Is Finding Out Your Baby’s Genetics A New Responsibility of Parenting?
A doctor pricks the heel of a newborn for a blood test.
Hours after a baby is born, its heel is pricked with a lancet. Drops of the infant's blood are collected on a porous card, which is then mailed to a state laboratory. The dried blood spots are screened for around thirty conditions, including phenylketonuria (PKU), the metabolic disorder that kick-started this kind of newborn screening over 60 years ago. In the U.S., parents are not asked for permission to screen their child. Newborn screening programs are public health programs, and the assumption is that no good parent would refuse a screening test that could identify a serious yet treatable condition in their baby.
Learning as much as you can about your child's health might seem like a natural obligation of parenting. But it's an assumption that I think needs to be much more closely examined.
Today, with the introduction of genome sequencing into clinical medicine, some are asking whether newborn screening goes far enough. As the cost of sequencing falls, should parents take a more expansive look at their children's health, learning not just whether they have a rare but treatable childhood condition, but also whether they are at risk for untreatable conditions or for diseases that, if they occur at all, will strike only in adulthood? Should genome sequencing be a part of every newborn's care?
It's an idea that appeals to Anne Wojcicki, the founder and CEO of the direct-to-consumer genetic testing company 23andMe, who in a 2016 interview with The Guardian newspaper predicted that having newborns tested would soon be considered standard practice—"as critical as testing your cholesterol"—and a new responsibility of parenting. Wojcicki isn't the only one excited to see everyone's genes examined at birth. Francis Collins, director of the National Institutes of Health and perhaps the most prominent advocate of genomics in the United States, has written that he is "almost certain … that whole-genome sequencing will become part of new-born screening in the next few years." Whether that would happen through state-mandated screening programs, or as part of routine pediatric care—or perhaps as a direct-to-consumer service that parents purchase at birth or receive as a baby-shower gift—is not clear.
Learning as much as you can about your child's health might seem like a natural obligation of parenting. But it's an assumption that I think needs to be much more closely examined, both because the results that genome sequencing can return are more complex and more uncertain than one might expect, and because parents are not actually responsible for their child's lifelong health and well-being.
What is a parent supposed to do about such a risk except worry?
Existing newborn screening tests look for the presence of rare conditions that, if identified early in life, before the child shows any symptoms, can be effectively treated. Sequencing could identify many of these same kinds of conditions (and it might be a good tool if it could be targeted to those conditions alone), but it would also identify gene variants that confer an increased risk rather than a certainty of disease. Occasionally that increased risk will be significant. About 12 percent of women in the general population will develop breast cancer during their lives, while those who have a harmful BRCA1 or BRCA2 gene variant have around a 70 percent chance of developing the disease. But for many—perhaps most—conditions, the increased risk associated with a particular gene variant will be very small. Researchers have identified over 600 genes that appear to be associated with schizophrenia, for example, but any one of those confers only a tiny increase in risk for the disorder. What is a parent supposed to do about such a risk except worry?
Sequencing results are uncertain in other important ways as well. While we now have the ability to map the genome—to create a read-out of the pairs of genetic letters that make up a person's DNA—we are still learning what most of it means for a person's health and well-being. Researchers even have a name for gene variants they think might be associated with a disease or disorder, but for which they don't have enough evidence to be sure. They are called "variants of unknown (or uncertain) significance (VUS), and they pop up in most people's sequencing results. In cancer genetics, where much research has been done, about 1 in 5 gene variants are reclassified over time. Most are downgraded, which means that a good number of VUS are eventually designated benign.
While one parent might reasonably decide to learn about their child's risk for a condition about which nothing can be done medically, a different, yet still thoroughly reasonable, parent might prefer to remain ignorant so that they can enjoy the time before their child is afflicted.
Then there's the puzzle of what to do about results that show increased risk or even certainty for a condition that we have no idea how to prevent. Some genomics advocates argue that even if a result is not "medically actionable," it might have "personal utility" because it allows parents to plan for their child's future needs, to enroll them in research, or to connect with other families whose children carry the same genetic marker.
