COVID Variants Are Like “a Thief Changing Clothes” – and Our Camera System Barely Exists

Being able to track variants of concern in real time is crucial to our ability to stay ahead of the virus.
Whether it's "natural selection" as Darwin called it, or it's "mutating" as the X-Men called it, living organisms change over time, developing thumbs or more efficient protein spikes, depending on the organism and the demands of its environment. The coronavirus that causes COVID-19, SARS-CoV-2, is not an exception, and now, after the virus has infected millions of people around the globe for more than a year, scientists are beginning to see those changes.
The notorious variants that have popped up include B.1.1.7, sometimes called the UK variant, as well as P.1 and B.1.351, which seem to have emerged in Brazil and South Africa respectively. As vaccinations are picking up pace, officials are warning that now
is not the time to become complacent or relax restrictions because the variants aren't well understood.
Some appear to be more transmissible, and deadlier, while others can evade the immune system's defenses better than earlier versions of the virus, potentially undermining the effectiveness of vaccines to some degree. Genomic surveillance, the process of sequencing the genetic code of the virus widely to observe changes and patterns, is a critical way that scientists can keep track of its evolution and work to understand how the variants might affect humans.
"It's like a thief changing clothes"
It's important to note that viruses mutate all the time. If there were funding and personnel to sequence the genome of every sample of the virus, scientists would see thousands of mutations. Not every variant deserves our attention. The vast majority of mutations are not important at all, but recognizing those that are is a crucial tool in getting and staying ahead of the virus. The work of sequencing, analyzing, observing patterns, and using public health tools as necessary is complicated and confusing to those without years of specialized training.
Jeremy Kamil, associate professor of microbiology and immunology at LSU Health Shreveport, in Louisiana, says that the variants developing are like a thief changing clothes. The thief goes in your house, steals your stuff, then leaves and puts on a different shirt and a wig, in the hopes you won't recognize them. Genomic surveillance catches the "thief" even in those different clothes.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context.
Understanding variants, both the uninteresting ones and the potentially concerning ones, gives public health officials and researchers at different levels a useful set of tools. Locally, knowing which variants are circulating in the community helps leaders know whether mask mandates and similar measures should be implemented or discontinued, or whether businesses and schools can open relatively safely.
There's more to it than observing new variants
Analysis is complex, particularly when it comes to understanding which variants are of concern. "So the question is always if a mutation becomes common, is that a random occurrence?" says Phoebe Lostroh, associate professor of molecular biology at Colorado College. "Or is the variant the result of some kind of selection because the mutation changes some property about the virus that makes it reproduce more quickly than variants of the virus that don't have that mutation? For a virus, [mutations can affect outcomes like] how much it replicates inside a person's body, how much somebody breathes it out, whether the particles that somebody might breathe in get smaller and can lead to greater transmission."
Along with all of those factors, accurate and useful genomic surveillance requires an understanding of where variants are occurring, how they are related, and an examination of why they might be prevalent.
For example, if a potentially worrisome variant appears in a community and begins to spread very quickly, it's not time to raise a public health alarm until several important questions have been answered, such as whether the variant is spreading due to specific events, or if it's happening because the mutation has allowed the virus to infect people more efficiently. Kamil offered a hypothetical scenario to explain: Imagine that a member of a community became infected and the virus mutated. That person went to church and three more people were infected, but one of them went to a karaoke bar and while singing infected 100 other people. Examining the conditions under which the virus has spread is, therefore, an essential part of untangling whether a mutation itself made the virus more transmissible or if an infected person's behaviors contributed to a local outbreak.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context. Genomic sequencing can help with that, but only when it's coordinated. When the same mutation occurs frequently, but is localized to one region, it's a concern, but when the same mutation happens in different places at the same time, it's much more likely that the "virus is learning that's a good mutation," explains Kamil.
The process is called convergent evolution, and it was a fascinating topic long before COVID. Just as your heritage can be traced through DNA, so can that of viruses, and when separate lineages develop similar traits it's almost like scientists can see evolution happening in real time. A mutation to SARS-CoV-2 that happens in more than one place at once is a mutation that makes it easier in some way for the virus to survive and that is when it may become alarming. The widespread, documented variants P.1 and B.1.351 are examples of convergence because they share some of the same virulent mutations despite having developed thousands of miles apart.
