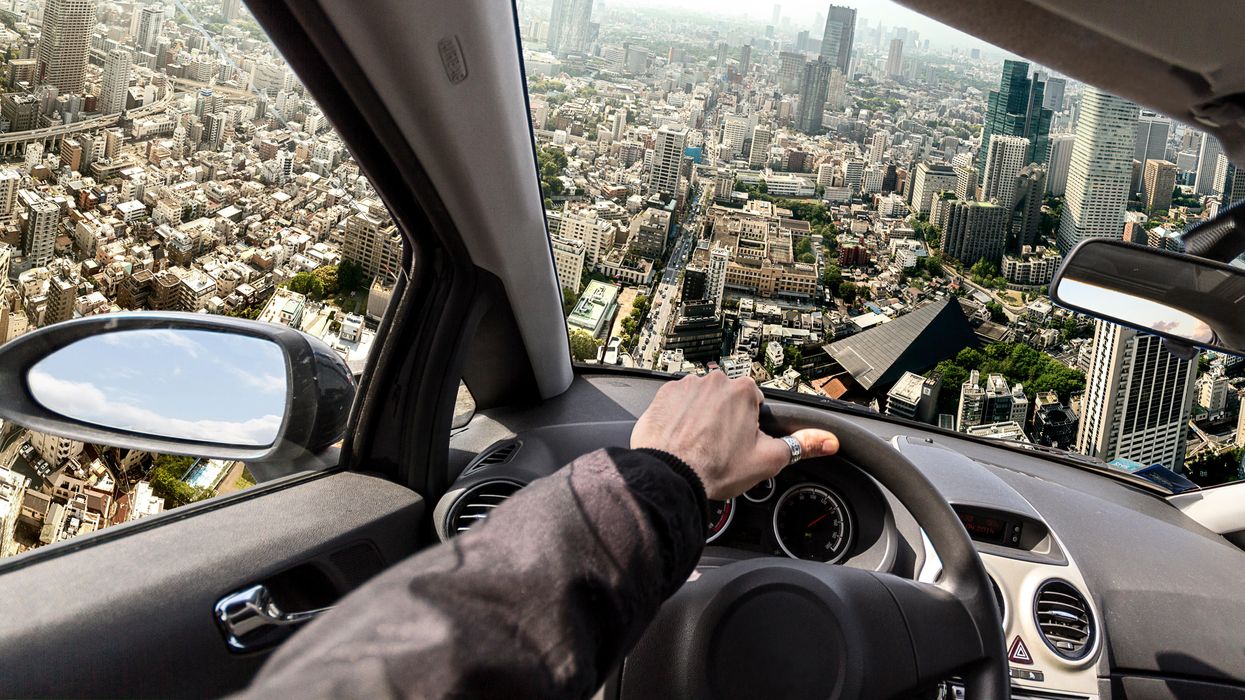
COVID Variants Are Like “a Thief Changing Clothes” – and Our Camera System Barely Exists

Being able to track variants of concern in real time is crucial to our ability to stay ahead of the virus.
Whether it's "natural selection" as Darwin called it, or it's "mutating" as the X-Men called it, living organisms change over time, developing thumbs or more efficient protein spikes, depending on the organism and the demands of its environment. The coronavirus that causes COVID-19, SARS-CoV-2, is not an exception, and now, after the virus has infected millions of people around the globe for more than a year, scientists are beginning to see those changes.
The notorious variants that have popped up include B.1.1.7, sometimes called the UK variant, as well as P.1 and B.1.351, which seem to have emerged in Brazil and South Africa respectively. As vaccinations are picking up pace, officials are warning that now
is not the time to become complacent or relax restrictions because the variants aren't well understood.
Some appear to be more transmissible, and deadlier, while others can evade the immune system's defenses better than earlier versions of the virus, potentially undermining the effectiveness of vaccines to some degree. Genomic surveillance, the process of sequencing the genetic code of the virus widely to observe changes and patterns, is a critical way that scientists can keep track of its evolution and work to understand how the variants might affect humans.
"It's like a thief changing clothes"
It's important to note that viruses mutate all the time. If there were funding and personnel to sequence the genome of every sample of the virus, scientists would see thousands of mutations. Not every variant deserves our attention. The vast majority of mutations are not important at all, but recognizing those that are is a crucial tool in getting and staying ahead of the virus. The work of sequencing, analyzing, observing patterns, and using public health tools as necessary is complicated and confusing to those without years of specialized training.
