New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
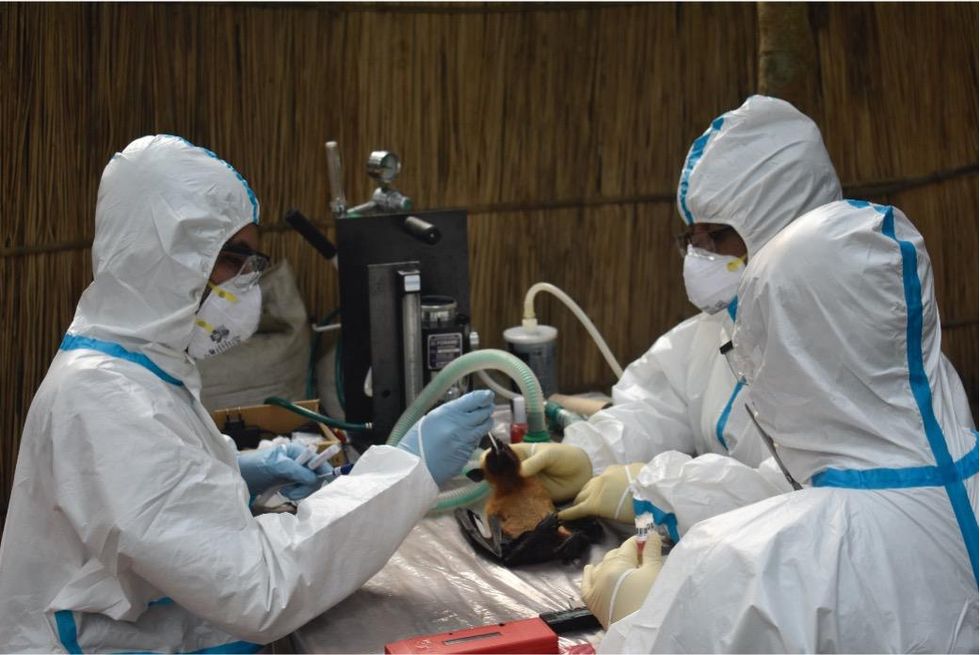
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
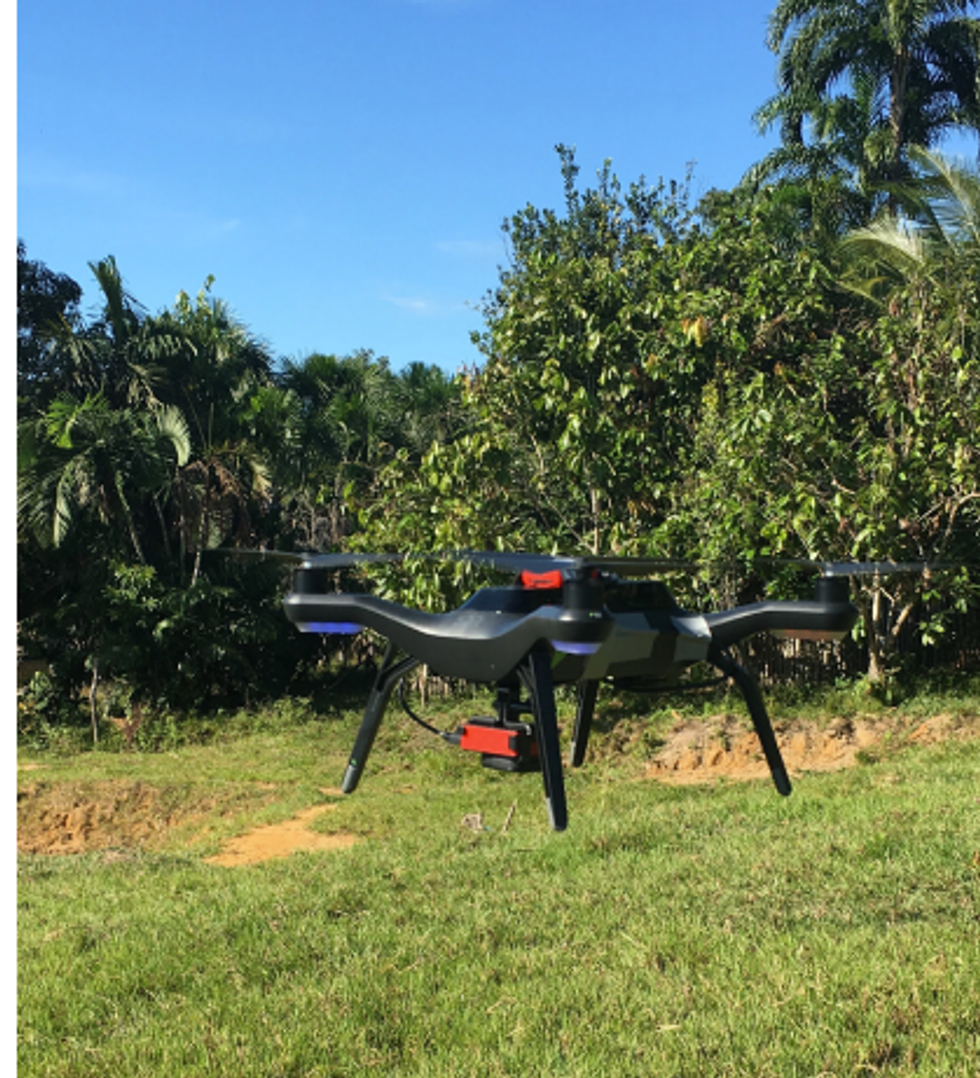

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
As gene therapies and small molecule drugs are being studied in clinical trials, companies increasingly see the value in hiring patients to help explain the potential benefits.
As a child, Wendy Borsari participated in a health study at Boston Children’s Hospital. She was involved because heart disease and sudden cardiac arrest ran in her family as far back as seven generations. When she was 18, however, the study’s doctors told her that she had a perfectly healthy heart and didn’t have to worry.
A couple of years after graduating from college, though, the Boston native began to experience episodes of near fainting. During any sort of strenuous exercise, my blood pressure would drop instead of increasing, she recalls.
She was diagnosed at 24 with hypertrophic cardiomyopathy. Although HCM is a commonly inherited heart disease, Borsari’s case resulted from a rare gene mutation, the MYH7 gene. Her mother had been diagnosed at 27, and Borsari had already lost her grandmother and two maternal uncles to the condition. After her own diagnosis, Borsari spent most of her free time researching the disease and “figuring out how to have this condition and still be the person I wanted to be,” she says.
Then, her son was found to have the genetic mutation at birth and diagnosed with HCM at 15. Her daughter, also diagnosed at birth, later suffered five cardiac arrests.
That changed Borsari’s perspective. She decided to become a patient advocate. “I didn’t want to just be a patient with the condition,” she says. “I wanted to be more involved with the science and the biopharmaceutical industry so I could be active in helping to make it better for other patients.”
She consulted on patient advocacy for a pharmaceutical and two foundations before coming to a company called Tenaya in 2021.