However, even variants that are emerging in different places at the same time don't present the kind of threat SARS-CoV-2 did in 2019. "This is nature," says Kamil. "It just means that this virus will not easily be driven to extinction or complete elimination by vaccines." Although a person who has already had COVID-19 can be reinfected with a variant, "it is almost always much milder disease" than the original infection, Kamil adds. Rather than causing full-fledged disease, variants have the potiental to "penetrate herd immunity, spreading relatively quietly among people who have developed natural immunity or been vaccinated, until the virus finds someone who has no immunity yet, and that person would be at risk of hospitalization-grade severe disease or death."
Surveillance and predictions
According to Lostroh, genomic surveillance can help scientists predict what's going to happen. "With the British strain, for instance, that's more transmissible, you can measure how fast it's doubling in the population and you can sort of tell whether we should take more measures against this mutation. Should we shut things down a little longer because that mutation is present in the population? That could be really useful if you did enough sampling in the population that you knew where it was," says Lostroh. If, for example, the more transmissible strain was present in 50 percent of cases, but in another county or state it was barely present, it would allow for rolling lockdowns instead of sweeping measures.
Variants are also extremely important when it comes to the development, manufacture, and distribution of vaccines. "You're also looking at medical countermeasures, such as whether your vaccine is still effective, or if your antiviral needs to be updated," says Lane Warmbrod, a senior analyst and research associate at Johns Hopkins Center for Health Security.
Properly funded and extensive genomic surveillance could eventually help control endemic diseases, too, like the seasonal flu, or other common respiratory infections. Kamil says he envisions a future in which genomic surveillance allows for prediction of sickness just as the weather is predicted today. "It's a 51 for infection today at the San Francisco Airport. There's been detection of some respiratory viruses," he says, offering an example. He says that if you're a vulnerable person, if you're immune-suppressed for some reason, you may want to wear a mask based on the sickness report.
The U.S. has the ability, but lacks standards
The benefits of widespread genomic surveillance are clear, and the United States certainly has the necessary technology, equipment, and personnel to carry it out. But, it's not happening at the speed and extent it needs to for the country to gain the benefits.
"The numbers are improving," said Kamil. "We're probably still at less than half a percent of all the samples that have been taken have been sequenced since the beginning of the pandemic."
Although there's no consensus on how many sequences is ideal for a robust surveillance program, modeling performed by the company Illumina suggests about 5 percent of positive tests should be sequenced. The reasons the U.S. has lagged in implementing a sequencing program are complex and varied, but solvable.
Perhaps the most important element that is currently missing is leadership. In order to conduct an effective genomic surveillance program, there need to be standards. The Johns Hopkins Center for Health Security recently published a paper with recommendations as to what kinds of elements need to be standardized in order to make the best use of sequencing technology and analysis.
"Along with which bioinformatic pipelines you're going to use to do the analyses, which sequencing strategy protocol are you going to use, what's your sampling strategy going to be, how is the data is going to be reported, what data gets reported," says Warmbrod. Currently, there's no guidance from the CDC on any of those things. So, while scientists can collect and report information, they may be collecting and reporting different information that isn't comparable, making it less useful for public health measures and vaccine updates.
Globally, one of the most important tools in making the information from genomic surveillance useful is GISAID, a platform designed for scientists to share -- and, importantly, to be credited for -- their data regarding genetic sequences of influenza. Originally, it was launched as a database of bird flu sequences, but has evolved to become an essential tool used by the WHO to make flu vaccine virus recommendations each year. Scientists who share their credentials have free access to the database, and anyone who uses information from the database must credit the scientist who uploaded that information.
Safety, logistics, and funding matter
Scientists at university labs and other small organizations have been uploading sequences to GISAID almost from the beginning of the pandemic, but their funding is generally limited, and there are no standards regarding information collection or reporting. Private, for-profit labs haven't had motivation to set up sequencing programs, although many of them have the logistical capabilities and funding to do so. Public health departments are understaffed, underfunded, and overwhelmed.
University labs may also be limited by safety concerns. The SARS-CoV-2 virus is dangerous, and there's a question of how samples should be transported to labs for sequencing.
Larger, for-profit organizations often have the tools and distribution capabilities to safely collect and sequence samples, but there hasn't been a profit motive. Genomic sequencing is less expensive now than ever before, but even at $100 per sample, the cost adds up -- not to mention the cost of employing a scientist with the proper credentials to analyze the sequence.
The path forward
The recently passed COVID-19 relief bill does have some funding to address genomic sequencing. Specifically, the American Rescue Plan Act includes $1.75 billion in funding for the Centers for Disease Control and Prevention's Advanced Molecular Detection (AMD) program. In an interview last month, CDC Director Rochelle Walensky said that the additional funding will be "a dial. And we're going to need to dial it up." AMD has already announced a collaboration called the Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance (SPHERES) Initiative that will bring together scientists from public health, academic, clinical, and non-profit laboratories across the country with the goal of accelerating sequencing.
