New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
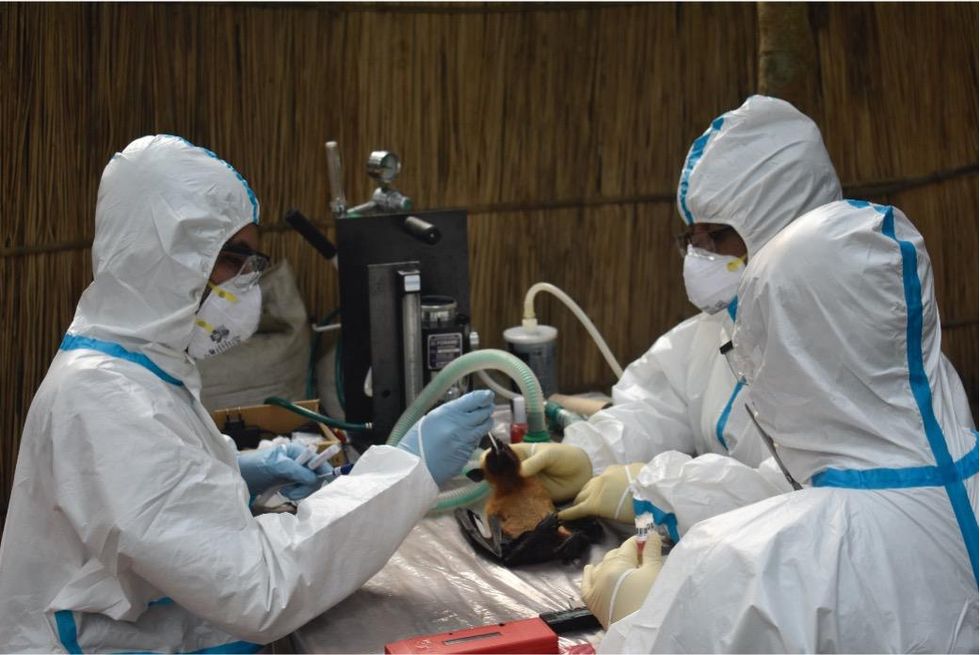
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
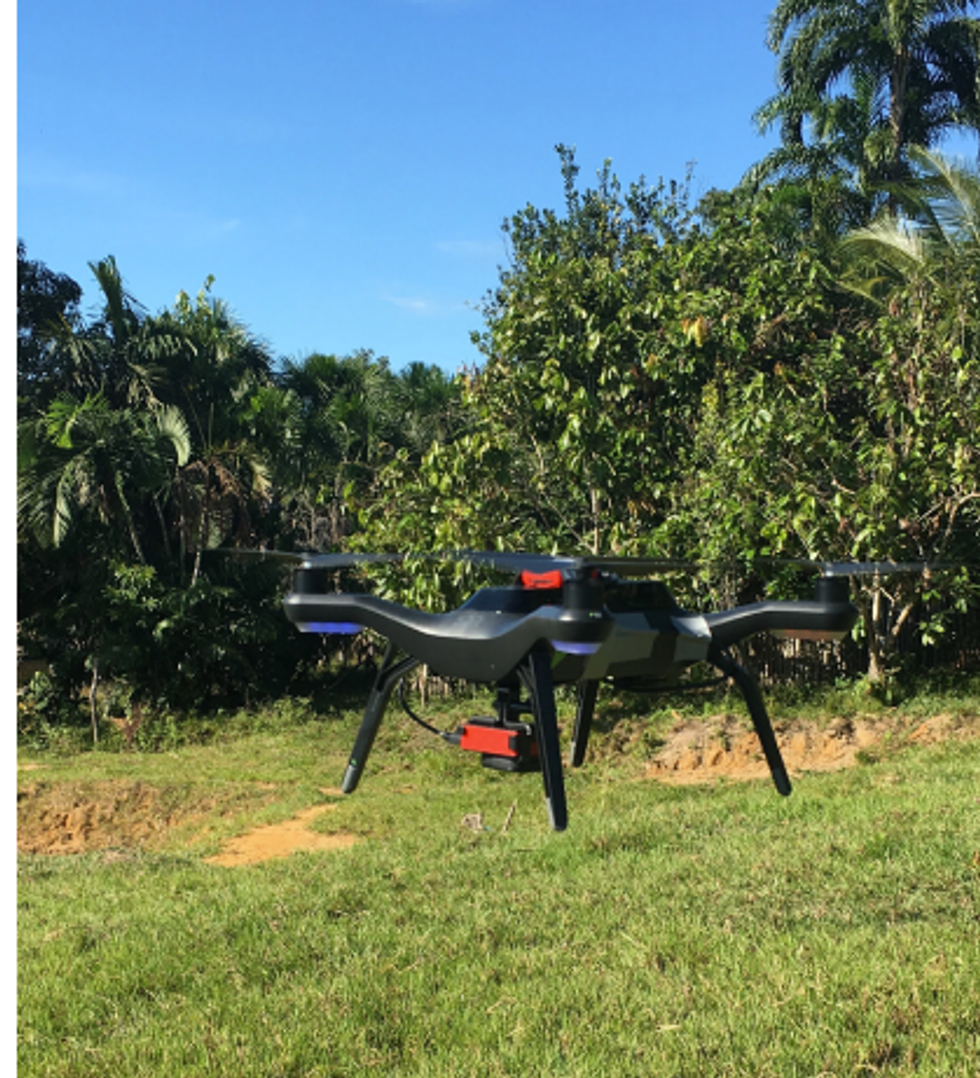

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
In this new video, we capture examples from history when people rejected scientific breakthroughs at first, only for the march of progress to continue.
