New tools could catch disease outbreaks earlier - or predict them

A team at the Catalan Institution for Research and Advanced Studies is utilizing drones and weather stations to collect data on how mosquito breeding patterns are changing in response to climate shifts.
Every year, the villages which lie in the so-called ‘Nipah belt’— which stretches along the western border between Bangladesh and India, brace themselves for the latest outbreak. For since 1998, when Nipah virus—a form of hemorrhagic fever most common in Bangladesh—first spilled over into humans, it has been a grim annual visitor to the people of this region.
With a 70 percent fatality rate, no vaccine, and no known treatments, Nipah virus has been dubbed in the Western world as ‘the worst disease no one has ever heard of.’ Currently, outbreaks tend to be relatively contained because it is not very transmissible. The virus circulates throughout Asia in fruit eating bats, and only tends to be passed on to people who consume contaminated date palm sap, a sweet drink which is harvested across Bangladesh.
But as SARS-CoV-2 has shown the world, this can quickly change.
“Nipah virus is among what virologists call ‘the Big 10,’ along with things like Lassa fever and Crimean Congo hemorrhagic fever,” says Noam Ross, a disease ecologist at New York-based non-profit EcoHealth Alliance. “These are pretty dangerous viruses from a lethality perspective, which don’t currently have the capacity to spread into broader human populations. But that can evolve, and you could very well see a variant emerge that has human-human transmission capability.”
That’s not an overstatement. Surveys suggest that mammals harbour about 40,000 viruses, with roughly a quarter capable of infecting humans. The vast majority never get a chance to do so because we don’t encounter them, but climate change can alter that. Recent studies have found that as animals relocate to new habitats due to shifting environmental conditions, the coming decades will bring around 300,000 first encounters between species which normally don’t interact, especially in tropical Africa and southeast Asia. All these interactions will make it far more likely for hitherto unknown viruses to cross paths with humans.
That’s why for the last 16 years, EcoHealth Alliance has been conducting ongoing viral surveillance projects across Bangladesh. The goal is to understand why Nipah is so much more prevalent in the western part of the country, compared to the east, and keep a watchful eye out for new Nipah strains as well as other dangerous pathogens like Ebola.
"There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them," says Cat Lippi, medical geography researcher at the University of Florida.
Until very recently this kind of work has been hampered by the limitations of viral surveillance technology. The PREDICT project, a $200 million initiative funded by the United States Agency for International Development, which conducted surveillance across the Amazon Basin, Congo Basin and extensive parts of South and Southeast Asia, relied upon so-called nucleic acid assays which enabled scientists to search for the genetic material of viruses in animal samples.
However, the project came under criticism for being highly inefficient. “That approach requires a big sampling effort, because of the rarity of individual infections,” says Ross. “Any particular animal may be infected for a couple of weeks, maybe once or twice in its lifetime. So if you sample thousands and thousands of animals, you'll eventually get one that has an Ebola virus infection right now.”
Ross explains that there is now far more interest in serological sampling—the scientific term for the process of drawing blood for antibody testing. By searching for the presence of antibodies in the blood of humans and animals, scientists have a greater chance of detecting viruses which started circulating recently.
Despite the controversy surrounding EcoHealth Alliance’s involvement in so-called gain of function research—experiments that study whether viruses might mutate into deadlier strains—the organization’s separate efforts to stay one step ahead of pathogen evolution are key to stopping the next pandemic.
“Having really cheap and fast surveillance is really important,” says Ross. “Particularly in a place where there's persistent, low level, moderate infections that potentially have the ability to develop into more epidemic or pandemic situations. It means there’s a pathway that something more dangerous can come through."
Scientists are searching for the presence of antibodies in the blood of humans and animals in hopes to detect viruses that recently started circulating.
EcoHealth Alliance
In Bangladesh, EcoHealth Alliance is attempting to do this using a newer serological technology known as a multiplex Luminex assay, which tests samples against a panel of known antibodies against many different viruses. It collects what Ross describes as a ‘footprint of information,’ which allows scientists to tell whether the sample contains the presence of a known pathogen or something completely different and needs to be investigated further.