“One of our core values as a company is putting patients first,” says Tenaya's CEO, Faraz Ali. “We thought of no better way to put our money where our mouth is than by bringing in somebody who is affected and whose family is affected by a genetic form of cardiomyopathy to have them make sure we’re incorporating the voice of the patient.”
Biomedical corporations and government research agencies are now incorporating patient advocacy more than ever, says Alice Lara, president and CEO of the Sudden Arrhythmia Death Syndromes Foundation in Salt Lake City, Utah. These organizations have seen the effectiveness of including patient voices to communicate and exemplify the benefits that key academic research institutions have shown in their medical studies.
“From our side of the aisle,” Lara says, “what we know as patient advocacy organizations is that educated patients do a lot better. They have a better course in their therapy and their condition, and understanding the genetics is important because all of our conditions are genetic.”
Founded in 2016, Tenaya is advancing gene therapies and small molecule drugs in clinical trials for both prevalent and rare forms of heart disease, says Ali, the CEO.
The firm's first small molecule, now in a Phase 1 clinical trial, is intended to treat heart failure with preserved ejection fraction, where the amount of blood pumped by the heart is reduced due to the heart chambers becoming weak or stiff. The condition accounts for half or more of all heart failure in the U.S., according to Ali, and is growing quickly because it's closely associated with diabetes. It’s also linked with metabolic syndrome, or a cluster of conditions including high blood pressure, high blood sugar, excess body fat around the waist, and abnormal cholesterol levels.
“We have a novel molecule that is first in class and, to our knowledge, best in class to tackle that, so we’re very excited about the clinical trial,” Ali says.
