New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
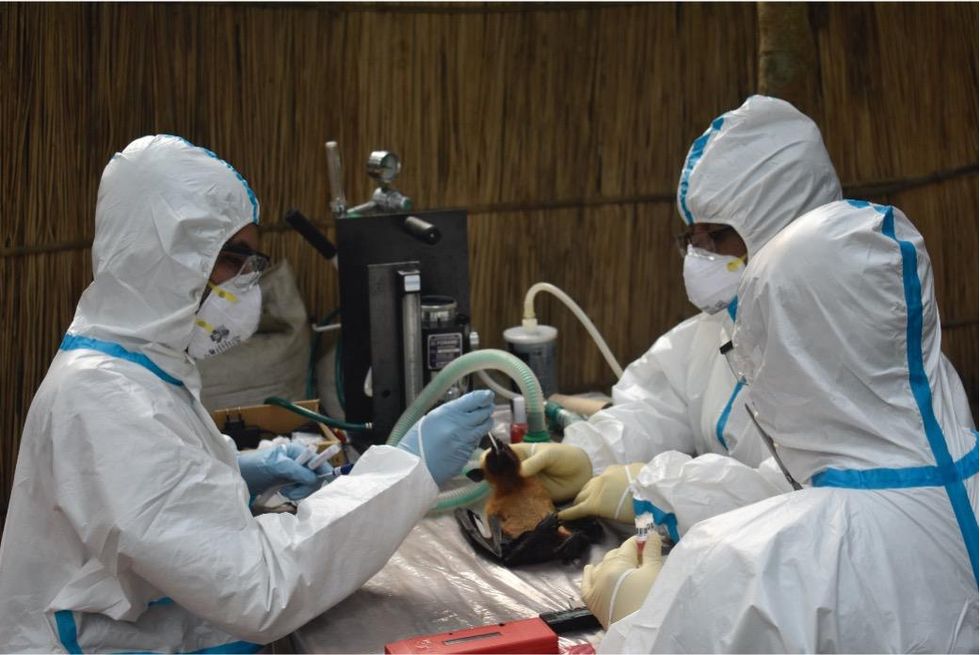
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.
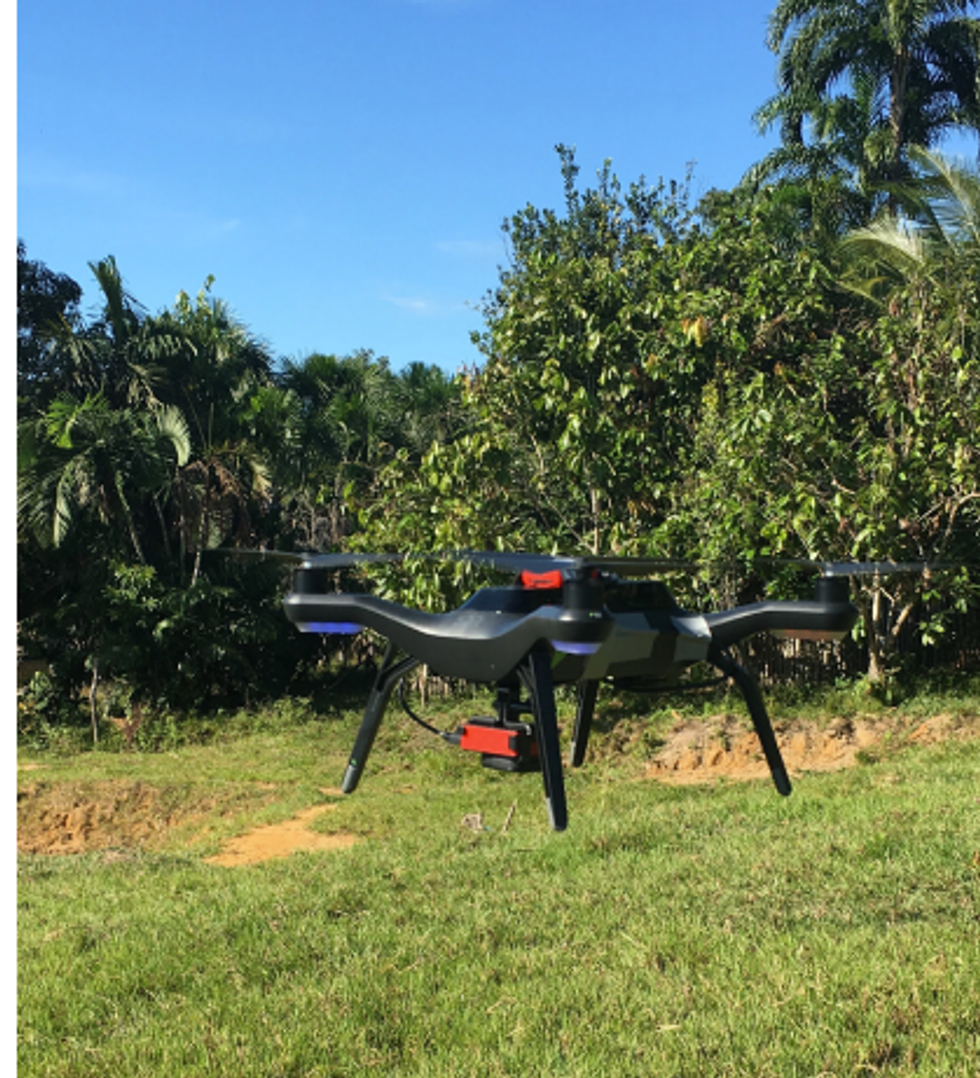

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Life is Emerging: Review of Siddhartha Mukherjee’s Song of the Cell
A new book by Pulitzer-winning physician-scientist Siddhartha Mukherjee will be released from Simon & Schuster on October 25, 2022.
The DNA double helix is often the image spiraling at the center of 21st century advances in biomedicine and the growing bioeconomy. And yet, DNA is molecularly inert. DNA, the code for genes, is not alive and is not strictly necessary for life. Ought life be at the center of our communication of living systems? Is not the Cell a superior symbol of life and our manipulation of living systems?
A code for life isn’t a code without the life that instantiates it. A code for life must be translated. The cell is the basic unit of that translation. The cell is the minimal viable package of life as we know it. Therefore, cell biology is at the center of biomedicine’s greatest transformations, suggests Pulitzer-winning physician-scientist Siddhartha Mukherjee in his latest book, The Song of the Cell: The Exploration of Medicine and the New Human.
The Song of the Cell begins with the discovery of cells and of germ theory, featuring characters such as Louis Pasteur and Robert Koch, who brought the cell “into intimate contact with pathology and medicine.” This intercourse would transform biomedicine, leading to the insight that we can treat disease by thinking at the cellular level. The slightest rearrangement of sick cells might be the path toward alleviating suffering for the organism: eroding the cell walls of a bacterium while sparing our human