Scientists redesign bacteria to tackle the antibiotic resistance crisis
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
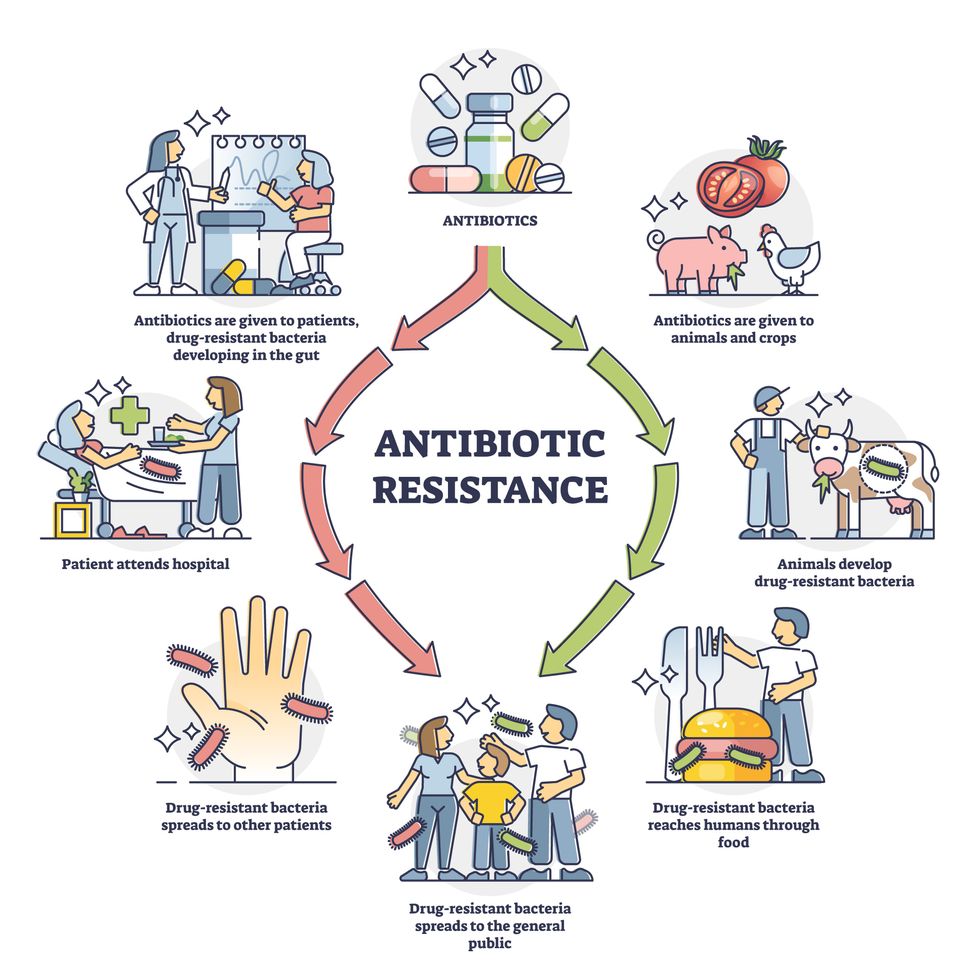
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
As Our AI Systems Get Better, So Must We
In order to build the future we want, we must also become ever better humans, explains the futurist Jamie Metzl in this opinion essay.
As the power and capability of our AI systems increase by the day, the essential question we now face is what constitutes peak human. If we stay where we are while the AI systems we are unleashing continually get better, they will meet and then exceed our capabilities in an ever-growing number of domains. But while some technology visionaries like Elon Musk call for us to slow down the development of AI systems to buy time, this approach alone will simply not work in our hyper-competitive world, particularly when the potential benefits of AI are so great and our frameworks for global governance are so weak. In order to build the future we want, we must also become ever better humans.
The list of activities we once saw as uniquely human where AIs have now surpassed us is long and growing. First, AI systems could beat our best chess players, then our best Go players, then our best champions of multi-player poker. They can see patterns far better than we can, generate medical and other hypotheses most human specialists miss, predict and map out new cellular structures, and even generate beautiful, and, yes, creative, art.
A recent paper by Microsoft researchers analyzing the significant leap in capabilities in OpenAI’s latest AI bot, ChatGPT-4, asserted that the algorithm can “solve novel and difficult tasks that span mathematics, coding, vision, medicine, law, psychology and more, without needing any special prompting.” Calling this functionality “strikingly close to human-level performance,” the authors conclude it “could reasonably be viewed as an early (yet still incomplete) version of an artificial general intelligence (AGI) system.”