Finding a certain gene variant in one child might inform parents' decisions about whether to have another—and if they do, about whether to use reproductive technologies or prenatal testing to select against that variant in a future child. I have no doubt that for some parents these personal utility arguments are persuasive, but notice how far we've now strayed from the serious yet treatable conditions that motivated governments to set up newborn screening programs, and to mandate such testing for all.
Which brings me to the other problem with the call for sequencing newborn babies: the idea that even if it's not what the law requires, it's what good parents should do. That idea is very compelling when we're talking about sequencing results that show a serious threat to the child's health, especially when interventions are available to prevent or treat that condition. But as I have shown, many sequencing results are not of this type.
While one parent might reasonably decide to learn about their child's risk for a condition about which nothing can be done medically, a different, yet still thoroughly reasonable, parent might prefer to remain ignorant so that they can enjoy the time before their child is afflicted. This parent might decide that the worry—and the hypervigilence it could inspire in them—is not in their child's best interest, or indeed in their own. This parent might also think that it should be up to the child, when he or she is older, to decide whether to learn about his or her risk for adult-onset conditions, especially given that many adults at high familial risk for conditions like Alzheimer's or Huntington's disease choose never to be tested. This parent will value the child's future autonomy and right not to know more than they value the chance to prepare for a health risk that won't strike the child until 40 or 50 years in the future.
Parents are not obligated to learn about their children's risk for a condition that cannot be prevented, has a small risk of occurring, or that would appear only in adulthood.
Contemporary understandings of parenting are famously demanding. We are asked to do everything within our power to advance our children's health and well-being—to act always in our children's best interests. Against that backdrop, the need to sequence every newborn baby's genome might seem obvious. But we should be skeptical. Many sequencing results are complex and uncertain. Parents are not obligated to learn about their children's risk for a condition that cannot be prevented, has a small risk of occurring, or that would appear only in adulthood. To suggest otherwise is to stretch parental responsibilities beyond the realm of childhood and beyond factors that parents can control.
The Mind-Blowing Promise of Neural Implants
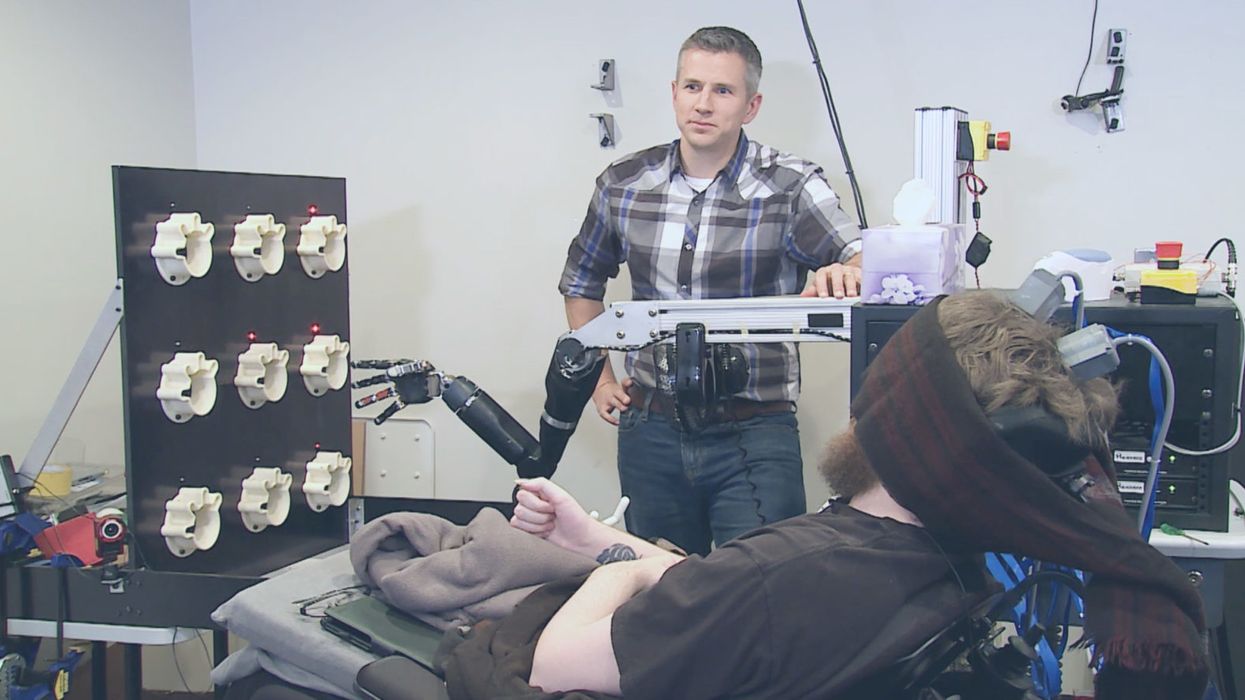
A patient with an implanted neural device that connects to a prosthetic arm can sense, while blindfolded, which of the mechanical fingers are being touched.
You may not have heard of DARPA, the research branch of the Pentagon. But you're definitely familiar with some of the technology it has pioneered, like the Internet, Siri, and handheld GPS.