Jeremy Kamil, associate professor of microbiology and immunology at LSU Health Shreveport, in Louisiana, says that the variants developing are like a thief changing clothes. The thief goes in your house, steals your stuff, then leaves and puts on a different shirt and a wig, in the hopes you won't recognize them. Genomic surveillance catches the "thief" even in those different clothes.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context.
Understanding variants, both the uninteresting ones and the potentially concerning ones, gives public health officials and researchers at different levels a useful set of tools. Locally, knowing which variants are circulating in the community helps leaders know whether mask mandates and similar measures should be implemented or discontinued, or whether businesses and schools can open relatively safely.
There's more to it than observing new variants
Analysis is complex, particularly when it comes to understanding which variants are of concern. "So the question is always if a mutation becomes common, is that a random occurrence?" says Phoebe Lostroh, associate professor of molecular biology at Colorado College. "Or is the variant the result of some kind of selection because the mutation changes some property about the virus that makes it reproduce more quickly than variants of the virus that don't have that mutation? For a virus, [mutations can affect outcomes like] how much it replicates inside a person's body, how much somebody breathes it out, whether the particles that somebody might breathe in get smaller and can lead to greater transmission."
Along with all of those factors, accurate and useful genomic surveillance requires an understanding of where variants are occurring, how they are related, and an examination of why they might be prevalent.
For example, if a potentially worrisome variant appears in a community and begins to spread very quickly, it's not time to raise a public health alarm until several important questions have been answered, such as whether the variant is spreading due to specific events, or if it's happening because the mutation has allowed the virus to infect people more efficiently. Kamil offered a hypothetical scenario to explain: Imagine that a member of a community became infected and the virus mutated. That person went to church and three more people were infected, but one of them went to a karaoke bar and while singing infected 100 other people. Examining the conditions under which the virus has spread is, therefore, an essential part of untangling whether a mutation itself made the virus more transmissible or if an infected person's behaviors contributed to a local outbreak.
One of the tricky things about variants is recognizing the point at which they move from interesting, to concerning at a local level, to dangerous in a larger context. Genomic sequencing can help with that, but only when it's coordinated. When the same mutation occurs frequently, but is localized to one region, it's a concern, but when the same mutation happens in different places at the same time, it's much more likely that the "virus is learning that's a good mutation," explains Kamil.
The process is called convergent evolution, and it was a fascinating topic long before COVID. Just as your heritage can be traced through DNA, so can that of viruses, and when separate lineages develop similar traits it's almost like scientists can see evolution happening in real time. A mutation to SARS-CoV-2 that happens in more than one place at once is a mutation that makes it easier in some way for the virus to survive and that is when it may become alarming. The widespread, documented variants P.1 and B.1.351 are examples of convergence because they share some of the same virulent mutations despite having developed thousands of miles apart.
However, even variants that are emerging in different places at the same time don't present the kind of threat SARS-CoV-2 did in 2019. "This is nature," says Kamil. "It just means that this virus will not easily be driven to extinction or complete elimination by vaccines." Although a person who has already had COVID-19 can be reinfected with a variant, "it is almost always much milder disease" than the original infection, Kamil adds. Rather than causing full-fledged disease, variants have the potiental to "penetrate herd immunity, spreading relatively quietly among people who have developed natural immunity or been vaccinated, until the virus finds someone who has no immunity yet, and that person would be at risk of hospitalization-grade severe disease or death."
Surveillance and predictions
According to Lostroh, genomic surveillance can help scientists predict what's going to happen. "With the British strain, for instance, that's more transmissible, you can measure how fast it's doubling in the population and you can sort of tell whether we should take more measures against this mutation. Should we shut things down a little longer because that mutation is present in the population? That could be really useful if you did enough sampling in the population that you knew where it was," says Lostroh. If, for example, the more transmissible strain was present in 50 percent of cases, but in another county or state it was barely present, it would allow for rolling lockdowns instead of sweeping measures.
Variants are also extremely important when it comes to the development, manufacture, and distribution of vaccines. "You're also looking at medical countermeasures, such as whether your vaccine is still effective, or if your antiviral needs to be updated," says Lane Warmbrod, a senior analyst and research associate at Johns Hopkins Center for Health Security.
Properly funded and extensive genomic surveillance could eventually help control endemic diseases, too, like the seasonal flu, or other common respiratory infections. Kamil says he envisions a future in which genomic surveillance allows for prediction of sickness just as the weather is predicted today. "It's a 51 for infection today at the San Francisco Airport. There's been detection of some respiratory viruses," he says, offering an example. He says that if you're a vulnerable person, if you're immune-suppressed for some reason, you may want to wear a mask based on the sickness report.