cells; inventing a medium that coaxes sperm and egg to dance into cellular union for in vitro fertilization (IVF); designing molecular missiles that home to the receptors decorating the exterior of cancer cells; teaching adult skin cells to remember their embryonic state for regenerative medicines.
Mukherjee uses the bulk of the book to elucidate key cell types in the human body, along with their “connective relationships” that enable key organs and organ systems to function. This includes the immune system, the heart, the brain, and so on. Mukherjee’s distinctive style features compelling anecdotes and human stories that animate the scientific (and unscientific) processes that have led to our current state of understanding. In his chapter on neurons and the brain, for example, he integrates Santiago Ramon y Cajal’s meticulous black ink sketches of neurons into Mukherjee’s own personal encounter with clinical depression. In one lucid section, he interviews Dr. Helen Mayberg, a pioneering neurologist who takes seriously the descriptive power of her patients’ metaphors, as they suffer from “caves,” “holes,” “voids,” and “force fields” that render their lives gray. Dr. Mayberg aims to stimulate patients’ neuronal cells in a manner that brings back the color.

Beyond exposing the insight and inventiveness that has arisen out of cell-based thinking, it seems that Mukherjee’s bigger project is an epistemological one. The early chapters of The Song of the Cell continually hint at the potential for redefining the basic unit of biology as the cell rather than the gene. The choice to center biomedicine around cells is, above all, a conspicuous choice not to center it around genes (the subject of Mukherjee’s previous book, The Gene), because genes dominate popular science communication.
