Scientists redesign bacteria to tackle the antibiotic resistance crisis
Probiotic bacteria can be engineered to fight antibiotic-resistant superbugs by releasing chemicals that kill them.
In 1945, almost two decades after Alexander Fleming discovered penicillin, he warned that as antibiotics use grows, they may lose their efficiency. He was prescient—the first case of penicillin resistance was reported two years later. Back then, not many people paid attention to Fleming’s warning. After all, the “golden era” of the antibiotics age had just began. By the 1950s, three new antibiotics derived from soil bacteria — streptomycin, chloramphenicol, and tetracycline — could cure infectious diseases like tuberculosis, cholera, meningitis and typhoid fever, among others.
Today, these antibiotics and many of their successors developed through the 1980s are gradually losing their effectiveness. The extensive overuse and misuse of antibiotics led to the rise of drug resistance. The livestock sector buys around 80 percent of all antibiotics sold in the U.S. every year. Farmers feed cows and chickens low doses of antibiotics to prevent infections and fatten up the animals, which eventually causes resistant bacterial strains to evolve. If manure from cattle is used on fields, the soil and vegetables can get contaminated with antibiotic-resistant bacteria. Another major factor is doctors overprescribing antibiotics to humans, particularly in low-income countries. Between 2000 to 2018, the global rates of human antibiotic consumption shot up by 46 percent.
In recent years, researchers have been exploring a promising avenue: the use of synthetic biology to engineer new bacteria that may work better than antibiotics. The need continues to grow, as a Lancet study linked antibiotic resistance to over 1.27 million deaths worldwide in 2019, surpassing HIV/AIDS and malaria. The western sub-Saharan Africa region had the highest death rate (27.3 people per 100,000).
Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
To make it worse, our remedy pipelines are drying up. Out of the 18 biggest pharmaceutical companies, 15 abandoned antibiotic development by 2013. According to the AMR Action Fund, venture capital has remained indifferent towards biotech start-ups developing new antibiotics. In 2019, at least two antibiotic start-ups filed for bankruptcy. As of December 2020, there were 43 new antibiotics in clinical development. But because they are based on previously known molecules, scientists say they are inadequate for treating multidrug-resistant bacteria. Researchers warn that if nothing changes, by 2050, antibiotic resistance could kill 10 million people annually.
The rise of synthetic biology
To circumvent this dire future, scientists have been working on alternative solutions using synthetic biology tools, meaning genetically modifying good bacteria to fight the bad ones.
From the time life evolved on earth around 3.8 billion years ago, bacteria have engaged in biological warfare. They constantly strategize new methods to combat each other by synthesizing toxic proteins that kill competition.
For example, Escherichia coli produces bacteriocins or toxins to kill other strains of E.coli that attempt to colonize the same habitat. Microbes like E.coli (which are not all pathogenic) are also naturally present in the human microbiome. The human microbiome harbors up to 100 trillion symbiotic microbial cells. The majority of them are beneficial organisms residing in the gut at different compositions.
The chemicals that these “good bacteria” produce do not pose any health risks to us, but can be toxic to other bacteria, particularly to human pathogens. For the last three decades, scientists have been manipulating bacteria’s biological warfare tactics to our collective advantage.
In the late 1990s, researchers drew inspiration from electrical and computing engineering principles that involve constructing digital circuits to control devices. In certain ways, every cell in living organisms works like a tiny computer. The cell receives messages in the form of biochemical molecules that cling on to its surface. Those messages get processed within the cells through a series of complex molecular interactions.
Synthetic biologists can harness these living cells’ information processing skills and use them to construct genetic circuits that perform specific instructions—for example, secrete a toxin that kills pathogenic bacteria. “Any synthetic genetic circuit is merely a piece of information that hangs around in the bacteria’s cytoplasm,” explains José Rubén Morones-Ramírez, a professor at the Autonomous University of Nuevo León, Mexico. Then the ribosome, which synthesizes proteins in the cell, processes that new information, making the compounds scientists want bacteria to make. “The genetic circuit remains separated from the living cell’s DNA,” Morones-Ramírez explains. When the engineered bacteria replicates, the genetic circuit doesn’t become part of its genome.
Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut.
In 2000, Boston-based researchers constructed an E.coli with a genetic switch that toggled between turning genes on and off two. Later, they built some safety checks into their bacteria. “To prevent unintentional or deleterious consequences, in 2009, we built a safety switch in the engineered bacteria’s genetic circuit that gets triggered after it gets exposed to a pathogen," says James Collins, a professor of biological engineering at MIT and faculty member at Harvard University’s Wyss Institute. “After getting rid of the pathogen, the engineered bacteria is designed to switch off and leave the patient's body.”
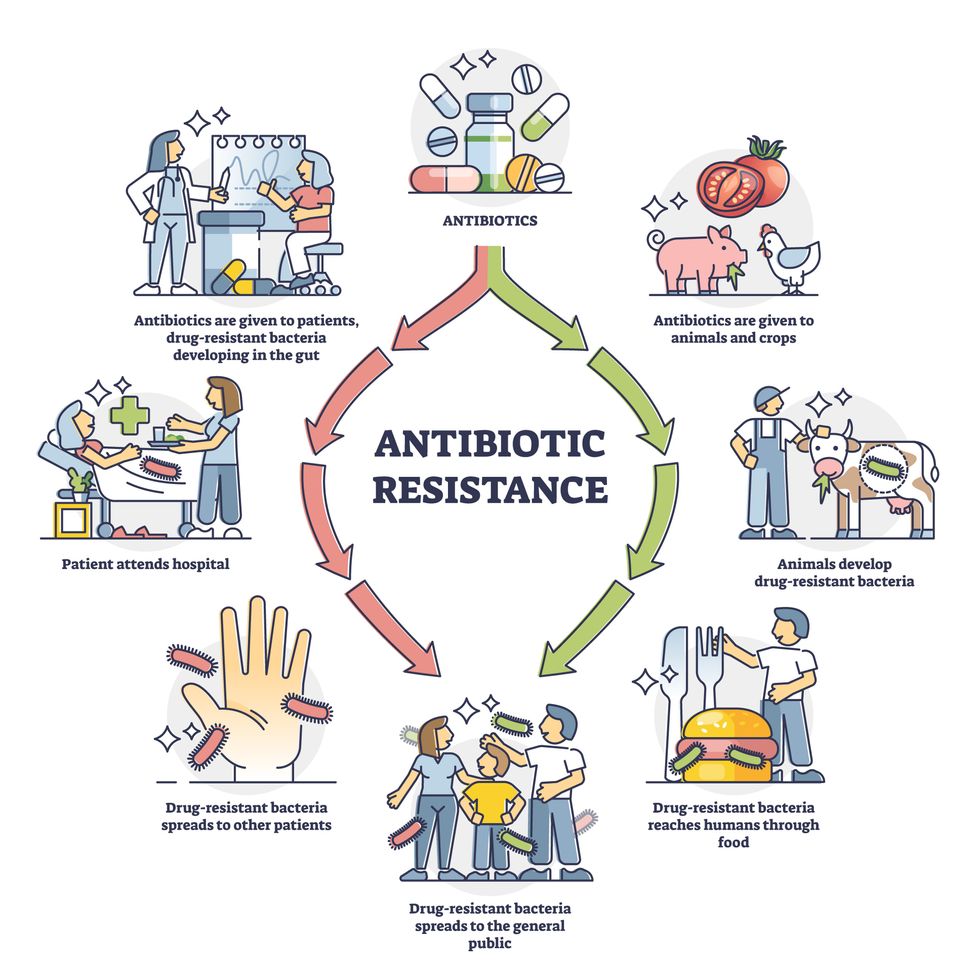
Overuse and misuse of antibiotics causes resistant strains to evolve
Adobe Stock
Seek and destroy
As the field of synthetic biology developed, scientists began using engineered bacteria to tackle superbugs. They first focused on Vibrio cholerae, which in the 19th and 20th century caused cholera pandemics in India, China, the Middle East, Europe, and Americas. Like many other bacteria, V. cholerae communicate with each other via quorum sensing, a process in which the microorganisms release different signaling molecules, to convey messages to its brethren. Highly intelligent by bacterial standards, some multidrug resistant V. cholerae strains can also “collaborate” with other intestinal bacterial species to gain advantage and take hold of the gut. When untreated, cholera has a mortality rate of 25 to 50 percent and outbreaks frequently occur in developing countries, especially during floods and droughts.