The U.S. has the ability, but lacks standards
The benefits of widespread genomic surveillance are clear, and the United States certainly has the necessary technology, equipment, and personnel to carry it out. But, it's not happening at the speed and extent it needs to for the country to gain the benefits.
"The numbers are improving," said Kamil. "We're probably still at less than half a percent of all the samples that have been taken have been sequenced since the beginning of the pandemic."
Although there's no consensus on how many sequences is ideal for a robust surveillance program, modeling performed by the company Illumina suggests about 5 percent of positive tests should be sequenced. The reasons the U.S. has lagged in implementing a sequencing program are complex and varied, but solvable.
Perhaps the most important element that is currently missing is leadership. In order to conduct an effective genomic surveillance program, there need to be standards. The Johns Hopkins Center for Health Security recently published a paper with recommendations as to what kinds of elements need to be standardized in order to make the best use of sequencing technology and analysis.
"Along with which bioinformatic pipelines you're going to use to do the analyses, which sequencing strategy protocol are you going to use, what's your sampling strategy going to be, how is the data is going to be reported, what data gets reported," says Warmbrod. Currently, there's no guidance from the CDC on any of those things. So, while scientists can collect and report information, they may be collecting and reporting different information that isn't comparable, making it less useful for public health measures and vaccine updates.
Globally, one of the most important tools in making the information from genomic surveillance useful is GISAID, a platform designed for scientists to share -- and, importantly, to be credited for -- their data regarding genetic sequences of influenza. Originally, it was launched as a database of bird flu sequences, but has evolved to become an essential tool used by the WHO to make flu vaccine virus recommendations each year. Scientists who share their credentials have free access to the database, and anyone who uses information from the database must credit the scientist who uploaded that information.
Safety, logistics, and funding matter
Scientists at university labs and other small organizations have been uploading sequences to GISAID almost from the beginning of the pandemic, but their funding is generally limited, and there are no standards regarding information collection or reporting. Private, for-profit labs haven't had motivation to set up sequencing programs, although many of them have the logistical capabilities and funding to do so. Public health departments are understaffed, underfunded, and overwhelmed.
University labs may also be limited by safety concerns. The SARS-CoV-2 virus is dangerous, and there's a question of how samples should be transported to labs for sequencing.
Larger, for-profit organizations often have the tools and distribution capabilities to safely collect and sequence samples, but there hasn't been a profit motive. Genomic sequencing is less expensive now than ever before, but even at $100 per sample, the cost adds up -- not to mention the cost of employing a scientist with the proper credentials to analyze the sequence.
The path forward
The recently passed COVID-19 relief bill does have some funding to address genomic sequencing. Specifically, the American Rescue Plan Act includes $1.75 billion in funding for the Centers for Disease Control and Prevention's Advanced Molecular Detection (AMD) program. In an interview last month, CDC Director Rochelle Walensky said that the additional funding will be "a dial. And we're going to need to dial it up." AMD has already announced a collaboration called the Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance (SPHERES) Initiative that will bring together scientists from public health, academic, clinical, and non-profit laboratories across the country with the goal of accelerating sequencing.
Such a collaboration is a step toward following the recommendations in the paper Warmbrod coauthored. Building capacity now, creating a network of labs, and standardizing procedures will mean improved health in the future. "I want to be optimistic," she says. "The good news is there are a lot of passionate, smart, capable people who are continuing to work with government and work with different stakeholders." She cautions, however, that without a national strategy we won't succeed.
"If we maximize the potential and create that framework now, we can also use it for endemic diseases," she says. "It's a very helpful system for more than COVID if we're smart in how we plan it."
In this week's Friday Five, an old diabetes drug finds an exciting new purpose. Plus, how to make the cities of the future less toxic, making old mice younger with EVs, a new reason for mysterious stillbirths - and much more.
The Friday Five covers five stories in research that you may have missed this week. There are plenty of controversies and troubling ethical issues in science – and we get into many of them in our online magazine – but this news roundup focuses on scientific creativity and progress to give you a therapeutic dose of inspiration headed into the weekend.
Here is the promising research covered in this week's Friday Five:
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
- How to make cities of the future less noisy
- An old diabetes drug could have a new purpose: treating an irregular heartbeat
- A new reason for mysterious stillbirths
- Making old mice younger with EVs
- No pain - or mucus - no gain
And an honorable mention this week: How treatments for depression can change the structure of the brain
Researchers at Stanford have found that the virus that causes Covid-19 can infect fat cells, which could help explain why obesity is linked to worse outcomes for those who catch Covid-19.