What makes people turn against science? After two years of a global pandemic, the world has never felt more divided on questions of science. But this is not a new phenomenon. People have resisted scientific and technological advances throughout history.
This video by Leaps.org, with support from the Gordon and Betty Moore Foundation, captures noteworthy examples from history when people rejected science. What do these cases have in common? Scientific breakthroughs can be revolutionary, but revolutions can be disorienting and anxiety-producing. They transform our livelihoods, culture and even our understanding of what it means to be human. But there's reason for optimism. Many of history’s controversies were overcome. Science has a way of enduring, because it changes things for the better.
Scientists search for a universal coronavirus vaccine
Stephen Zeichner, an infectious disease specialist at the University of Virginia Medical Center, has made progress with an early stage universal coronavirus vaccine.
The Covid-19 pandemic had barely begun when VBI Vaccines, a biopharmaceutical company based in Cambridge, Massachusetts, initiated their search for a universal coronavirus vaccine.
It was March 2020, and while most pharmaceutical companies were scrambling to initiate vaccine programs which specifically targeted the SARS-CoV-2 virus, VBI’s executives were already keen to look at the broader picture.
Having observed the SARS and MERS coronavirus outbreaks over the last two decades, Jeff Baxter, CEO of VBI Vaccines, was aware that SARS-CoV-2 is unlikely to be the last coronavirus to move from an animal host into humans. “It's absolutely apparent that the future is to create a vaccine which gives more broad protection against not only pre-existing coronaviruses, but those that will potentially make the leap into humans in future,” says Baxter.
It was a prescient decision. Over the last two years, more biotechs and pharma companies have joined the search to find a vaccine which might be able to protect against all coronaviruses, along with dozens of academic research groups. Last September, the US National Institutes of Health dedicated $36 million specifically to pan-coronavirus vaccine research, while the global Coalition for Epidemic Preparedness Innovations (CEPI) has earmarked $200 million towards the effort.
Until October 2021, the very concept of whether it might be
theoretically possible to vaccinate against multiple coronaviruses remained an open question. But then a groundbreaking study renewed optimism.
The emergence of new variants of Covid-19 over the past year, particularly the highly mutated Omicron variant, has added greater impetus to find broader spectrum vaccines. But until October 2021, the very concept of whether it might be theoretically possible to vaccinate against multiple coronaviruses remained an open question. After all, scientists have spent decades trying to develop a similar vaccine for influenza with little success.
But then a groundbreaking study from renowned virologist Linfa Wang, who runs the emerging infectious diseases program at Duke-National University of Singapore Medical School, provided renewed optimism.
Wang found that eight SARS survivors who had been injected with the Pfizer/BioNTech Covid-19 vaccine had neutralising antibodies in their blood against SARS, the Alpha, Beta and Delta variants of SARS-CoV-2, and five other coronaviruses which reside in bats and pangolins. He concluded that the combination of past coronavirus infection, and immunization with a messenger RNA vaccine, had resulted in a wider spectrum of protection than might have been expected.
“This is a significant study because it showed that pre-existing immunity to one coronavirus could help with the elicitation of cross-reactive antibodies when immunizing with a second coronavirus,” says Kevin Saunders, Director of Research at the Duke Human Vaccine Institute in North Carolina, which is developing a universal coronavirus vaccine. “It provides a strategy to perhaps broaden the immune response against coronaviruses.”
In the next few months, some of the first data is set to emerge looking at whether this kind of antibody response could be elicited by a single universal coronavirus vaccine. In April 2021, scientists at the Walter Reed Army Institute of Research in Silver Spring, Maryland, launched a Phase I clinical trial of their vaccine, with a spokesman saying that it was successful, and the full results will be announced soon.
The Walter Reed researchers have already released preclinical data, testing the vaccine in non-human primates where it was found to have immunising capabilities against a range of Covid-19 variants as well as the original SARS virus. If the Phase I trial displays similar efficacy, a larger Phase II trial will begin later this year.
Two different approaches
Broadly speaking, scientists are taking two contrasting approaches to the task of finding a universal coronavirus vaccine. The Walter Reed Army Institute of Research, VBI Vaccines – who plan to launch their own clinical trial in the summer – and the Duke Human Vaccine Institute – who are launching a Phase I trial in early 2023 – are using a soccer-ball shaped ferritin nanoparticle studded with different coronavirus protein fragments.