Sometimes, however, V. cholerae makes mistakes. In 2008, researchers at Cornell University observed that when quorum sensing V. cholerae accidentally released high concentrations of a signaling molecule called CAI-1, it had a counterproductive effect—the pathogen couldn’t colonize the gut.
So the group, led by John March, professor of biological and environmental engineering, developed a novel strategy to combat V. cholerae. They genetically engineered E.coli to eavesdrop on V. cholerae communication networks and equipped it with the ability to release the CAI-1 molecules. That interfered with V. cholerae progress. Two years later, the Cornell team showed that V. cholerae-infected mice treated with engineered E.coli had a 92 percent survival rate.
These findings inspired researchers to sic the good bacteria present in foods like yogurt and kimchi onto the drug-resistant ones.
Three years later in 2011, Singapore-based scientists engineered E.coli to detect and destroy Pseudomonas aeruginosa, an often drug-resistant pathogen that causes pneumonia, urinary tract infections, and sepsis. Once the genetically engineered E.coli found its target through its quorum sensing molecules, it then released a peptide, that could eradicate 99 percent of P. aeruginosa cells in a test-tube experiment. The team outlined their work in a Molecular Systems Biology study.
“At the time, we knew that we were entering new, uncharted territory,” says lead author Matthew Chang, an associate professor and synthetic biologist at the National University of Singapore and lead author of the study. “To date, we are still in the process of trying to understand how long these microbes stay in our bodies and how they might continue to evolve.”
More teams followed the same path. In a 2013 study, MIT researchers also genetically engineered E.coli to detect P. aeruginosa via the pathogen’s quorum-sensing molecules. It then destroyed the pathogen by secreting a lab-made toxin.
Probiotics that fight
A year later in 2014, a Nature study found that the abundance of Ruminococcus obeum, a probiotic bacteria naturally occurring in the human microbiome, interrupts and reduces V.cholerae’s colonization— by detecting the pathogen’s quorum sensing molecules. The natural accumulation of R. obeum in Bangladeshi adults helped them recover from cholera despite living in an area with frequent outbreaks.
The findings from 2008 to 2014 inspired Collins and his team to delve into how good bacteria present in foods like yogurt and kimchi can attack drug-resistant bacteria. In 2018, Collins and his team developed the engineered probiotic strategy. They tweaked a bacteria commonly found in yogurt called Lactococcus lactis to treat cholera.
Engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut. --José Rubén Morones-Ramírez.
More scientists followed with more experiments. So far, researchers have engineered various probiotic organisms to fight pathogenic bacteria like Staphylococcus aureus (leading cause of skin, tissue, bone, joint and blood infections) and Clostridium perfringens (which causes watery diarrhea) in test-tube and animal experiments. In 2020, Russian scientists engineered a probiotic called Pichia pastoris to produce an enzyme called lysostaphin that eradicated S. aureus in vitro. Another 2020 study from China used an engineered probiotic bacteria Lactobacilli casei as a vaccine to prevent C. perfringens infection in rabbits.
In a study last year, Ramírez’s group at the Autonomous University of Nuevo León, engineered E. coli to detect quorum-sensing molecules from Methicillin-resistant Staphylococcus aureus or MRSA, a notorious superbug. The E. coli then releases a bacteriocin that kills MRSA. “An antibiotic is just a molecule that is not intelligent,” says Ramírez. “On the other hand, engineered bacteria can be trained to target pathogens when they are at their most vulnerable metabolic stage in the human gut.”