"Now we're going to try to go from this proof-of-concept all the way to commercial technologies that can powerfully affect patients' lives."
Last week in National Harbor, Maryland, DARPA celebrated its 60th anniversary by showcasing its latest breakthroughs and emerging research programs, one of which centers around using neurotechnology to enhance the capabilities of the human brain. This technology is initially being developed to help warfighters and veterans, but its success could have enormous implications for civilian patients and, eventually, mainstream consumers.
The field is moving ahead rapidly. Fifteen years ago, a monkey named Aurora used a brain-machine interface to control a cursor on a computer screen. In 2014, DARPA's mind-controlled prosthetic arm for amputees won approval from the Food and Drug Administration.
Since then, DARPA has continued to push neurotechnology to new heights. Here are three of their research programs that are showing promise in early human testing:
1) A NEURAL IMPLANT HELP MANAGE PSYCHIATRIC ILLNESS
More than 2.2 million veterans and 44 million civilians are living with some form of psychiatric illness, and medications don't work for everyone. DARPA set out to create new options for people living with debilitating anxiety, depression, and PTSD.
"We can get somebody back to normal. It's a whole new set of tools for physicians," said Justin Sanchez, Director of the Biological Technologies Office at DARPA.
He told the audience about a woman living with both epilepsy and extreme anxiety, who has a direct neural interface that reads her brain's signals in real time and can be modulated with stimulation. He shared a recent video of her testing the device:
"Now we're going to try to go from this proof-of-concept all the way to commercial technologies that can powerfully affect patients' lives," Sanchez said.
2) A NEURAL IMPLANT TO HELP IMPROVE MEMORY
"We are right at the cusp" of improving memory recall with direct neural interfaces, Sanchez said.
All day long, our brains shift between poor and good memory states. A brain-computer interface can read the signals of populations of neurons in the lateral temporal cortex. The device continuously monitors the state of the brain and delivers stimulation within a fraction of a second after detecting a poor memory state, to improve the person's memory performance.
The improved memory lasts only seconds, so the system "delivers stimulation as needed in a closed loop to keep the performance in a good state, because of this natural variability of performance," said Dan Rizzuto, founder of NiaTherapeutics, whose technology was developed with support from DARPA and the United States BRAIN Initiative.
Check out this recently shot video of a patient testing the device, which Sanchez called "a breakthrough moment":
About 400 patients have been tested with this technology so far. In a pilot study whose data have not yet been published, patients with traumatic brain injury showed improvement in recall of around 28 percent, according to Rizzuto.