Such a collaboration is a step toward following the recommendations in the paper Warmbrod coauthored. Building capacity now, creating a network of labs, and standardizing procedures will mean improved health in the future. "I want to be optimistic," she says. "The good news is there are a lot of passionate, smart, capable people who are continuing to work with government and work with different stakeholders." She cautions, however, that without a national strategy we won't succeed.
"If we maximize the potential and create that framework now, we can also use it for endemic diseases," she says. "It's a very helpful system for more than COVID if we're smart in how we plan it."
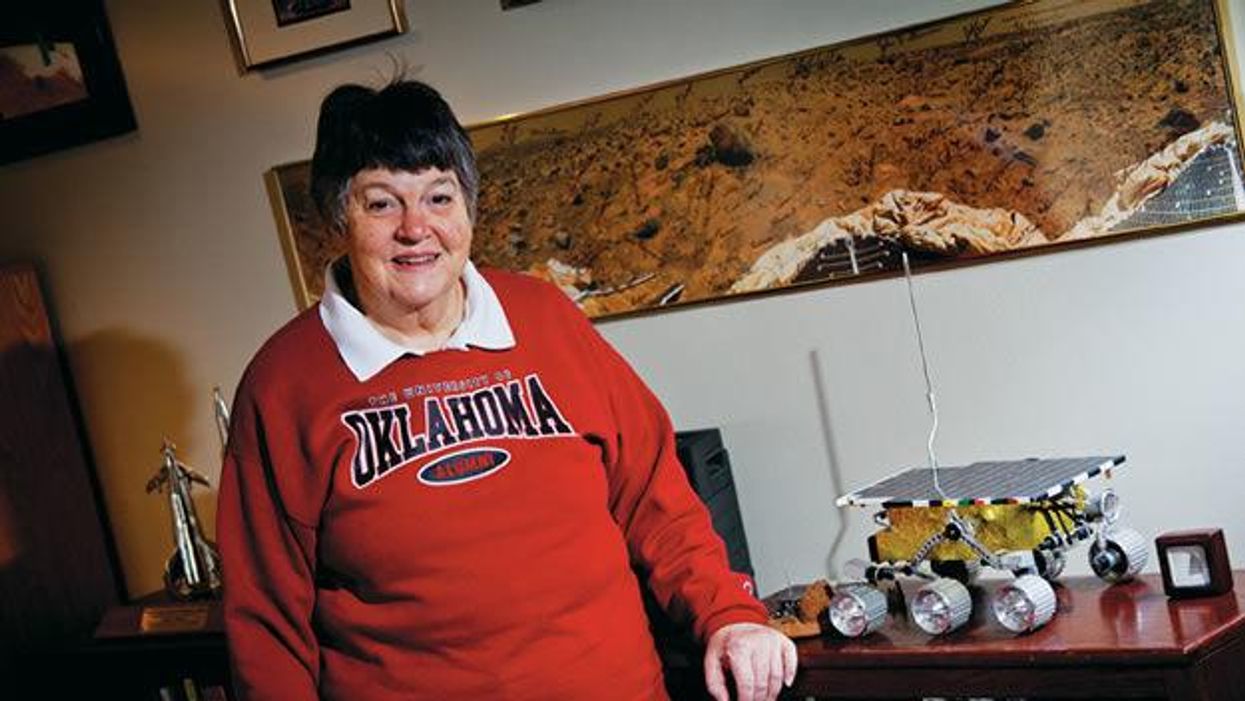
Donna Shirley pictured at her home in Tulsa, with a model of the Sojourner rover she was in charge of that explored Mars.
When NASA's Perseverance rover landed successfully on Mars on February 18, 2021, calling it "one giant leap for mankind" – as Neil Armstrong said when he set foot on the moon in 1969 – would have been inaccurate. This year actually marked the fifth time the U.S. space agency has put a remote-controlled robotic exploration vehicle on the Red Planet. And it was a female engineer named Donna Shirley who broke new ground for women in science as the manager of both the Mars Exploration Program and the 30-person team that built Sojourner, the first rover to land on Mars on July 4, 1997.