This choice of cells over genes is most welcome. Cells are alive. Genes are not. Letters—such as the As, Cs, Gs, and Ts that represent the nucleotides of DNA, which make up our genes—must be synthesized into a word or poem or song that offers a glimpse into deeper truths. A key idea embedded in this thinking is that of emergence. Whether in ancient myth or modern art, creation tends to be an emergent process, not a linearly coded script. The cell is our current best guess for the basic unit of life’s emergence, turning a finite set of chemical building blocks—nucleic acids, proteins, sugars, fats—into a replicative, evolving system for fighting stasis and entropy. The cell’s song is one for our times, for it is the song of biology’s emergence out of chemistry and physics, into the “frenetically active process” of homeostasis.
Re-centering our view of biology has practical consequences, too, for how we think about diagnosing and treating disease, and for inventing new medicines. Centering cells presents a challenge: which type of cell to place at the center? Rather than default to the apparent simplicity of DNA as a symbol because it represents the one master code for life, the tension in defining the diversity of cells—a mapping process still far from complete in cutting-edge biology laboratories—can help to create a more thoughtful library of cellular metaphors to shape both the practice and communication of biology.
Further, effective problem solving is often about operating at the right level, or the right scale. The cell feels like appropriate level at which to interrogate many of the diseases that ail us, because the senses that guide our own perceptions of sickness and health—the smoldering pain of inflammation, the tunnel vision of a migraine, the dizziness of a fluttering heart—are emergent.
This, unfortunately, is sort of where Mukherjee leaves the reader, under-exploring the consequences of a biology of emergence. Many practical and profound questions have to do with the ways that each scale of life feeds back on the others. In a tome on Cells and “the future human” I wished that Mukherjee had created more space for seeking the ways that cells will shape and be shaped by the future, of humanity and otherwise.
We are entering a phase of real-world bioengineering that features the modularization of cellular parts within cells, of cells within organs, of organs within bodies, and of bodies within ecosystems. In this reality, we would be unwise to assume that any whole is the mere sum of its parts.
For example, when discussing the regenerative power of pluripotent stem cells, Mukherjee raises the philosophical thought experiment of the Delphic boat, also known as the Ship of Theseus. The boat is made of many pieces of wood, each of which is replaced for repairs over the years, with the boat’s structure unchanged. Eventually none of the boat’s original wood remains: Is it the same boat?
Mukherjee raises the Delphic boat in one paragraph at the end of the chapter on stem cells, as a metaphor related to the possibility of stem cell-enabled regeneration in perpetuity. He does not follow any of the threads of potential answers. Given the current state of cellular engineering, about which Mukherjee is a world expert from his work as a physician-scientist, this book could have used an entire section dedicated to probing this question and, importantly, the ways this thought experiment falls apart.
We are entering a phase of real-world bioengineering that features the modularization of cellular parts within cells, of cells within organs, of organs within bodies, and of bodies within ecosystems. In this reality, we would be unwise to assume that any whole is the mere sum of its parts. Wholeness at any one of these scales of life—organelle, cell, organ, body, ecosystem—is what is at stake if we allow biological reductionism to assume away the relation between those scales.
In other words, Mukherjee succeeds in providing a masterful and compelling narrative of the lives of many of the cells that emerge to enliven us. Like his previous books, it is a worthwhile read for anyone curious about the role of cells in disease and in health. And yet, he fails to offer the broader context of The Song of the Cell.
As leading agronomist and essayist Wes Jackson has written, “The sequence of amino acids that is at home in the human cell, when produced inside the bacterial cell, does not fold quite right. Something about the E. coli internal environment affects the tertiary structure of the protein and makes it inactive. The whole in this case, the E. coli cell, affects the part—the newly made protein. Where is the priority of part now?” [1]
Beyond the ways that different kingdoms of life translate the same genetic code, the practical situation for humanity today relates to the ways that the different disciplines of modern life use values and culture to influence our genes, cells, bodies, and environment. It may be that humans will soon become a bit like the Delphic boat, infused with the buzz of fresh cells to repopulate different niches within our bodies, for healthier, longer lives. But in biology, as in writing, a mixed metaphor can cause something of a cacophony. For we are not boats with parts to be replaced piecemeal. And nor are whales, nor alpine forests, nor topsoil. Life isn’t a sum of parts, and neither is a song that rings true.
