11.5 billion per year.
What would one do with that sh$t?
We could convert it into high-value goods. Sewage can be used to generate electricity, fertilizer, and drinking water. The technologies not only exist but are getting better and more efficient all the time. Some exciting examples include biodigesters and urine diversion (or peecycling) systems that can produce fertilizer and biogas, essentially natural gas. The United Nations estimates that the biogas produced from poop could provide electricity for 138 million homes. And the recovered and cleaned water can be used for irrigation, laundry and flushing toilets. It can even be refined to the point that it is safe for drinking water – just ask the folks in Orange County, CA who have been doing so for the last few decades.
How do we deal with all the human-made pollutants in our sewage? There is technology for that too. Called pyrolysis, it heats up sludge to high temperatures in the absence of oxygen, which causes most of the substances to degrade and fall apart.
There are solutions to the problems—as long as we acknowledge that the problems exist. The fact that you are reading this means that you are part of the solution already. The next time you flush your toilet, think about where this output may flow. Does your septic system work properly? Does your local treatment plant discharge raw sewage on rainy days? Can that plant implement newer technologies that can upcycle waste? These questions are part of re-potty training the world, one household at a time. And together, these households are the force that can turn back the toxic sewage tide. And keep our oceans blue.
The U.S. must fund more biotech innovation – or other countries will catch up faster than you think
In the coming years, U.S. market share in biotech will decline unless the federal government makes investments to improve the quality and quantity of U.S. research, writes the author.
The U.S. has approximately 58 percent of the market share in the biotech sector, followed by China with 11 percent. However, this market share is the result of several years of previous research and development (R&D) – it is a present picture of what happened in the past. In the future, this market share will decline unless the federal government makes investments to improve the quality and quantity of U.S. research in biotech.
The effectiveness of current R&D can be evaluated in a variety of ways such as monies invested and the number of patents filed. According to the UNESCO Institute for Statistics, the U.S. spends approximately 2.7 percent of GDP on R&D ($476,459.0M), whereas China spends 2 percent ($346,266.3M). However, investment levels do not necessarily translate into goods that end up contributing to innovation.
Patents are a better indication of innovation. The biotech industry relies on patents to protect their investments, making patenting a key tool in the process of translating scientific discoveries that can ultimately benefit patients. In 2020, China filed 1,497,159 patents, a 6.9 percent increase in growth rate. In contrast, the U.S. filed 597,172, a 3.9 percent decline. When it comes to patents filed, China has approximately 45 percent of the world share compared to 18 percent for the U.S.
So how did we get here? The nature of science in academia allows scientists to specialize by dedicating several years to advance discovery research and develop new inventions that can then be licensed by biotech companies. This makes academic science critical to innovation in the U.S. and abroad.
Academic scientists rely on government and foundation grants to pay for R&D, which includes salaries for faculty, investigators and trainees, as well as monies for infrastructure, support personnel and research supplies. Of particular interest to academic scientists to cover these costs is government support such as Research Project Grants, also known as R01 grants, the oldest grant mechanism from the National Institutes of Health. Unfortunately, this funding mechanism is extremely competitive, as applications have a success rate of only about 20 percent. To maximize the chances of getting funded, investigators tend to limit the innovation of their applications, since a project that seems overambitious is discouraged by grant reviewers.
Considering the difficulty in obtaining funding, the limited number of opportunities for scientists to become independent investigators capable of leading their own scientific projects, and the salaries available to pay for scientists with a doctoral degree, it is not surprising that the U.S. is progressively losing its workforce for innovation.
This approach affects the future success of the R&D enterprise in the U.S. Pursuing less innovative work tends to produce scientific results that are more obvious than groundbreaking, and when a discovery is obvious, it cannot be patented, resulting in fewer inventions that go on to benefit patients. Even though there are governmental funding options available for scientists in academia focused on more groundbreaking and translational projects, those options are less coveted by academic scientists who are trying to obtain tenure and long-term funding to cover salaries and other associated laboratory expenses. Therefore, since only a small percent of projects gets funded, the likelihood of scientists interested in pursuing academic science or even research in general keeps declining over time.
Efforts to raise the number of individuals who pursue a scientific education are paying off. However, the number of job openings for those trainees to carry out independent scientific research once they graduate has proved harder to increase. These limitations are not just in the number of faculty openings to pursue academic science, which are in part related to grant funding, but also the low salary available to pay those scientists after they obtain their doctoral degree, which ranges from $53,000 to $65,000, depending on years of experience.
Thus, considering the difficulty in obtaining funding, the limited number of opportunities for scientists to become independent investigators capable of leading their own scientific projects, and the salaries available to pay for scientists with a doctoral degree, it is not surprising that the U.S. is progressively losing its workforce for innovation, which results in fewer patents filed.
Perhaps instead of encouraging scientists to propose less innovative projects in order to increase their chances of getting grants, the U.S. government should give serious consideration to funding investigators for their potential for success -- or the success they have already achieved in contributing to the advancement of science. Such a funding approach should be tiered depending on career stage or years of experience, considering that 42 years old is the median age at which the first R01 is obtained. This suggests that after finishing their training, scientists spend 10 years before they establish themselves as independent academic investigators capable of having the appropriate funds to train the next generation of scientists who will help the U.S. maintain or even expand its market share in the biotech industry for years to come. Patenting should be given more weight as part of the academic endeavor for promotion purposes, or governmental investment in research funding should be increased to support more than just 20 percent of projects.
Remaining at the forefront of biotech innovation will give us the opportunity to not just generate more jobs, but it will also allow us to attract the brightest scientists from all over the world. This talented workforce will go on to train future U.S. scientists and will improve our standard of living by giving us the opportunity to produce the next generation of therapies intended to improve human health.
This problem cannot rely on just one solution, but what is certain is that unless there are more creative changes in funding approaches for scientists in academia, eventually we may be saying “remember when the U.S. was at the forefront of biotech innovation?”