The first phase of the trial is being performed with healthy participants, rather than people with the disease, to establish safety and tolerability. The researchers can also look for the drug in blood samples, which could tell them whether it's reaching its target. Ali estimates that, if the company can establish safety and that it engages the right parts of the body, it will likely begin dosing patients with the disease in 2024.
Tenaya’s therapy delivers a healthy copy of the gene so that it makes a copy of the protein missing from the patients' hearts because of their mutation. The study will start with adult patients, then pivot potentially to children and even newborns, Ali says, “where there is an even greater unmet need because the disease progresses so fast that they have no options.”
Although this work still has a long way to go, Ali is excited about the potential because the gene therapy achieved positive results in the preclinical mouse trial. This animal trial demonstrated that the treatment reduced enlarged hearts, reversed electrophysiological abnormalities, and improved the functioning of the heart by increasing the ejection fraction after the single-dose of gene therapy. That measurement remained stable to the end of the animals’ lives, roughly 18 months, Ali says.
He’s also energized by the fact that heart disease has “taken a page out of the oncology playbook” by leveraging genetic research to develop more precise and targeted drugs and gene therapies.
“Now we are talking about a potential cure of a disease for which there was no cure and using a very novel concept,” says Melind Desai of the Cleveland Clinic.
Tenaya’s second program focuses on developing a gene therapy to mitigate the leading cause of hypertrophic cardiomyopathy through a specific gene called MYPBC3. The disease affects approximately 600,000 patients in the U.S. This particular genetic form, Ali explains, affects about 115,000 in the U.S. alone, so it is considered a rare disease.
“There are infants who are dying within the first weeks to months of life as a result of this mutation,” he says. “There are also adults who start having symptoms in their 20s, 30s and 40s with early morbidity and mortality.” Tenaya plans to apply before the end of this year to get the FDA’s approval to administer an investigational drug for this disease humans. If approved, the company will begin to dose patients in 2023.
“We now understand the genetics of the heart much better,” he says. “We now understand the leading genetic causes of hypertrophic myopathy, dilated cardiomyopathy and others, so that gives us the ability to take these large populations and stratify them rationally into subpopulations.”
Melind Desai, MD, who directs Cleveland Clinic’s Hypertrophic Cardiomyopathy Center, says that the goal of Tenaya’s second clinical study is to help improve the basic cardiac structure in patients with hypertrophic cardiomyopathy related to the MYPBC3 mutation.
“Now we are talking about a potential cure of a disease for which there was no cure and using a very novel concept,” he says. “So this is an exciting new frontier of therapeutic investigation for MYPBC3 gene-positive patients with a chance for a cure.
Neither of Tenaya’s two therapies address the gene mutation that has affected Borsari and her family. But Ali sees opportunity down the road to develop a gene therapy for her particular gene mutation, since it is the second leading cause of cardiomyopathy. Treating the MYH7 gene is especially challenging because it requires gene editing or silencing, instead of just replacing the gene.

Wendy Borsari was diagnosed at age 24 with a commonly inherited heart disease. She joined Tenaya as a patient advocate in 2021.
Wendy Borsari
“If you add a healthy gene it will produce healthy copies,” Ali explains, “but it won’t stop the bad effects of the mutant protein the gene produces. You can only do that by silencing the gene or editing it out, which is a different, more complicated approach.”
Euan Ashley, professor of medicine and genetics at Stanford University and founding director of its Center for Inherited Cardiovascular Disease, is confident that we will see genetic therapies for heart disease within the next decade.
“We are at this really exciting moment in time where we have diseases that have been under-recognized and undervalued now being attacked by multiple companies with really modern tools,” says Ashley, author of The Genome Odyssey. “Gene therapies are unusual in the sense that they can reverse the cause of the disease, so we have the enticing possibility of actually reversing or maybe even curing these diseases.”
Although no one is doing extensive research into a gene therapy for her particular mutation yet, Borsari remains hopeful, knowing that companies such as Tenaya are moving in that direction.
“I know that’s now on the horizon,” she says. “It’s not just some pipe dream, but will happen hopefully in my lifetime or my kids’ lifetime to help them.”
Chicken that is grown entirely in a laboratory, without harming a single bird, could be sold in supermarkets in the coming months. But critics say the doubts about lab-grown meat have not been appropriately explored.
