The Shiny–and Potentially Dangerous—New Tool for Predicting Human Behavior

Studies of twins have played an important role in determining that genetic differences play a role in the development of differences in behavior.
[Editor's Note: This essay is in response to our current Big Question, which we posed to experts with different perspectives: "How should DNA tests for intelligence be used, if at all, by parents and educators?"]
Imagine a world in which pregnant women could go to the doctor and obtain a simple inexpensive genetic test of their unborn child that would allow them to predict how tall he or she would eventually be. The test might also tell them the child's risk for high blood pressure or heart disease.
Can we use DNA not to understand, but to predict who is going to be intelligent or extraverted or mentally ill?
Even more remarkable -- and more dangerous -- the test might predict how intelligent the child would be, or how far he or she could be expected to go in school. Or heading further out, it might predict whether he or she will be an alcoholic or a teetotaler, or straight or gay, or… you get the idea. Is this really possible? If it is, would it be a good idea? Answering these questions requires some background in a scientific field called behavior genetics.
Differences in human behavior -- intelligence, personality, mental illness, pretty much everything -- are related to genetic differences among people. Scientists have known this for 150 years, ever since Darwin's half-cousin Francis Galton first applied Shakespeare's phrase, "Nature and Nurture" to the scientific investigation of human differences. We knew about the heritability of behavior before Mendel's laws of genetics had been re-discovered at the end of the last century, and long before the structure of DNA was discovered in the 1950s. How could discoveries about genetics be made before a science of genetics even existed?
The answer is that scientists developed clever research designs that allowed them to make inferences about genetics in the absence of biological knowledge about DNA. The best-known is the twin study: identical twins are essentially clones, sharing 100 percent of their DNA, while fraternal twins are essentially siblings, sharing half. To the extent that identical twins are more similar for some trait than fraternal twins, one can infer that heredity is playing a role. Adoption studies are even more straightforward. Is the personality of an adopted child more like the biological parents she has never seen, or the adoptive parents who raised her?
Twin and adoption studies played an important role in establishing beyond any reasonable doubt that genetic differences play a role in the development of differences in behavior, but they told us very little about how the genetics of behavior actually worked. When the human genome was finally sequenced in the early 2000s, and it became easier and cheaper to obtain actual DNA from large samples of people, scientists anticipated that we would soon find the genes for intelligence, mental illness, and all the other behaviors that were known to be "heritable" in a general way.
But to everyone's amazement, the genes weren't there. It turned out that there are thousands of genes related to any given behavior, so many that they can't be counted, and each one of them has such a tiny effect that it can't be tied to meaningful biological processes. The whole scientific enterprise of understanding the genetics of behavior seemed ready to collapse, until it was rescued -- sort of -- by a new method called polygenic scores, PGS for short. Polygenic scores abandon the old task of finding the genes for complex human behavior, replacing it with black-box prediction: can we use DNA not to understand, but to predict who is going to be intelligent or extraverted or mentally ill?
Prediction from observing parents works better, and is far easier and cheaper, than anything we can do with DNA.
PGS are the shiny new toy of human genetics. From a technological standpoint they are truly amazing, and they are useful for some scientific applications that don't involve making decisions about individual people. We can obtain DNA from thousands of people, estimate the tiny relationships between individual bits of DNA and any outcome we want — height or weight or cardiac disease or IQ — and then add all those tiny effects together into a single bell-shaped score that can predict the outcome of interest. In theory, we could do this from the moment of conception.
Polygenic scores for height already work pretty well. Physicians are debating whether the PGS for heart disease are robust enough to be used in the clinic. For some behavioral traits-- the most data exist for educational attainment -- they work well enough to be scientifically interesting, if not practically useful. For traits like personality or sexual orientation, the prediction is statistically significant but nowhere close to practically meaningful. No one knows how much better any of these predictions are likely to get.
Without a doubt, PGS are an amazing feat of genomic technology, but the task they accomplish is something scientists have been able to do for a long time, and in fact it is something that our grandparents could have done pretty well. PGS are basically a new way to predict a trait in an individual by using the same trait in the individual's parents — a way of observing that the acorn doesn't fall far from the tree.