[1] Wes Jackson, "Visions and Assumptions," in Nature as Measure (p. 52-53).
You won't score many glamor points for using a neti pot, but it could be a worthwhile asset in fighting Covid-19, according to the author's experience and recent research.
Twice a day, morning and night, I use a neti pot to send a warm saltwater solution coursing through one nostril and out the other to flush out debris and pathogens. I started many years ago because of sinus congestion and infections and it has greatly reduced those problems. Along with vaccination when it became available, it seems to have helped with protecting me from developing Covid-19 symptoms despite being of an age and weight that puts me squarely at risk.
Now that supposition of protection has been backed up with evidence from a solidly designed randomized clinical trial. It found that irrigating your sinuses twice a day with a simple saltwater solution can lead to an 8.5-fold reduction in hospitalization from Covid-19. The study is another example of recent research that points to easy and inexpensive ways to help protect yourself and help control the epidemic.
Amy Baxter, the physician researcher behind the study at Augusta University, Medical College of Georgia, began the study in 2020, before a vaccine or monoclonal antibodies became available to counter the virus. She wanted to be able to offer another line of defense for people with limited access to healthcare.
The nasal cavity is the front door that the SARS-CoV-2 virus typically uses to enter the body, latching on to the ACE2 receptors on cells lining those tissue compartments to establish infection. Once the virus replicates here, infection spreads into the lungs and often other parts of the body, including the brain and gut. Some studies have shown that a mouthwash could reduce the viral load, but any effect on disease progression was less clear. Baxter reasoned that reducing the amount of virus in the nose might give the immune system a better chance to react and control that growth before it got out of hand.
She decided to test this approach in patients who had just tested positive for Covid-19, were over 55 years of age, and often had other risk factors for developing serious symptoms. It was the quickest and easiest way to get results. A traditional prevention study would have required many more volunteers, taken a longer period of follow up, and cost money she did not have.
The trial enrolled 79 participants within 24 hours of testing positive for Covid-19, and they agreed to follow the regimen of twice daily nasal irrigation. They were followed for 28 days. One patient was hospitalized; a 1.27% rate compared with 11% in a national sample control group of similar age people who tested positive for Covid-19. Patients who strictly adhered to nasal irrigation had fewer, shorter and less severe symptoms than people in the study who missed some of their saline rinses.
Baxter initially made the results of her clinical trial available as a preprint in the summer of 2021 and was dismayed when many of the comments were from anti-vaxxers who argued this was a reason why you did not need to get vaccinated. That was not her intent.
There are several mechanisms that explain why warm saltwater is so effective. First and most obvious is the physical force of the water that sweeps away debris just as a rainstorm sends trash into a street gutter and down a storm drain. It also lubricates the cilia, small hair-like structures whose job it is to move detritus away from cells for expulsion. Cilia are rich in ACE2 receptors and keeping them moving makes it harder for the virus to latch on to the receptors.
It turns out the saline has a direct effect on the virus itself. SARS-CoV-2 becomes activated when an enzyme called furin snips off part of its molecular structure, which allows the virus to grab on to the ACE2 receptor, but saline inhibits this process. Once inside a cell the virus replicates best in a low salt environment, but nasal cells absorb salt from the irrigation, which further slows viral replication, says Baxter.
Finally, “salt improves the jellification of liquid, it makes better and stickier mucus so that you can get those virus out,” she explains, lamenting, “Nobody cares about snot. I do now.”
She initially made the results of her clinical trial available as a preprint in the summer of 2021 and was dismayed when many of the comments were from anti-vaxxers who argued this was a reason why you did not need to get vaccinated. That was not her intent. Two journals rejected the paper, and Baxter believes getting caught up in the polarizing politics of Covid-19 was an important part of the reason why. She says that editors “didn't want to be associated with something that was being used by anti-vaxxers.” She strongly supports vaccination but realizes that additional and alternative approaches also are needed.
Premeasured packets of saline are inexpensive and can be purchased at any drug store. They are safe to use several times a day. Say you’re vaccinated but were in a situation where you fear you might have been exposed to SARS-CoV-2; an extra irrigation will clear out your sinuses and may reduce the risk of that possible exposure.
Baxter plans no further study in this area. She is returning to her primary research focus, which is pain control and reducing opioid use, but she hopes that others will expand on what she had done.