Last November, when the U.S. Food and Drug Administration disclosed that chicken from a California firm called UPSIDE Foods did not raise safety concerns, it drily upended how humans have obtained animal protein for thousands of generations.
“The FDA is ready to work with additional firms developing cultured animal cell food and production processes to ensure their food is safe and lawful,” the agency said in a statement at the time.
Assuming UPSIDE obtains clearances from the U.S. Department of Agriculture, its chicken – grown entirely in a laboratory without harming a single bird – could be sold in supermarkets in the coming months.
“Ultimately, we want our products to be available everywhere meat is sold, including retail and food service channels,” a company spokesperson said. The upscale French restaurant Atelier Crenn in San Francisco will have UPSIDE chicken on its menu once it is approved, she added.
Known as lab-grown or cultured meat, a product such as UPSIDE’s is created using stem cells and other tissue obtained from a chicken, cow or other livestock. Those cells are then multiplied in a nutrient-dense environment, usually in conjunction with a “scaffold” of plant-based materials or gelatin to give them a familiar form, such as a chicken breast or a ribeye steak. A Dutch company called Mosa Meat claims it can produce 80,000 hamburgers derived from a cluster of tissue the size of a sesame seed.
Critics say the doubts about lab-grown meat and the possibility it could merge “Brave New World” with “The Jungle” and “Soylent Green” have not been appropriately explored.
That’s a far cry from when it took months of work to create the first lab-grown hamburger a decade ago. That minuscule patty – which did not contain any fat and was literally plucked from a Petri dish to go into a frying pan – cost about $325,000 to produce.
Just a decade later, an Israeli company called Future Meat said it can produce lab-grown meat for about $1.70 per pound. It plans to open a production facility in the U.S. sometime in 2023 and distribute its products under the brand name “Believer.”
Costs for production have sunk so low that researchers at Carnegie Mellon University in Pittsburgh expect sometime in early 2024 to produce lab-grown Wagyu steak to showcase the viability of growing high-end cuts of beef cheaply. The Carnegie Mellon team is producing its Wagyu using a consumer 3-D printer bought secondhand on eBay and modified to print the highly marbled flesh using a method developed by the university. The device costs $200 – about the same as a pound of Wagyu in the U.S. The initiative’s modest five-figure budget was successfully crowdfunded last year.
“The big cost is going to be the cells (which are being extracted by a cow somewhere in Pennsylvania), but otherwise printing doesn’t add much to the process,” said Rosalyn Abbott, a Carnegie Mellon assistant professor of bioengineering who is co-leader on the project. “But it adds value, unlike doing this with ground meat.”
Lab-Grown Meat’s Promise
Proponents of lab-grown meat say it will cut down on traditional agriculture, which has been a leading contributor to deforestation, water shortages and contaminated waterways from animal waste, as well as climate change.
An Oxford University study from 2011 concludes lab-grown meat could have greenhouse emissions 96 percent lower compared to traditionally raised livestock. Moreover, proponents of lab-grown meat claim that the suffering of animals would decline dramatically, as they would no longer need to be warehoused and slaughtered. A recently opened 26-story high-rise in China dedicated to the raising and slaughtering of pigs illustrates the current plight of livestock in stark terms.
Scientists may even learn how to tweak lab-grown meat to make it more nutritious. Natural red meat is high in saturated fat and, if it’s eaten too often, can lead to chronic diseases. In lab versions, the saturated fat could be swapped for healthier, omega-3 fatty acids.
But critics say the doubts about lab-grown meat and the possibility it could merge “Brave New World” with “The Jungle” and “Soylent Green” have not been appropriately explored.
A Slippery Slope?
Some academics who have studied the moral and ethical issues surrounding lab-grown meat believe it will have a tough path ahead gaining acceptance by consumers. Should it actually succeed in gaining acceptance, many ethical questions must be answered.
“People might be interested” in lab-grown meat, perhaps as a curiosity, said Carlos Alvaro, an associate professor of philosophy at the New York City College of Technology, part of the City University of New York. But the allure of traditionally sourced meat has been baked – or perhaps grilled – into people’s minds for so long that they may not want to make the switch. Plant-based meat provides a recent example of the uphill battle involved in changing old food habits, with Beyond Meat’s stock prices dipping nearly 80 percent in 2022.
"There are many studies showing that people don’t really care about the environment (to that extent)," Alvaro said. "So I don’t know how you would convince people to do this because of the environment.”