For Shirley, the Mars Pathfinder mission was the climax of her 32-year career at NASA's Jet Propulsion Laboratory (JPL) in Pasadena, California. The Oklahoma-born scientist, who earned her Master's degree in aerospace engineering from the University of Southern California, saw her profile skyrocket with media appearances from CNN to the New York Times, and her autobiography Managing Martians came out in 1998. Now 79 and living in a Tulsa retirement community, she still embraces her status as a female pioneer.
"Periodically, I'll hear somebody say they got into the space program because of me, and that makes me feel really good," Shirley told Leaps.org. "I look at the mission control area, and there are a lot of women in there. I'm quite pleased I was able to break the glass ceiling."
Her $25-million, 25-pound microrover – powered by solar energy and designed to get rock samples and test soil chemistry for evidence of life – was named after Sojourner Truth, a 19th-century Black abolitionist and women's rights activist. Unlike Mars Pathfinder, Shirley didn't have to travel more than 131 million miles to reach her goal, but her path to scientific fame as a woman sometimes resembled an asteroid field.
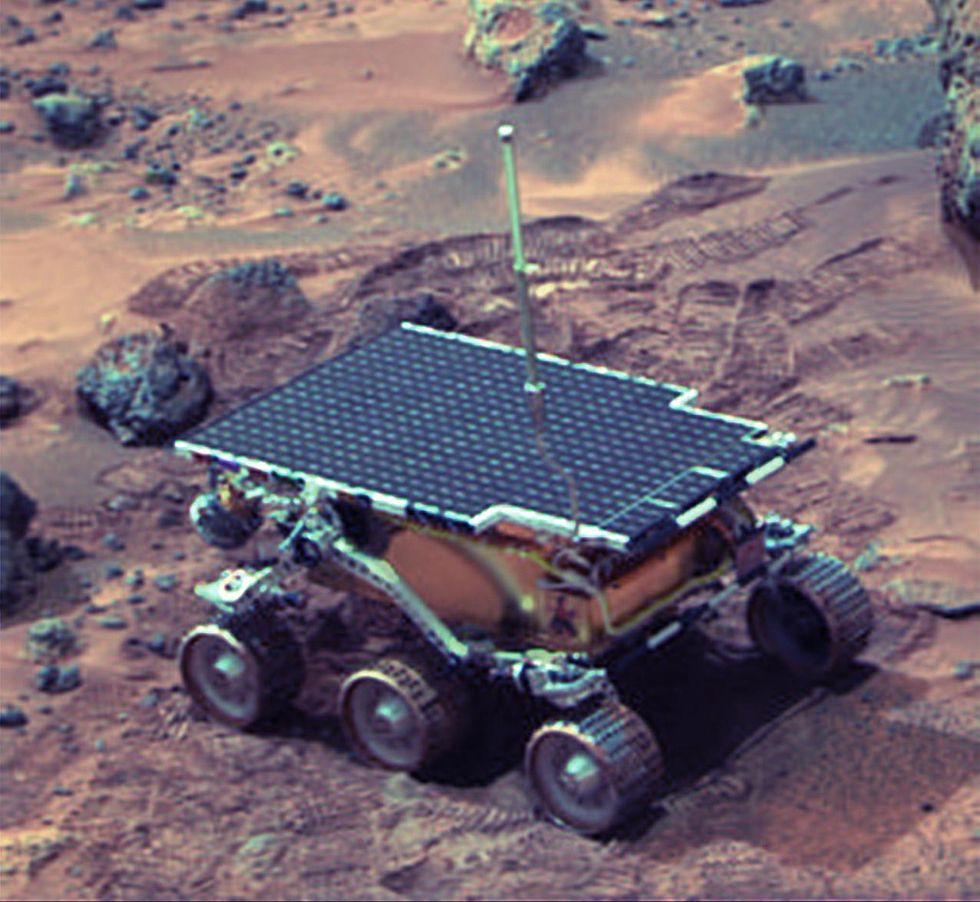 The Sojourner Rover in 1997 on Mars.NASA/JPL
The Sojourner Rover in 1997 on Mars.NASA/JPLAs a high-IQ tomboy growing up in Wynnewood, Oklahoma (pop. 2,300), Shirley yearned to escape. She decided to become an engineer at age 10 and took flying lessons at 15. Her extraterrestrial aspirations were fueled by Ray Bradbury's The Martian Chronicles and Arthur C. Clarke's The Sands of Mars. Yet when she entered the University of Oklahoma (OU) in 1958, her freshman academic advisor initially told her: "Girls can't be engineers." She ignored him.