Collins and Timothy Lu, an associate professor of biological engineering at MIT, found that engineered E. coli can help treat other conditions—such as phenylketonuria, a rare metabolic disorder, that causes the build-up of an amino acid phenylalanine. Their start-up Synlogic aims to commercialize the technology, and has completed a phase 2 clinical trial.
Circumventing the challenges
The bacteria-engineering technique is not without pitfalls. One major challenge is that beneficial gut bacteria produce their own quorum-sensing molecules that can be similar to those that pathogens secrete. If an engineered bacteria’s biosensor is not specific enough, it will be ineffective.
Another concern is whether engineered bacteria might mutate after entering the gut. “As with any technology, there are risks where bad actors could have the capability to engineer a microbe to act quite nastily,” says Collins of MIT. But Collins and Ramírez both insist that the chances of the engineered bacteria mutating on its own are virtually non-existent. “It is extremely unlikely for the engineered bacteria to mutate,” Ramírez says. “Coaxing a living cell to do anything on command is immensely challenging. Usually, the greater risk is that the engineered bacteria entirely lose its functionality.”
However, the biggest challenge is bringing the curative bacteria to consumers. Pharmaceutical companies aren’t interested in antibiotics or their alternatives because it’s less profitable than developing new medicines for non-infectious diseases. Unlike the more chronic conditions like diabetes or cancer that require long-term medications, infectious diseases are usually treated much quicker. Running clinical trials are expensive and antibiotic-alternatives aren’t lucrative enough.
“Unfortunately, new medications for antibiotic resistant infections have been pushed to the bottom of the field,” says Lu of MIT. “It's not because the technology does not work. This is more of a market issue. Because clinical trials cost hundreds of millions of dollars, the only solution is that governments will need to fund them.” Lu stresses that societies must lobby to change how the modern healthcare industry works. “The whole world needs better treatments for antibiotic resistance.”
How the body's immune resilience affects our health and lifespan
Immune cells battle an infection.
Story by Big Think
It is a mystery why humans manifest vast differences in lifespan, health, and susceptibility to infectious diseases. However, a team of international scientists has revealed that the capacity to resist or recover from infections and inflammation (a trait they call “immune resilience”) is one of the major contributors to these differences.
Immune resilience involves controlling inflammation and preserving or rapidly restoring immune activity at any age, explained Weijing He, a study co-author. He and his colleagues discovered that people with the highest level of immune resilience were more likely to live longer, resist infection and recurrence of skin cancer, and survive COVID and sepsis.
Measuring immune resilience
The researchers measured immune resilience in two ways. The first is based on the relative quantities of two types of immune cells, CD4+ T cells and CD8+ T cells. CD4+ T cells coordinate the immune system’s response to pathogens and are often used to measure immune health (with higher levels typically suggesting a stronger immune system). However, in 2021, the researchers found that a low level of CD8+ T cells (which are responsible for killing damaged or infected cells) is also an important indicator of immune health. In fact, patients with high levels of CD4+ T cells and low levels of CD8+ T cells during SARS-CoV-2 and HIV infection were the least likely to develop severe COVID and AIDS.
Individuals with optimal levels of immune resilience were more likely to live longer.
In the same 2021 study, the researchers identified a second measure of immune resilience that involves two gene expression signatures correlated with an infected person’s risk of death. One of the signatures was linked to a higher risk of death; it includes genes related to inflammation — an essential process for jumpstarting the immune system but one that can cause considerable damage if left unbridled. The other signature was linked to a greater chance of survival; it includes genes related to keeping inflammation in check. These genes help the immune system mount a balanced immune response during infection and taper down the response after the threat is gone. The researchers found that participants who expressed the optimal combination of genes lived longer.
Immune resilience and longevity
The researchers assessed levels of immune resilience in nearly 50,000 participants of different ages and with various types of challenges to their immune systems, including acute infections, chronic diseases, and cancers. Their evaluation demonstrated that individuals with optimal levels of immune resilience were more likely to live longer, resist HIV and influenza infections, resist recurrence of skin cancer after kidney transplant, survive COVID infection, and survive sepsis.