The children of tall people tend to be tall. Children of excellent athletes are athletic; children of smart people are smart; children of people with heart disease are at risk, themselves. Not every time, of course, but that is how imperfect prediction works: children of tall parents vary in their height like anyone else, but on average they are taller than the rest of us. Prediction from observing parents works better, and is far easier and cheaper, than anything we can do with DNA.
But wait a minute. Prediction from parents isn't strictly genetic. Smart parents not only pass on their genes to their kids, but they also raise them. Smart families are privileged in thousands of ways — they make more money and can send their kids to better schools. The same is true for PGS.
The ability of a genetic score to predict educational attainment depends not only on examining the relationship between certain genes and how far people go in school, but also on every personal and social characteristic that helps or hinders education: wealth, status, discrimination, you name it. The bottom line is that for any kind of prediction of human behavior, separation of genetic from environmental prediction is very difficult; ultimately it isn't possible.
Still, experts are already discussing how to use PGS to make predictions for children, and even for embryos.
This is a reminder that we really have no idea why either parents or PGS predict as well or as poorly as they do. It is easy to imagine that a PGS for educational attainment works because it is summarizing genes that code for efficient neurological development, bigger brains, and swifter problem solving, but we really don't know that. PGS could work because they are associated with being rich, or being motivated, or having light skin. It's the same for predicting from parents. We just don't know.
Still, experts are already discussing how to use PGS to make predictions for children, and even for embryos.
For example, maybe couples could fertilize multiple embryos in vitro, test their DNA, and select the one with the "best" PGS on some trait. This would be a bad idea for a lot of reasons. Such scores aren't effective enough to be very useful to parents, and to the extent they are effective, it is very difficult to know what other traits might be selected for when parents try to prioritize intelligence or attractiveness. People will no doubt try it anyway, and as a matter of reproductive freedom I can't think of any way to stop them. Fortunately, the practice probably won't have any great impact one way or another.
That brings us to the ethics of PGS, particularly in the schools. Imagine that when a child enrolls in a public school, an IQ test is given to her biological parents. Children with low-IQ parents are statistically more likely to have low IQs themselves, so they could be assigned to less demanding classrooms or vocational programs. Hopefully we agree that this would be unethical, but let's think through why.
First of all, it would be unethical because we don't know why the parents have low IQs, or why their IQs predict their children's. The parents could be from a marginalized ethnic group, recognizable by their skin color and passed on genetically to their children, so discriminating based on a parent's IQ would just be a proxy for discriminating based on skin color. Such a system would be no more than a social scientific gloss on an old-fashioned program for perpetuating economic and cognitive privilege via the educational system.
People deserve to be judged on the basis of their own behavior, not a genetic test.
Assigning children to classrooms based on genetic testing would be no different, although it would have the slight ethical advantage of being less effective. The PGS for educational attainment could reflect brain-efficiency, but it could also depend on skin color, or economic advantage, or personality, or literally anything that is related in any way to economic success. Privileging kids with higher genetic scores would be no different than privileging children with smart parents. If schools really believe that a psychological trait like IQ is important for school placement, the sensible thing is to administer the children an actual IQ test – not a genetic test.
IQ testing has its own issues, of course, but at least it involves making decisions about individuals based on their own observable characteristics, rather than on characteristics of their parents or their genome. If decisions must be made, if resources must be apportioned, people deserve to be judged on the basis of their own behavior, the content of their character. Since it can't be denied that people differ in all sorts of relevant ways, this is what it means for all people to be created equal.
[Editor's Note: Read another perspective in the series here.]
Jamie Rettinger with his now fiance Amie Purnel-Davis, who helped him through the clinical trial.
Jamie Rettinger was still in his thirties when he first noticed a tiny streak of brown running through the thumbnail of his right hand. It slowly grew wider and the skin underneath began to deteriorate before he went to a local dermatologist in 2013. The doctor thought it was a wart and tried scooping it out, treating the affected area for three years before finally removing the nail bed and sending it off to a pathology lab for analysis.
"I have some bad news for you; what we removed was a five-millimeter melanoma, a cancerous tumor that often spreads," Jamie recalls being told on his return visit. "I'd never heard of cancer coming through a thumbnail," he says. None of his doctors had ever mentioned it either. "I just thought I was being treated for a wart." But nothing was healing and it continued to bleed.
A few months later a surgeon amputated the top half of his thumb. Lymph node biopsy tested negative for spread of the cancer and when the bandages finally came off, Jamie thought his medical issues were resolved.