He estimates that potential FDA approval of the device for patients with traumatic brain injury is still 7 to 8 years away. The technology holds the potential to help many other kinds of patients as well.
"We believe this device could also be used to treat Alzheimer's because it's not specific to any brain pathology but based on a deep understanding of the way human memory works," Rizzuto said.
3) A NEURAL IMPLANT TO REVOLUTIONIZE PROSTHETICS FOR WARFIGHTERS AND VETERANS
Since 2006, DARPA has run a program to revolutionize prosthetics. The latest advances allow amputees to actually feel again with their bionic limbs.
Sensors in a prosthetic hand relay information to an interface in the brain that allows the person to detect which of their "fingers" are being touched, while their eyes are closed:
WHAT COMES NEXT?
DARPA is now turning its attention to non-surgical, non-invasive neurotechnology. Researchers hope to use advanced sensor technology to detect signals from neurons without putting any electrodes directly inside the brain. Under the direction of program manager Dr. Al Emondi, the N³ program is about to launch soon and plans to run for four or five years.
"We haven't even scratched the surface of what a human brain's capability is," said Dr. Geoffrey Ling, the Founding Director of the Biological Technologies Office. "When we can make this a non-invasive consumer technology, this will explode. It will take on a life of its own."
Then, inevitably, the hard questions will follow.
As Sanchez put it: "Will society consider some form of neural enhancement a personal choice like braces? Could there be a disturbing gap for people who have neurotech and those who don't? We must come together and all think over the horizon. How the story unfolds ultimately depends on all of us."
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
Prostate Cancer Treatments Are Racing Ahead. Here’s a Big Reason Why.
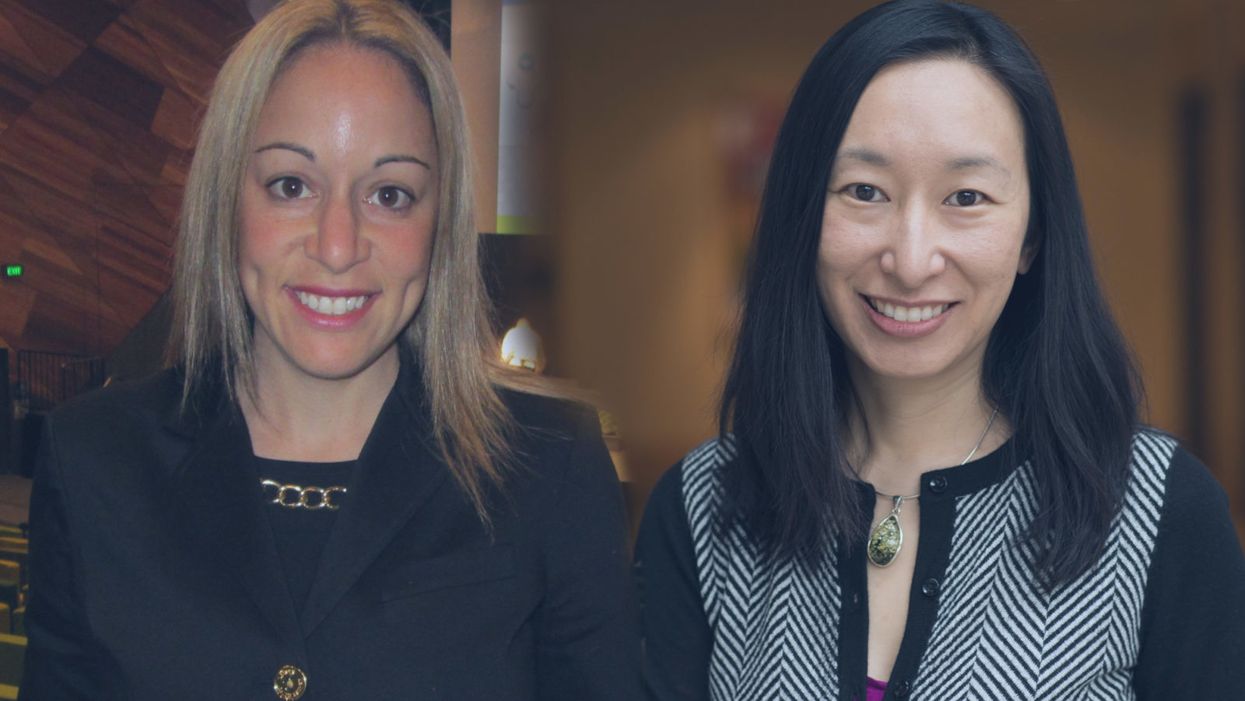
Dr. Stacy Loeb (left) and Dr. Heather Cheng were both recipients of a special early-career grant from the Prostate Cancer Foundation for young researchers.