The concept of AGI has been around for decades. In its common use, the term suggests a time when individual machines can do many different things at a human level, not just one thing like playing Go or analyzing radiological images. Debating when AGI might arrive, a favorite pastime of computer scientists for years, now has become outdated.
We already have AI algorithms and chatbots that can do lots of different things. Based on the generalist definition, in other words, AGI is essentially already here.
Unfettered by the evolved capacity and storage constraints of our brains, AI algorithms can access nearly all of the digitized cultural inheritance of humanity since the dawn of recorded history and have increasing access to growing pools of digitized biological data from across the spectrum of life.
Once we recognize that both AI systems and humans have unique superpowers, the essential question becomes what each of us can do better than the other and what humans and AIs can best do in active collaboration. The future of our species will depend upon our ability to safely, dynamically, and continually figure that out.
With these ever-larger datasets, rapidly increasing computing and memory power, and new and better algorithms, our AI systems will keep getting better faster than most of us can today imagine. These capabilities have the potential to help us radically improve our healthcare, agriculture, and manufacturing, make our economies more productive and our development more sustainable, and do many important things better.
Soon, they will learn how to write their own code. Like human children, in other words, AI systems will grow up. But even that doesn’t mean our human goose is cooked.
Just like dolphins and dogs, these alternate forms of intelligence will be uniquely theirs, not a lesser or greater version of ours. There are lots of things AI systems can't do and will never be able to do because our AI algorithms, for better and for worse, will never be human. Our embodied human intelligence is its own thing.
Our human intelligence is uniquely ours based on the capacities we have developed in our 3.8-billion-year journey from single cell organisms to us. Our brains and bodies represent continuous adaptations on earlier models, which is why our skeletal systems look like those of lizards and our brains like most other mammals with some extra cerebral cortex mixed in. Human intelligence isn’t just some type of disembodied function but the inextricable manifestation of our evolved physical reality. It includes our sensory analytical skills and all of our animal instincts, intuitions, drives, and perceptions. Disembodied machine intelligence is something different than what we have evolved and possess.
Because of this, some linguists including Noam Chomsky have recently argued that AI systems will never be intelligent as long as they are just manipulating symbols and mathematical tokens without any inherent understanding. Nothing could be further from the truth. Anyone interacting with even first-generation AI chatbots quickly realizes that while these systems are far from perfect or omniscient and can sometimes be stupendously oblivious, they are surprisingly smart and versatile and will get more so… forever. We have little idea even how our own minds work, so judging AI systems based on their output is relatively close to how we evaluate ourselves.
Anyone not awed by the potential of these AI systems is missing the point. AI’s newfound capacities demand that we work urgently to establish norms, standards, and regulations at all levels from local to global to manage the very real risks. Pausing our development of AI systems now doesn’t make sense, however, even if it were possible, because we have no sufficient ways of uniformly enacting such a pause, no plan for how we would use the time, and no common framework for addressing global collective challenges like this.
But if all we feel is a passive awe for these new capabilities, we will also be missing the point.
Human evolution, biology, and cultural history are not just some kind of accidental legacy, disability, or parlor trick, but our inherent superpower. Our ancestors outcompeted rivals for billions of years to make us so well suited to the world we inhabit and helped build. Our social organization at scale has made it possible for us to forge civilizations of immense complexity, engineer biology and novel intelligence, and extend our reach to the stars. Our messy, embodied, intuitive, social human intelligence is roughly mimicable by AI systems but, by definition, never fully replicable by them.
Once we recognize that both AI systems and humans have unique superpowers, the essential question becomes what each of us can do better than the other and what humans and AIs can best do in active collaboration. We still don't know. The future of our species will depend upon our ability to safely, dynamically, and continually figure that out.
As we do, we'll learn that many of our ideas and actions are made up of parts, some of which will prove essentially human and some of which can be better achieved by AI systems. Those in every walk of work and life who most successfully identify the optimal contributions of humans, AIs, and the two together, and who build systems and workflows empowering humans to do human things, machines to do machine things, and humans and machines to work together in ways maximizing the respective strengths of each, will be the champions of the 21st century across all fields.
The dawn of the age of machine intelligence is upon us. It’s a quantum leap equivalent to the domestication of plants and animals, industrialization, electrification, and computing. Each of these revolutions forced us to rethink what it means to be human, how we live, and how we organize ourselves. The AI revolution will happen more suddenly than these earlier transformations but will follow the same general trajectory. Now is the time to aggressively prepare for what is fast heading our way, including by active public engagement, governance, and regulation.