By using this technology to sample human and animal populations across the country, they hope to gain an idea of whether there are any novel Nipah virus variants or strains from the same family, as well as other deadly viral families like Ebola.
This is just one of several novel tools being used for viral discovery in surveillance projects around the globe. Multiple research groups are taking PREDICT’s approach of looking for novel viruses in animals in various hotspots. They collect environmental DNA—mucus, faeces or shed skin left behind in soil, sediment or water—which can then be genetically sequenced.
Five years ago, this would have been a painstaking work requiring bringing collected samples back to labs. Today, thanks to the vast amounts of money spent on new technologies during COVID-19, researchers now have portable sequencing tools they can take out into the field.
Christopher Jerde, a researcher at the UC Santa Barbara Marine Science Institute, points to the Oxford Nanopore MinION sequencer as one example. “I tried one of the early versions of it four years ago, and it was miserable,” he says. “But they’ve really improved, and what we’re going to be able to do in the next five to ten years will be amazing. Instead of having to carefully transport samples back to the lab, we're going to have cigar box-shaped sequencers that we take into the field, plug into a laptop, and do the whole sequencing of an organism.”
In the past, viral surveillance has had to be very targeted and focused on known families of viruses, potentially missing new, previously unknown zoonotic pathogens. Jerde says that the rise of portable sequencers will lead to what he describes as “true surveillance.”
“Before, this was just too complex,” he says. “It had to be very focused, for example, looking for SARS-type viruses. Now we’re able to say, ‘Tell us all the viruses that are here?’ And this will give us true surveillance – we’ll be able to see the diversity of all the pathogens which are in these spots and have an understanding of which ones are coming into the population and causing damage.”
But being able to discover more viruses also comes with certain challenges. Some scientists fear that the speed of viral discovery will soon outpace the human capacity to analyze them all and assess the threat that they pose to us.
“I think we're already there,” says Jason Ladner, assistant professor at Northern Arizona University’s Pathogen and Microbiome Institute. “If you look at all the papers on the expanding RNA virus sphere, there are all of these deposited partial or complete viral sequences in groups that we just don't know anything really about yet.” Bats, for example, carry a myriad of viruses, whose ability to infect human cells we understand very poorly.
Cultivating these viruses under laboratory conditions and testing them on organoids— miniature, simplified versions of organs created from stem cells—can help with these assessments, but it is a slow and painstaking work. One hope is that in the future, machine learning could help automate this process. The new SpillOver Viral Risk Ranking platform aims to assess the risk level of a given virus based on 31 different metrics, while other computer models have tried to do the same based on the similarity of a virus’s genomic sequence to known zoonotic threats.
However, Ladner says that these types of comparisons are still overly simplistic. For one thing, scientists are still only aware of a few hundred zoonotic viruses, which is a very limited data sample for accurately assessing a novel pathogen. Instead, he says that there is a need for virologists to develop models which can determine viral compatibility with human cells, based on genomic data.
“One thing which is really useful, but can be challenging to do, is understand the cell surface receptors that a given virus might use,” he says. “Understanding whether a virus is likely to be able to use proteins on the surface of human cells to gain entry can be very informative.”
As the Earth’s climate heats up, scientists also need to better model the so-called vector borne diseases such as dengue, Zika, chikungunya and yellow fever. Transmitted by the Aedes mosquito residing in humid climates, these blights currently disproportionally affect people in low-income nations. But predictions suggest that as the planet warms and the pests find new homes, an estimated one billion people who currently don’t encounter them might be threatened by their bites by 2080. “When it comes to mosquito-borne diseases we have to worry about shifts in suitable habitat,” says Cat Lippi, a medical geography researcher at the University of Florida. “As climate patterns change on these big scales, we expect to see shifts in where people will be at risk for contracting these diseases.”