VBI Vaccines is looking to elicit broader immune responses by combining SARS, SARS-CoV-2 and MERS spike proteins on the same nanoparticle. Dave Anderson, chief scientific officer at VBI Vaccines, explains that the idea is that by showing the immune system these three spike proteins at the same time, it can help train it to identify and respond to subtle differences between coronavirus strains.
The Duke Human Vaccine Institute is utilising the same method, but rather than including the entire spike proteins from different coronaviruses, they are only including the receptor binding domain (RBD) fragment from each spike protein. “We designed our vaccine to focus the immune system on a site of vulnerability for the virus, which is the receptor binding domain,” says Saunders. “Since the RBD is small, arraying multiple RBDs on a nanoparticle is a straight-forward approach. The goal is to generate immunity to many different subgenuses of viruses so that there will be cross-reactivity with new or unknown coronaviruses.”
But the other strategy is to create a vaccine which contains regions of the viral protein structure which are conserved between all coronavirus strains. This is something which scientists have tried to do for a universal influenza vaccine, but it is thought to be more feasible for coronaviruses because they mutate at a slower rate and are more constrained in the ways that they can evolve.
DIOSynVax, a biotech based in Cambridge, United Kingdom, announced in a press release earlier this month that they are partnering with CEPI to use their computational predictive modelling techniques to identify common structures between all of the SARS coronaviruses which do not mutate, and thus present good vaccine targets.
Stephen Zeichner, an infectious disease specialist at the University of Virginia Medical Center, has created an early stage vaccine using the fusion peptide region – another part of the coronavirus spike protein that aids the virus’s entry into host cells – which so far appears to be highly conserved between all coronaviruses.
So far Zeichner has trialled this version of the vaccine in pigs, where it provided protection against a different coronavirus called porcine epidemic diarrhea virus, which he described as very promising as this virus is from a different family called alphacoronaviruses, while SARS-CoV-2 is a betacoronavirus.
“If a betacoronavirus fusion peptide vaccine designed from SARS-CoV-2 can protect pigs against clinical disease from an alphacoronavirus, then that suggests that an analogous vaccine would enable broad protection against many, many different coronaviruses,” he says.
The road ahead
But while some of the early stage results are promising, researchers are fully aware of the scale of the challenge ahead of them. Although CEPI have declared an aim of having a licensed universal coronavirus vaccine available by 2024-2025, Zeichner says that such timelines are ambitious in the extreme.
“I was incredibly impressed at the speed at which the mRNA coronavirus vaccines were developed for SARS-CoV-2,” he says. “That was faster than just about anybody anticipated. On the other hand, I think a universal coronavirus vaccine is more equivalent to the challenge of developing an HIV vaccine and we're 35 years into that effort without success. We know a lot more now than before, and maybe it will be easier than we think. But I think the route to a universal vaccine is harder than an individual vaccine, so I wouldn’t want to put money on a timeline prediction.”
The major challenge for scientists is essentially designing a vaccine for a future threat which is not even here yet. As such, there are no guidelines on what safety data would be required to license such a vaccine, and how researchers can demonstrate that it truly provides efficacy against all coronaviruses, even those which have not yet jumped to humans.
The teams working on this problem have already devised some ingenious ways of approaching the challenge. VBI Vaccines have taken the genetic sequences of different coronaviruses found in bats and pangolins, from publicly available databases, and inserted them into what virologists call a pseudotype virus – one which has been engineered so it does not have enough genetic material to replicate.
This has allowed them to test the neutralising antibodies that their vaccine produces against these coronaviruses in test tubes, under safe lab conditions. “We have literally just been ordering the sequences, and making synthetic viruses that we can use to test the antibody responses,” says Anderson.
However, some scientists feel that going straight to a universal coronavirus vaccine is likely to be too complex. Instead they say that we should aim for vaccines which are a little more specific. Pamela Bjorkman, a structural biologist at the California Institute of Technology, suggests that pan-coronavirus vaccines which protect against SARS-like betacoronaviruses such as SARS or SARS-CoV-2, or MERS-like betacoronaviruses, may be more realistic.
“I think a vaccine to protect against all coronaviruses is likely impossible since there are so many varieties,” she says. “Perhaps trying to narrow down the scope is advisable.”
But if the mission to develop a universal coronavirus vaccine does succeed, it will be one of the most remarkable feats in the annals of medical science. In January, US chief medical advisor Anthony Fauci urged for greater efforts to be devoted towards this goal, one which scientists feel would be the biological equivalent of the race to develop the first atomic bomb
“The development of an effective universal coronavirus vaccine would be equally groundbreaking, as it would have global applicability and utility,” says Saunders. “Coronaviruses have caused multiple deadly outbreaks, and it is likely that another outbreak will occur. Having a vaccine that prevents death from a future outbreak would be a tremendous achievement in global health.”
He agrees that it will require creativity on a remarkable scale: “The universal coronavirus vaccine will also require ingenuity and perseverance comparable to that needed for the Manhattan project.”