AI systems will not replace us, but, like these earlier technology-driven revolutions, they will force us to become different humans as we co-evolve with our technology. We will never reach peak human in our ongoing evolutionary journey, but we’ve got to manage this transition wisely to build the type of future we’d like to inhabit.
Alongside our ascending AIs, we humans still have a lot of climbing to do.
Based on recent research, new therapies could promote a mix of viruses in the intestines to help prevent diseases of aging.
Story by Big Think
Our gut microbiome plays a substantial role in our health and well-being. Most research, however, focuses on bacteria, rather than the viruses that hide within them. Now, research from the University of Copenhagen, newly published in Nature Microbiology, found that people who live past age 100 have a greater diversity of bacteria-infecting viruses in their intestines than younger people. Furthermore, they found that the viruses are linked to changes in bacterial metabolism that may support mucosal integrity and resistance to pathogens.
The microbiota and aging
In the early 1970s, scientists discovered that the composition of our gut microbiota changes as we age. Recent studies have found that the changes are remarkably predictable and follow a pattern: The microbiota undergoes rapid, dramatic changes as toddlers transition to solid foods; further changes become less dramatic during childhood as the microbiota strikes a balance between the host and the environment; and as that balance is achieved, the microbiota remains mostly stable during our adult years (ages 18-60). However, that stability is lost as we enter our elderly years, and the microbiome undergoes dramatic reorganization. This discovery led scientists to question what causes this change and what effect it has on health.
Centenarians have a distinct gut community enriched in microorganisms that synthesize potent antimicrobial molecules that can kill multidrug-resistant pathogens.
“We are always eager to find out why some people live extremely long lives. Previous research has shown that the intestinal bacteria of old Japanese citizens produce brand-new molecules that make them resistant to pathogenic — that is, disease-promoting — microorganisms. And if their intestines are better protected against infection, well, then that is probably one of the things that cause them to live longer than others,” said Joachim Johansen, a researcher at the University of Copenhagen.
In 2021, a team of Japanese scientists set out to characterize the effect of this change on older people’s health. They specifically wanted to determine if people who lived to be over 100 years old — that is, centenarians — underwent changes that provided them with unique benefits. They discovered centenarians have a distinct gut community enriched in microorganisms that synthesize potent antimicrobial molecules that can kill multidrug-resistant pathogens, including Clostridioides difficile and Enterococcus faecium. In other words, the late-life shift in microbiota reduces an older person’s susceptibility to common gut pathogens.
Viruses can change alter the genes of bacteria
Although the late-in-life microbiota change could be beneficial to health, it remained unclear what facilitated this shift. To solve this mystery, Johansen and his colleagues turned their attention to an often overlooked member of the microbiome: viruses. “Our intestines contain billions of viruses living inside bacteria, and they could not care less about human cells; instead, they infect the bacterial cells. And seeing as there are hundreds of different types of bacteria in our intestines, there are also lots of bacterial viruses,” said Simon Rasmussen, Johansen’s research advisor.
Centenarians had a more diverse virome, including previously undescribed viral genera.
For decades, scientists have explored the possibility of phage therapy — that is, using viruses that infect bacteria (called bacteriophages or simply phages) to kill pathogens. However, bacteriophages can also enhance the bacteria they infect. For example, they can provide genes that help their bacterial host attack other bacteria or provide new metabolic capabilities. Both of these can change which bacteria colonize the gut and, in turn, protect against certain disease states.
Intestinal viruses give bacteria new abilities
Johansen and his colleagues were interested in what types of viruses centenarians had in their gut and whether those viruses carried genes that altered metabolism. They compared fecal samples of healthy centenarians (100+ year-olds) with samples from younger patients (18-100 year-olds). They found that the centenarians had a more diverse virome, including previously undescribed viral genera.
They also revealed an enrichment of genes supporting key steps in the sulfate metabolic pathway. The authors speculate that this translates to increased levels of microbially derived sulfide, which may lead to health-promoting outcomes, such as supporting mucosal integrity and resistance to potential pathogens.
“We have learned that if a virus pays a bacterium a visit, it may actually strengthen the bacterium. The viruses we found in the healthy Japanese centenarians contained extra genes that could boost the bacteria,” said Johansen.
Simon Rasmussen added, “If you discover bacteria and viruses that have a positive effect on the human intestinal flora, the obvious next step is to find out whether only some or all of us have them. If we are able to get these bacteria and their viruses to move in with the people who do not have them, more people could benefit from them.”
This article originally appeared on Big Think, home of the brightest minds and biggest ideas of all time.
Sign up for Big Think’s newsletter