Public health practitioners and government decision-makers need tools to make climate-informed decisions about the evolving threat of different infectious diseases. Some projects are already underway. An ongoing collaboration between the Catalan Institution for Research and Advanced Studies and researchers in Brazil and Peru is utilizing drones and weather stations to collect data on how mosquitoes change their breeding patterns in response to climate shifts. This information will then be fed into computer algorithms to predict the impact of mosquito-borne illnesses on different regions.

The team at the Catalan Institution for Research and Advanced Studies is using drones and weather stations to collect data on how mosquito breeding patterns change due to climate shifts.
Gabriel Carrasco
Lippi says that similar models are urgently needed to predict how changing climate patterns affect respiratory, foodborne, waterborne and soilborne illnesses. The UK-based Wellcome Trust has allocated significant assets to fund such projects, which should allow scientists to monitor the impact of climate on a much broader range of infections. “There are a lot of different infectious agents that are sensitive to climate change that don't have these sorts of software tools being developed for them,” she says.
COVID-19’s havoc boosted funding for infectious disease research, but as its threats begin to fade from policymakers’ focus, the money may dry up. Meanwhile, scientists warn that another major infectious disease outbreak is inevitable, potentially within the next decade, so combing the planet for pathogens is vital. “Surveillance is ultimately a really boring thing that a lot of people don't want to put money into, until we have a wide scale pandemic,” Jerde says, but that vigilance is key to thwarting the next deadly horror. “It takes a lot of patience and perseverance to keep looking.”
This article originally appeared in One Health/One Planet, a single-issue magazine that explores how climate change and other environmental shifts are increasing vulnerabilities to infectious diseases by land and by sea. The magazine probes how scientists are making progress with leaders in other fields toward solutions that embrace diverse perspectives and the interconnectedness of all lifeforms and the planet.
Hyperbaric oxygen therapy could treat Long COVID, new study shows
Hyperbaric oxygen therapy has been used in the past to help people with traumatic brain injury, stroke and other conditions involving wounds to the brain. Now, researchers at Shamir Medical Center in Tel Aviv are studying how it could treat Long Covid.
Long COVID is not a single disease, it is a syndrome or cluster of symptoms that can arise from exposure to SARS-CoV-2, a virus that affects an unusually large number of different tissue types. That's because the ACE2 receptor it uses to enter cells is common throughout the body, and inflammation from the immune response fighting that infection can damage surrounding tissue.
One of the most widely shared groups of symptoms is fatigue and what has come to be called “brain fog,” a difficulty focusing and an amorphous feeling of slowed mental functioning and capacity. Researchers have tied these COVID-related symptoms to tissue damage in specific sections of the brain and actual shrinkage in its size.
When Shai Efrati, medical director of the Sagol Center for Hyperbaric Medicine and Research in Tel Aviv, first looked at functional magnetic resonance images (fMRIs) of patients with what is now called long COVID, he saw “micro infarcts along the brain.” It reminded him of similar lesions in other conditions he had treated with hyperbaric oxygen therapy (HBOT). “Once we saw that, we said, this is the type of wound we can treat. It doesn't matter if the primary cause is mechanical injury like TBI [traumatic brain injury] or stroke … we know how to oxidize them.”Efrati came to HBOT almost by accident. The physician had seen how it had helped heal diabetic ulcers and improved the lives of other patients, but he was busy with his own research. Then the director of his Tel Aviv hospital threatened to shut down the small HBOT chamber unless Efrati took on administrative responsibility for it. He reluctantly agreed, a decision that shifted the entire focus of his research.
“The main difference between wounds in the leg and wounds in the brain is that one is something we can see, it's tangible, and the wound in the brain is hidden,” says Efrati. With fMRIs, he can measure how a limited supply of oxygen in blood is shuttled around to fuel activity in various parts of the brain. Years of research have mapped how specific areas of the brain control activity ranging from thinking to moving. An fMRI captures the brain area as it’s activated by supplies of oxygen; lack of activity after the same stimuli suggests damage has occurred in that tissue. Suddenly, what was hidden became visible to researchers using fMRI. It helped to make a diagnosis and measure response to treatment.