Melanoma is the deadliest form of skin cancer. About 85,000 people are diagnosed with it each year in the U.S. and more than 8,000 die of the cancer when it spreads to other parts of the body, according to the Centers for Disease Control and Prevention (CDC).
There are two peaks in diagnosis of melanoma; one is in younger women ages 30-40 and often is tied to past use of tanning beds; the second is older men 60+ and is related to outdoor activity from farming to sports. Light-skinned people have a twenty-times greater risk of melanoma than do people with dark skin.
"When I graduated from medical school, in 2005, melanoma was a death sentence" --Diwakar Davar.
Jamie had a follow up PET scan about six months after his surgery. A suspicious spot on his lung led to a biopsy that came back positive for melanoma. The cancer had spread. Treatment with a monoclonal antibody (nivolumab/Opdivo®) didn't prove effective and he was referred to the UPMC Hillman Cancer Center in Pittsburgh, a four-hour drive from his home in western Ohio.
An alternative monoclonal antibody treatment brought on such bad side effects, diarrhea as often as 15 times a day, that it took more than a week of hospitalization to stabilize his condition. The only options left were experimental approaches in clinical trials.
Early research
"When I graduated from medical school, in 2005, melanoma was a death sentence" with a cure rate in the single digits, says Diwakar Davar, 39, an oncologist at UPMC Hillman Cancer Center who specializes in skin cancer. That began to change in 2010 with introduction of the first immunotherapies, monoclonal antibodies, to treat cancer. The antibodies attach to PD-1, a receptor on the surface of T cells of the immune system and on cancer cells. Antibody treatment boosted the melanoma cure rate to about 30 percent. The search was on to understand why some people responded to these drugs and others did not.
At the same time, there was a growing understanding of the role that bacteria in the gut, the gut microbiome, plays in helping to train and maintain the function of the body's various immune cells. Perhaps the bacteria also plays a role in shaping the immune response to cancer therapy.
One clue came from genetically identical mice. Animals ordered from different suppliers sometimes responded differently to the experiments being performed. That difference was traced to different compositions of their gut microbiome; transferring the microbiome from one animal to another in a process known as fecal transplant (FMT) could change their responses to disease or treatment.
When researchers looked at humans, they found that the patients who responded well to immunotherapies had a gut microbiome that looked like healthy normal folks, but patients who didn't respond had missing or reduced strains of bacteria.
Davar and his team knew that FMT had a very successful cure rate in treating the gut dysbiosis of Clostridioides difficile, a persistant intestinal infection, and they wondered if a fecal transplant from a patient who had responded well to cancer immunotherapy treatment might improve the cure rate of patients who did not originally respond to immunotherapies for melanoma.
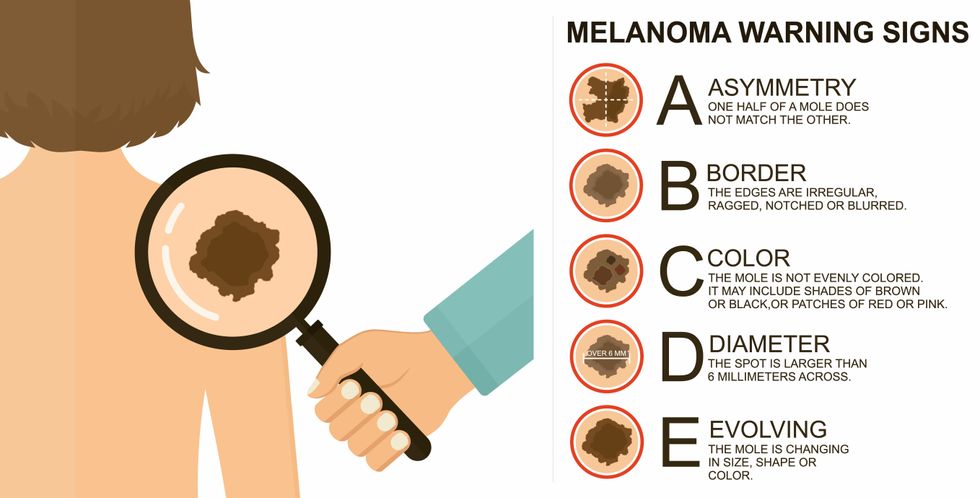
The ABCDE of melanoma detection
Adobe Stock
Clinical trial
"It was pretty weird, I was totally blasted away. Who had thought of this?" Jamie first thought when the hypothesis was explained to him. But Davar's explanation that the procedure might restore some of the beneficial bacterial his gut was lacking, convinced him to try. He quickly signed on in October 2018 to be the first person in the clinical trial.