Obesity is a risk factor for worse outcomes for a variety of medical conditions ranging from cancer to Covid-19. Most experts attribute it simply to underlying low-grade inflammation and added weight that make breathing more difficult.
Now researchers have found a more direct reason: SARS-CoV-2, the virus that causes Covid-19, can infect adipocytes, more commonly known as fat cells, and macrophages, immune cells that are part of the broader matrix of cells that support fat tissue. Stanford University researchers Catherine Blish and Tracey McLaughlin are senior authors of the study.
Most of us think of fat as the spare tire that can accumulate around the middle as we age, but fat also is present closer to most internal organs. McLaughlin's research has focused on epicardial fat, “which sits right on top of the heart with no physical barrier at all,” she says. So if that fat got infected and inflamed, it might directly affect the heart.” That could help explain cardiovascular problems associated with Covid-19 infections.
Looking at tissue taken from autopsy, there was evidence of SARS-CoV-2 virus inside the fat cells as well as surrounding inflammation. In fat cells and immune cells harvested from health humans, infection in the laboratory drove "an inflammatory response, particularly in the macrophages…They secreted proteins that are typically seen in a cytokine storm” where the immune response runs amok with potential life-threatening consequences. This suggests to McLaughlin “that there could be a regional and even a systemic inflammatory response following infection in fat.”
It is easy to see how the airborne SARS-CoV-2 virus infects the nose and lungs, but how does it get into fat tissue? That is a mystery and the source of ample speculation.
The macrophages studied by McLaughlin and Blish were spewing out inflammatory proteins, While the the virus within them was replicating, the new viral particles were not able to replicate within those cells. It was a different story in the fat cells. “When [the virus] gets into the fat cells, it not only replicates, it's a productive infection, which means the resulting viral particles can infect another cell,” including microphages, McLaughlin explains. It seems to be a symbiotic tango of the virus between the two cell types that keeps the cycle going.
It is easy to see how the airborne SARS-CoV-2 virus infects the nose and lungs, but how does it get into fat tissue? That is a mystery and the source of ample speculation.
Macrophages are mobile; they engulf and carry invading pathogens to lymphoid tissue in the lymph nodes, tonsils and elsewhere in the body to alert T cells of the immune system to the pathogen. Perhaps some of them also carry the virus through the bloodstream to more distant tissue.
ACE2 receptors are the means by which SARS-CoV-2 latches on to and enters most cells. They are not thought to be common on fat cells, so initially most researchers thought it unlikely they would become infected.
However, while some cell receptors always sit on the surface of the cell, other receptors are expressed on the surface only under certain conditions. Philipp Scherer, a professor of internal medicine and director of the Touchstone Diabetes Center at the University of Texas Southwestern Medical Center, suggests that, in people who have obesity, “There might be higher levels of dysfunctional [fat cells] that facilitate entry of the virus,” either through transiently expressed ACE2 or other receptors. Inflammatory proteins generated by macrophages might contribute to this process.
Another hypothesis is that viral RNA might be smuggled into fat cells as cargo in small bits of material called extracellular vesicles, or EVs, that can travel between cells. Other researchers have shown that when EVs express ACE2 receptors, they can act as decoys for SARS-CoV-2, where the virus binds to them rather than a cell. These scientists are working to create drugs that mimic this decoy effect as an approach to therapy.
Do fat cells play a role in Long Covid? “Fat cells are a great place to hide. You have all the energy you need and fat cells turn over very slowly; they have a half-life of ten years,” says Scherer. Observational studies suggest that acute Covid-19 can trigger the onset of diabetes especially in people who are overweight, and that patients taking medicines to regulate their diabetes “were actually quite protective” against acute Covid-19. Scherer has funding to study the risks and benefits of those drugs in animal models of Long Covid.
McLaughlin says there are two areas of potential concern with fat tissue and Long Covid. One is that this tissue might serve as a “big reservoir where the virus continues to replicate and is sent out” to other parts of the body. The second is that inflammation due to infected fat cells and macrophages can result in fibrosis or scar tissue forming around organs, inhibiting their function. Once scar tissue forms, the tissue damage becomes more difficult to repair.
Current Covid-19 treatments work by stopping the virus from entering cells through the ACE2 receptor, so they likely would have no effect on virus that uses a different mechanism. That means another approach will have to be developed to complement the treatments we already have. So the best advice McLaughlin can offer today is to keep current on vaccinations and boosters and lose weight to reduce the risk associated with obesity.