However, a person’s immune resilience fluctuates all the time. Study participants who had optimal immune resilience before common symptomatic viral infections like a cold or the flu experienced a shift in their gene expression to poor immune resilience within 48 hours of symptom onset. As these people recovered from their infection, many gradually returned to the more favorable gene expression levels they had before. However, nearly 30% who once had optimal immune resilience did not fully regain that survival-associated profile by the end of the cold and flu season, even though they had recovered from their illness.
Intriguingly, some people who are 90+ years old still have optimal immune resilience, suggesting that these individuals’ immune systems have an exceptional capacity to control inflammation and rapidly restore proper immune balance.
This could suggest that the recovery phase varies among people and diseases. For example, young female sex workers who had many clients and did not use condoms — and thus were repeatedly exposed to sexually transmitted pathogens — had very low immune resilience. However, most of the sex workers who began reducing their exposure to sexually transmitted pathogens by using condoms and decreasing their number of sex partners experienced an improvement in immune resilience over the next 10 years.
Immune resilience and aging
The researchers found that the proportion of people with optimal immune resilience tended to be highest among the young and lowest among the elderly. The researchers suggest that, as people age, they are exposed to increasingly more health conditions (acute infections, chronic diseases, cancers, etc.) which challenge their immune systems to undergo a “respond-and-recover” cycle. During the response phase, CD8+ T cells and inflammatory gene expression increase, and during the recovery phase, they go back down.
However, over a lifetime of repeated challenges, the immune system is slower to recover, altering a person’s immune resilience. Intriguingly, some people who are 90+ years old still have optimal immune resilience, suggesting that these individuals’ immune systems have an exceptional capacity to control inflammation and rapidly restore proper immune balance despite the many respond-and-recover cycles that their immune systems have faced.
Public health ramifications could be significant. Immune cell and gene expression profile assessments are relatively simple to conduct, and being able to determine a person’s immune resilience can help identify whether someone is at greater risk for developing diseases, how they will respond to treatment, and whether, as well as to what extent, they will recover.
A new injection is helping stave off RSV this season
The FDA approved a single-dose, long-acting injection to protect babies and toddlers from RSV over the fall and winter.
In November 2021, Mickayla Wininger’s then one-month-old son, Malcolm, endured a terrifying bout with RSV, the respiratory syncytial (sin-SISH-uhl) virus—a common ailment that affects all age groups. Most people recover from mild, cold-like symptoms in a week or two, but RSV can be life-threatening in others, particularly infants.
Wininger, who lives in southern Illinois, was dressing Malcolm for bed when she noticed what seemed to be a minor irregularity with this breathing. She and her fiancé, Gavin McCullough, planned to take him to the hospital the next day. The matter became urgent when, in the morning, the boy’s breathing appeared to have stopped.
After they dialed 911, Malcolm started breathing again, but he ended up being hospitalized three times for RSV and defects in his heart. Eventually, he recovered fully from RSV, but “it was our worst nightmare coming to life,” Wininger recalled.
It’s a scenario that the federal government is taking steps to prevent. In July, the Food and Drug Administration approved a single-dose, long-acting injection to protect babies and toddlers. The injection, called Beyfortus, or nirsevimab, became available this October. It reduces the incidence of RSV in pre-term babies and other infants for their first RSV season. Children at highest risk for severe RSV are those who were born prematurely and have either chronic lung disease of prematurity or congenital heart disease. In those cases, RSV can progress to lower respiratory tract diseases such as pneumonia and bronchiolitis, or swelling of the lung’s small airway passages.
Each year, RSV is responsible for 2.1 million outpatient visits among children younger than five-years-old, 58,000 to 80,000 hospitalizations in this age group, and between 100 and 300 deaths, according to the Centers for Disease Control and Prevention. Transmitted through close contact with an infected person, the virus circulates on a seasonal basis in most regions of the country, typically emerging in the fall and peaking in the winter.
In August, however, the CDC issued a health advisory on a late-summer surge in severe cases of RSV among young children in Florida and Georgia. The agency predicts "increased RSV activity spreading north and west over the following two to three months.”