Fecal donations go through the same safety procedures of screening for and inactivating diseases that are used in processing blood donations to make them safe for transfusion. The procedure itself uses a standard hollow colonoscope designed to screen for colon cancer and remove polyps. The transplant is inserted through the center of the flexible tube.
Most patients are sedated for procedures that use a colonoscope but Jamie doesn't respond to those drugs: "You can't knock me out. I was watching them on the TV going up my own butt. It was kind of unreal at that point," he says. "There were about twelve people in there watching because no one had seen this done before."
A test two weeks after the procedure showed that the FMT had engrafted and the once-missing bacteria were thriving in his gut. More importantly, his body was responding to another monoclonal antibody (pembrolizumab/Keytruda®) and signs of melanoma began to shrink. Every three months he made the four-hour drive from home to Pittsburgh for six rounds of treatment with the antibody drug.
"We were very, very lucky that the first patient had a great response," says Davar. "It allowed us to believe that even though we failed with the next six, we were on the right track. We just needed to tweak the [fecal] cocktail a little better" and enroll patients in the study who had less aggressive tumor growth and were likely to live long enough to complete the extensive rounds of therapy. Six of 15 patients responded positively in the pilot clinical trial that was published in the journal Science.
Davar believes they are beginning to understand the biological mechanisms of why some patients initially do not respond to immunotherapy but later can with a FMT. It is tied to the background level of inflammation produced by the interaction between the microbiome and the immune system. That paper is not yet published.
Surviving cancer
It has been almost a year since the last in his series of cancer treatments and Jamie has no measurable disease. He is cautiously optimistic that his cancer is not simply in remission but is gone for good. "I'm still scared every time I get my scans, because you don't know whether it is going to come back or not. And to realize that it is something that is totally out of my control."
"It was hard for me to regain trust" after being misdiagnosed and mistreated by several doctors he says. But his experience at Hillman helped to restore that trust "because they were interested in me, not just fixing the problem."
He is grateful for the support provided by family and friends over the last eight years. After a pause and a sigh, the ruggedly built 47-year-old says, "If everyone else was dead in my family, I probably wouldn't have been able to do it."
"I never hesitated to ask a question and I never hesitated to get a second opinion." But Jamie acknowledges the experience has made him more aware of the need for regular preventive medical care and a primary care physician. That person might have caught his melanoma at an earlier stage when it was easier to treat.
Davar continues to work on clinical studies to optimize this treatment approach. Perhaps down the road, screening the microbiome will be standard for melanoma and other cancers prior to using immunotherapies, and the FMT will be as simple as swallowing a handful of freeze-dried capsules off the shelf rather than through a colonoscopy. Earlier this year, the Food and Drug Administration approved the first oral fecal microbiota product for C. difficile, hopefully paving the way for more.
An older version of this hit article was first published on May 18, 2021
All organisms can repair damaged tissue, but none do it better than salamanders and newts. A surprising area of science could tell us how they manage this feat - and perhaps even help us develop a similar ability.
All organisms have the capacity to repair or regenerate tissue damage. None can do it better than salamanders or newts, which can regenerate an entire severed limb.
That feat has amazed and delighted man from the dawn of time and led to endless attempts to understand how it happens – and whether we can control it for our own purposes. An exciting new clue toward that understanding has come from a surprising source: research on the decline of cells, called cellular senescence.
Senescence is the last stage in the life of a cell. Whereas some cells simply break up or wither and die off, others transition into a zombie-like state where they can no longer divide. In this liminal phase, the cell still pumps out many different molecules that can affect its neighbors and cause low grade inflammation. Senescence is associated with many of the declining biological functions that characterize aging, such as inflammation and genomic instability.