In his lab at UCLA, Dr. Charles Sawyer discovered two drugs for metastatic prostate cancer that are now in routine use all over the world.
At the University of Washington at Seattle, Dr. Heather Cheng was part of a team that discovered the connection between BRCA2 mutations and advanced prostate cancer, and she recently opened a prostate cancer genetics clinic – a new frontier in the field.
At UT Southwestern Medical Center in Dallas, Dr. Nima Sharifi's pioneering research showed why certain drugs don't work in castrate-resistant prostate cancer, and now new therapies are being developed instead.
"We have good reason to believe that investing in young scientists is the way to go."
What Do These Researchers Share in Common?
They were all under 40 when they received a special grant for early-career scientists from the Prostate Cancer Foundation, the leading philanthropic organization that funds prostate cancer research. Experts say that the foundation's dedicated support for young innovators has been a game changer in contributing to the discovery of newer and better therapies for prostate cancer patients.
Howard Soule, the foundation's Executive Vice President and Chief Science Officer, was aware that many of the people who leave behind major legacies in science typically make their discoveries before age 40, like Albert Einstein, who was in his thirties when he published his paper on general relativity.
So back in 2007, the PCF decided to ramp up its support for young researchers.
"We have good reason to believe that investing in young scientists is the way to go, so we've created a program at PCF that is I believe is unique in the field," says Soule.
The Young Investigator Awards Program rigorously screens a pool of roughly 150 applicants for 20 to 25 awards that consist of funding for three years – and that's just the start.
"It's much more than sending them money," says Soule. "We celebrate them at annual meetings, we have a networking center with no equal in the field, and throughout the years of their three-year-award and basically forever, we create community. We are a safe place for them to land, they share data with us that's unpublished, and we provide a lot of feedback and stewardship to their donors."
Spotlighting Recipients: From the Study of Tumors to Twitter
Heather Cheng was in her thirties when she received her award three years ago. "It's been very, very helpful in allowing me to do the type of work I am really excited about doing," she says.
At the time, she had recently joined the faculty at the University of Washington after completing an MD/PhD medical scientist training program, internal medicine residency and hematology/oncology fellowship, and she was considering what new direction to take in her research. Several patients captured her imagination who were diagnosed at a very young age with metastatic prostate cancer, and "even though we had cool new drugs to extend life, these particular patients' cancers blew through everything."
"This is a new intersection because genetics has not been discussed in the context of care for men with prostate cancer that much."
She decided to make a niche out of understanding the connection between often early-onset aggressive prostate cancer and familial genetic risk, in order to improve treatment options for these patients. In 2016, Cheng launched a new clinic and invited any men to visit who have a family history of cancer and who are interested in genetic testing, or who have a known mutation and want to learn about treatment opportunities, or who want to know if their cancer tumor can be inherited.
"It's an open door to have a discussion because the technology and treatment potentials are so new," Cheng says. "There's a lot to learn."
It used to be that a doctor would ask a male patient about his family history, and if a mother had breast cancer at a young age, for example, and several other family members met the criteria for a genetic risk, then perhaps the patient had inherited a mutation in a cancer risk gene. But what to do next was unclear.