Years later, Shirley would combat such archaic thinking, succeeding at JPL with her creative, collaborative management style. "If you look at the literature, you'll find that teams that are either led by or heavily involved with women do better than strictly male teams," she noted.
However, her career trajectory stalled at OU. Burned out by her course load and distracted by a broken engagement to marry a fellow student, she switched her major to professional writing. After graduation, she applied her aeronautical background as a McDonnell Aircraft technical writer, but her boss, she says, harassed her and she faced gender-based hostility from male co-workers.
Returning to OU, Shirley finished off her engineering degree and became a JPL aerodynamist in 1966 after answering an ad in the St. Louis Post-Dispatch. At first, she was the only female engineer among the research center's 2,000-odd engineers. She wore many hats, from designing planetary atmospheric entry vehicles to picking the launch date of November 4, 1973 for Mariner 10's mission to Venus and Mercury.
By the mid-1980's, she was managing teams that focused on robotics and Mars, delivering creative solutions when NASA budget cuts loomed. In 1989, the same year the Sojourner microrover concept was born, President George H.W. Bush announced his Space Exploration Initiative, including plans for a human mission to Mars by 2019.
That target, of course, wasn't attained, despite huge advances in technology and our understanding of the Martian environment. Today, Shirley believes humans could land on Mars by 2030. She became the founding director of the Science Fiction Museum and Hall of Fame in Seattle in 2004 after leaving NASA, and to this day, she enjoys checking out pop culture portrayals of Mars landings – even if they're not always accurate.
After the novel The Martian was published in 2011, which later was adapted into the hit film starring Matt Damon, Shirley phoned author Andy Weir: "You've got a major mistake in here. It says there's a storm that tries to blow the rocket over. But actually, the Mars atmosphere is so thin, it would never blow a rocket over!"
Fearlessly speaking her mind and seeking the stars helped Donna Shirley make history. However, a 2019 Washington Post story noted: "Women make up only about a third of NASA's workforce. They comprise just 28 percent of senior executive leadership positions and are only 16 percent of senior scientific employees." Whether it's traveling to Mars or trending toward gender equality, we've still got a long way to go.
Announcing March Event: "COVID Vaccines and the Return to Life: Part 1"
Leading medical and scientific experts will discuss the latest developments around the COVID-19 vaccines at our March 11th event.
EVENT INFORMATION
DATE:
Thursday, March 11th, 2021 at 12:30pm - 1:45pm EST
On the one-year anniversary of the global declaration of the pandemic, this virtual event will convene leading scientific and medical experts to discuss the most pressing questions around the COVID-19 vaccines. Planned topics include the effect of the new circulating variants on the vaccines, what we know so far about transmission dynamics post-vaccination, how individuals can behave post-vaccination, the myths of "good" and "bad" vaccines as more alternatives come on board, and more. A public Q&A will follow the expert discussion.
CONTACT:
kira@goodinc.com
LOCATION:
Zoom webinar
SPEAKERS:

Dr. Paul Offit speaking at Communicating Vaccine Science.
commons.wikimedia.orgDr. Paul Offit, M.D., is the director of the Vaccine Education Center and an attending physician in infectious diseases at the Children's Hospital of Philadelphia. He is a co-inventor of the rotavirus vaccine for infants, and he has lent his expertise to the advisory committees that review data on new vaccines for the CDC and FDA.

Dr. Monica Gandhi
UCSF Health
Dr. Monica Gandhi, M.D., MPH, is Professor of Medicine and Associate Division Chief (Clinical Operations/ Education) of the Division of HIV, Infectious Diseases, and Global Medicine at UCSF/ San Francisco General Hospital.

Dr. Onyema Ogbuagu, MBBCh, FACP, FIDSA
Yale Medicine
Dr. Onyema Ogbuagu, MBBCh, is an infectious disease physician at Yale Medicine who treats COVID-19 patients and leads Yale's clinical studies around COVID-19. He ran Yale's trial of the Pfizer/BioNTech vaccine.

Dr. Eric Topol
Dr. Topol's Twitter
Dr. Eric Topol, M.D., is a cardiologist, scientist, professor of molecular medicine, and the director and founder of Scripps Research Translational Institute. He has led clinical trials in over 40 countries with over 200,000 patients and pioneered the development of many routinely used medications.
REGISTER NOW
This event is the first of a four-part series co-hosted by LeapsMag, the Aspen Institute Science & Society Program, and the Sabin–Aspen Vaccine Science & Policy Group, with generous support from the Gordon and Betty Moore Foundation and the Howard Hughes Medical Institute.

Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.