Infants are generally more susceptible to RSV than older people because their airways are very small, and their mechanisms to clear these passages are underdeveloped. RSV also causes mucus production and inflammation, which is more of a problem when the airway is smaller, said Jennifer Duchon, an associate professor of newborn medicine and pediatrics in the Icahn School of Medicine at Mount Sinai in New York.
In 2021 and 2022, RSV cases spiked, sending many to emergency departments. “RSV can cause serious disease in infants and some children and results in a large number of emergency department and physician office visits each year,” John Farley, director of the Office of Infectious Diseases in the FDA’s Center for Drug Evaluation and Research, said in a news release announcing the approval of the RSV drug. The decision “addresses the great need for products to help reduce the impact of RSV disease on children, families and the health care system.”
Sean O’Leary, chair of the committee on infectious diseases for the American Academy of Pediatrics, says that “we’ve never had a product like this for routine use in children, so this is very exciting news.” It is recommended for all kids under eight months old for their first RSV season. “I would encourage nirsevimab for all eligible children when it becomes available,” O’Leary said.
For those children at elevated risk of severe RSV and between the ages of 8 and 19 months, the CDC recommends one dose in their second RSV season.
The drug will be “really helpful to keep babies healthy and out of the hospital,” said O’Leary, a professor of pediatrics at the University of Colorado Anschutz Medical Campus/Children’s Hospital Colorado in Denver.
An antiviral drug called Synagis (palivizumab) has been an option to prevent serious RSV illness in high-risk infants since it was approved by the FDA in 1998. The injection must be given monthly during RSV season. However, its use is limited to “certain children considered at high risk for complications, does not help cure or treat children already suffering from serious RSV disease, and cannot prevent RSV infection,” according to the National Foundation for Infectious Diseases.
Until the approval this summer of the new monoclonal antibody, nirsevimab, there wasn’t a reliable method to prevent infection in most healthy infants.
Both nirsevimab and palivizumab are monoclonal antibodies that act against RSV. Monoclonal antibodies are lab-made proteins that mimic the immune system’s ability to fight off harmful pathogens such as viruses. A single intramuscular injection of nirsevimab preceding or during RSV season may provide protection.
The strategy with the new monoclonal antibody is “to extend protection to healthy infants who nonetheless are at risk because of their age, as well as infants with additional medical risk factors,” said Philippa Gordon, a pediatrician and infectious disease specialist in Brooklyn, New York, and medical adviser to Park Slope Parents, an online community support group.
No specific preventive measure is needed for older and healthier kids because they will develop active immunity, which is more durable. Meanwhile, older adults, who are also vulnerable to RSV, can receive one of two new vaccines. So can pregnant women, who pass on immunity to the fetus, Gordon said.
Until the approval this summer of the new monoclonal antibody, nirsevimab, there wasn’t a reliable method to prevent infection in most healthy infants, “nor is there any treatment other than giving oxygen or supportive care,” said Stanley Spinner, chief medical officer and vice president of Texas Children’s Pediatrics and Texas Children’s Urgent Care.
As with any virus, washing hands frequently and keeping infants and children away from sick people are the best defenses, Duchon said. This approach isn’t foolproof because viruses can run rampant in daycare centers, schools and parents’ workplaces, she added.
Mickayla Wininger, Malcolm’s mother, insists that family and friends wear masks, wash their hands and use hand sanitizer when they’re around her daughter and two sons. She doesn’t allow them to kiss or touch the children. Some people take it personally, but she would rather be safe than sorry.
Wininger recalls the severe anxiety caused by Malcolm's ordeal with RSV. After returning with her infant from his hospital stays, she was terrified to go to sleep. “My fiancé and I would trade shifts, so that someone was watching over our son 24 hours a day,” she said. “I was doing a night shift, so I would take caffeine pills to try and keep myself awake and would end up crashing early hours in the morning and wake up frantically thinking something happened to my son.”
Two years later, her anxiety has become more manageable, and Malcolm is doing well. “He is thriving now,” Wininger said. He recently had his second birthday and "is just the spunkiest boy you will ever meet. He looked death straight in the eyes and fought to be here today.”