“From my research, I understand that the taste (of lab-grown meat) is not quite there,” Alvaro said, noting that the amino acids, sugars and other nutrients required to grow cultivated meat do not mimic what livestock are fed. He also observed that the multiplication of cells as part of the process “really mimic cancer cells” in the way they grow, another off-putting thought for would-be consumers of the product.
Alvaro is also convinced the public will not buy into any argument that lab-grown meat is more environmentally friendly.
“If people care about the environment, they either try and consume considerably less meat and other animal products, or they go vegan or vegetarian,” he said. “But there are many studies showing that people don’t really care about the environment (to that extent). So I don’t know how you would convince people to do this because of the environment.”
Ben Bramble, a professor at Australian National University who previously held posts at Princeton and Trinity College in Ireland, takes a slightly different tack. He noted that “if lab-grown meat becomes cheaper, healthier, or tastier than regular meat, there will be a large market for it. If it becomes all of these things, it will dominate the market.”
However, Bramble has misgivings about that occurring. He believes a smooth transition from traditionally sourced meat to a lab-grown version would allow humans to elide over the decades of animal cruelty perpetrated by large-scale agriculture, without fully reckoning with and learning from this injustice.
“My fear is that if we all switch over to lab-grown meat because it has become cheaper, healthier, or tastier than regular meat, we might never come to realize what we have done, and the terrible things we are capable of,” he said. “This would be a catastrophe.”
Bramble’s writings about cultured meat also raise some serious moral conundrums. If, for example, animal meat may be cultivated without killing animals, why not create products from human protein?
Actually, that’s already happened.
It occurred in 2019, when Orkan Telhan, a professor of fine arts at the University of Pennsylvania, collaborated with two scientists to create an art exhibit at the Philadelphia Museum of Art on the future of foodstuffs.
Although the exhibit included bioengineered bread and genetically modified salmon, it was an installation called “Ouroboros Steak” that drew the most attention. That was comprised of pieces of human flesh grown in a lab from cultivated cells and expired blood products obtained from online sources.
The exhibit was presented as four tiny morsels of red meat – shaped in patterns suggesting an ouroboros, a dragon eating its own tail. They were placed in tiny individual saucers atop a larger plate and placemat with a calico pattern, suggesting an item to order in a diner. The artwork drew international headlines – as well as condemnation for Telhan’s vision.
Telhan’s artwork is intended to critique the overarching assumption that lab-grown meat will eventually replace more traditional production methods, as well as the lack of transparency surrounding many processed foodstuffs. “They think that this problem (from industrial-scale agriculture) is going be solved by this new technology,” Telhan said. “I am critical (of) that perspective.”
Unlike Bramble, Telhan is not against lab-grown meat, so long as its producers are transparent about the sourcing of materials and its cultivation. But he believes that large-scale agricultural meat production – which dates back centuries – is not going to be replaced so quickly.
“We see this again and again with different industries, like algae-based fuels. A lot of companies were excited about this, and promoted it,” Telhan said. “And years later, we know these fuels work. But to be able to displace the oil industry means building the infrastructure to scale takes billions of dollars, and nobody has the patience or money to do it.”
Alvaro concurred on this point, which he believes is already weakened because a large swath of consumers aren’t concerned about environmental degradation.
“They’re going to have to sell this big, but in order to convince people to do so, they have to convince them to eat this product instead of regular meat,” Alvaro said.
Hidden Tweaks?
Moreover, if lab-based meat does obtain a significant market share, Telhan suggested companies may do things to the product – such as to genetically modify it to become more profitable – and never notify consumers. That is a particular concern in the U.S., where regulations regarding such modifications are vastly more relaxed than in the European Union.
“I think that they have really good objectives, and they aspire to good objectives,” Telhan said. “But the system itself doesn't really allow for that much transparency.”
No matter what the future holds, sometime next year Carnegie Mellon is expected to hold a press conference announcing it has produced a cut of the world’s most expensive beef with the help of a modified piece of consumer electronics. It will likely take place at around the same time UPSIDE chicken will be available for purchase in supermarkets and restaurants, pending the USDA’s approvals.
Abbott, the Carnegie Mellon professor, suggested the future event will be both informative and celebratory.
“I think Carnegie Mellon would have someone potentially cook it for us,” she said. “Like have a really good chef in New York City do it.”