Oddly enough, newts are one of the few species that do not accumulate senescent cells as they age, according to research over several years by Maximina Yun. A research group leader at the Center for Regenerative Therapies Dresden and the Max Planck Institute of Molecular and Cell Biology and Genetics, in Dresden, Germany, Yun discovered that senescent cells were induced at some stages of regeneration of the salamander limb, “and then, as the regeneration progresses, they disappeared, they were eliminated by the immune system,” she says. “They were present at particular times and then they disappeared.”
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states.
Previous research on senescence in aging had suggested, logically enough, that applying those cells to the stump of a newly severed salamander limb would slow or even stop its regeneration. But Yun stood that idea on its head. She theorized that senescent cells might also play a role in newt limb regeneration, and she tested it by both adding and removing senescent cells from her animals. It turned out she was right, as the newt limbs grew back faster than normal when more senescent cells were included.
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states, which could then be turned into progenitors, a cell type in between stem cells and specialized cells, needed to regrow the muscle tissue of the missing limb. “We think that this ability to dedifferentiate is intrinsically a big part of why salamanders can regenerate all these very complex structures, which other organisms cannot,” she explains.
Yun sees regeneration as a two part problem. First, the cells must be able to sense that their neighbors from the lost limb are not there anymore. Second, they need to be able to produce the intermediary progenitors for regeneration, , to form what is missing. “Molecularly, that must be encoded like a 3D map,” she says, otherwise the new tissue might grow back as a blob, or liver, or fin instead of a limb.
Wound healing
Another recent study, this time at the Mayo Clinic, provides evidence supporting the role of senescent cells in regeneration. Looking closely at molecules that send information between cells in the wound of a mouse, the researchers found that senescent cells appeared near the start of the healing process and then disappeared as healing progressed. In contrast, persistent senescent cells were the hallmark of a chronic wound that did not heal properly. The function and significance of senescence cells depended on both the timing and the context of their environment.
The paper suggests that senescent cells are not all the same. That has become clearer as researchers have been able to identify protein markers on the surface of some senescent cells. The patterns of these proteins differ for some senescent cells compared to others. In biology, such physical differences suggest functional differences, so it is becoming increasingly likely there are subsets of senescent cells with differing functions that have not yet been identified.
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes.
Scientists initially thought that senescent cells couldn’t play a role in regeneration because they could no longer reproduce, says Anthony Atala, a practicing surgeon and bioengineer who leads the Wake Forest Institute for Regenerative Medicine in North Carolina. But Yun’s study points in the other direction. “What this paper shows clearly is that these cells have the potential to be involved in tissue regeneration [in newts]. The question becomes, will these cells be able to do the same in humans.”
As our knowledge of senescent cells increases, Atala thinks we need to embrace a new analogy to help understand them: humans in retirement. They “have acquired a lot of wisdom throughout their whole life and they can help younger people and mentor them to grow to their full potential. We're seeing the same thing with these cells,” he says. They are no longer putting energy into their own reproduction, but the signaling molecules they secrete “can help other cells around them to regenerate.”
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes. If so, it seems that our genes are unable to express this ability, perhaps as part of a tradeoff in acquiring other traits. It is a fertile area of research.
Dedifferentiation is likely to become an important process in the field of regenerative medicine. One extreme example: a lab has been able to turn back the clock and reprogram adult male skin cells into female eggs, a potential milestone in reproductive health. It will be more difficult to control just how far back one wishes to go in the cell's dedifferentiation – part way or all the way back into a stem cell – and then direct it down a different developmental pathway. Yun is optimistic we can learn these tricks from newts.
Senolytics
A growing field of research is using drugs called senolytics to remove senescent cells and slow or even reverse disease of aging.
“Senolytics are great, but senolytics target different types of senescence,” Yun says. “If senescent cells have positive effects in the context of regeneration, of wound healing, then maybe at the beginning of the regeneration process, you may not want to take them out for a little while.”
“If you look at pretty much all biological systems, too little or too much of something can be bad, you have to be in that central zone” and at the proper time, says Atala. “That's true for proteins, sugars, and the drugs that you take. I think the same thing is true for these cells. Why would they be different?”
Our growing understanding that senescence is not a single thing but a variety of things likely means that effective senolytic drugs will not resemble a single sledge hammer but more a carefully manipulated scalpel where some types of senescent cells are removed while others are added. Combinations and timing could be crucial, meaning the difference between regenerating healthy tissue, a scar, or worse.