Now, doctors are taking men with a diagnosis of prostate cancer, sequencing their inherited DNA or their tumors, and finding out if they have mutations that could guide their treatment plan. For example, medications called PARP inhibitors have shown encouraging early results for men with a BRCA2 gene mutation and are now in clinical trials for treating prostate cancer.
"This is a new intersection because genetics has not been discussed in the context of care for men with prostate cancer that much," Cheng says. "This has changed practice because changes to national cancer guidelines have happened in less than five years. The change has happened so quickly that the field is not completely prepared for implementation and clinical logistics."
Another young investigator, New York University urologist and prostate cancer researcher Stacy Loeb, received her award at age 36 two years ago. She realized that no one had scientifically studied how patients are using crowdsourcing platforms like GoFundMe and YouCaring to raise money for their treatments. In her research, she found that there are many more campaigns for breast cancer and that they are more successful in crowdfunding than the prostate cancer campaigns.
"We have identified some gaps in advocacy and awareness for prostate cancer – fewer people know about it or discuss it, but it is a leading cause of death of U.S. men, so it is important to get more people aware," Loeb notes.
In fact, today the PCF releases data from a survey of more than 2,000 U.S. adults that reveals widespread ignorance about the disease. Two-thirds of respondents, for example, did not know that men with early stage prostate cancer experience no symptoms, and many were unaware that screening begins with a simple blood test.
Besides studying patient behavior, Loeb also wanted to better understand how physicians and scientists are using social media, and how their participation on platforms like Twitter could be fostered to promote greater dissemination of knowledge. So she helped start a monthly prostate cancer journal club on Twitter, hosted through the PCF science account. The club features an important new research paper in the field each month, and she invites the authors of the paper to participate in a 48-hour online discussion.
"The Journal Club is a monthly thing at most institutions," she says, "but typically it's one institution with people from one department. What's better about this is we have people who are doctors, nurses, scientists, patients, stakeholders participating from all over the world."
Why Do Young Innovators Have an Edge?
The environment matters, for one.
"We all bring different life experiences to the table, we grew up in different eras, so we have different norms and tools at our disposal that weren't available," says Loeb, who was one of the early adopters of social media in the urology space. She now gives a lecture at the annual PCF retreat on how to use social media to advance one's scientific career.
"The more you're invested into a system, the less you may be able to recognize its limitations."
But the advantage of youth is not just greater familiarity with the newest tools. It's also the existential benefit of not being entrenched in the way it's always been.
"Often there is a healthy skepticism of what's come before," explains Dr. Joseph La Brie, a clinical psychologist and professor at psychology at Loyola Marymount University. "That's connected to not being wedded to a programmatic view of the problem. There's a freshness and creative outlook because they are looking at it with a new set of eyes, and there's a desire to make their mark on the field, to be unique and innovative and not just follow in somebody else's footsteps."
And as Cheng puts it, "The more you're invested into a system, the less you may be able to recognize its limitations."
But it's notoriously difficult for scientists to get funding for innovative ideas without having already published preliminary data, which is what the National Institutes of Health and other funding bodies like to see. Eliminating that hurdle is a big part of why PCF's approach has been so effective, according to a veteran of the field, Johns Hopkins urologist Dr. Kenneth Pienta; his own groundbreaking research has been supported by PCF since he was a young scientist in the '90s.
"Any granting mechanism that allows people to fund ideas without a lot of preliminary data is a good thing," he declares.
Support for creative young minds is crucial across all endeavors, beyond any single disease or discipline. At a recent conference showcasing emerging technology for DARPA, the research arm of the Defense Department, expert panelists in artificial intelligence were asked: What is the single most important thing to focus on over the next decade?
Robotics pioneer Dr. Rodney Brooks may have put it best: "Figure out how to fund some really radical young mavericks and see what happens."
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.