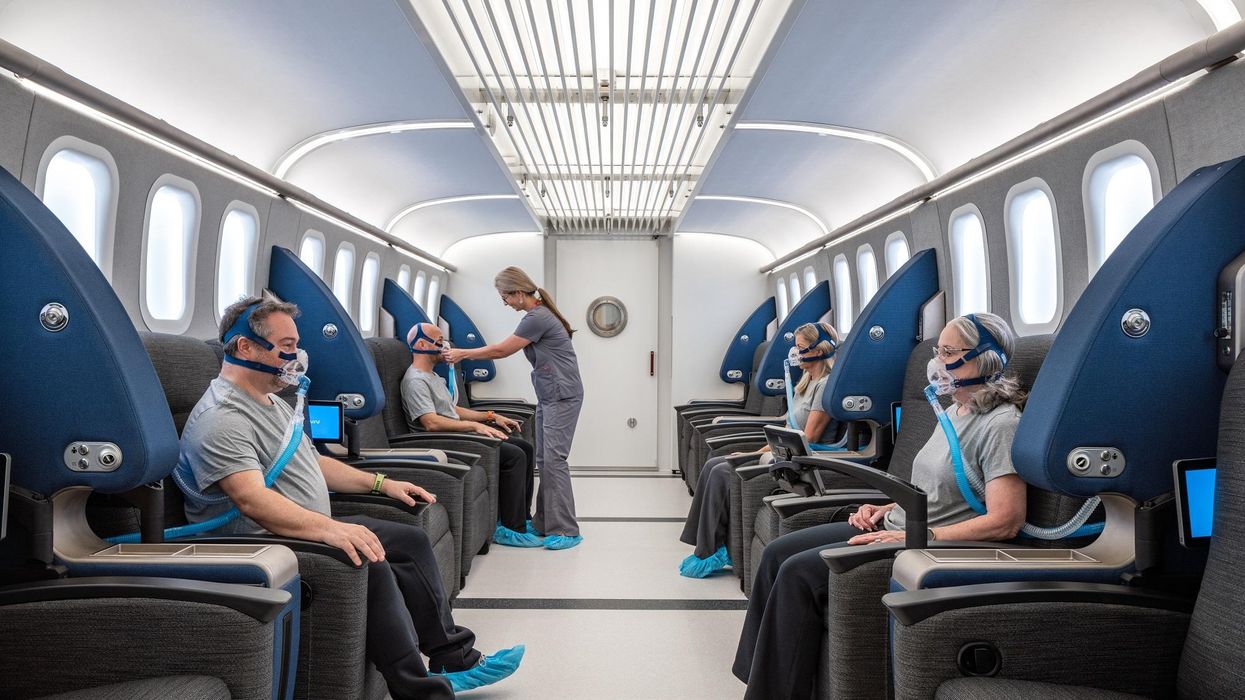
HBOT is not a single thing but rather a tool, a process or approach with variations depending on the condition being treated. It aims to increase the amount of oxygen that gets to damaged tissue and speed up healing. Regular air is about 21 percent oxygen. But inside the HBOT chamber the atmospheric pressure can be increased to up to three times normal pressure at sea level and the patient breathes pure oxygen through a mask; blood becomes saturated with much higher levels of oxygen. This can defuse through the damaged capillaries of a wound and promote healing.
The trial
Efrati’s clinical trials started in December 2020, barely a year after SARS-CoV-2 had first appeared in Israel. Patients who’d experienced cognitive issues after having COVID received 40 sessions in the chamber over a period of 60 days. In each session, they spent 90 minutes breathing through a mask at two atmospheres of pressure. While inside, they performed mental exercises to train the brain. The only difference between the two groups of patients was that one breathed pure oxygen while the other group breathed normal air. No one knew who was receiving which level of oxygen.
The results were striking. Before and after fMRIs showed significant repair of damaged tissue in the brain and functional cognition tests improved substantially among those who received pure oxygen. Importantly, 80 percent of patients said they felt back to “normal,” but Efrati says they didn't include patient evaluation in the paper because there was no baseline data to show how they functioned before COVID. After the study was completed, the placebo group was offered a new round of treatments using 100 percent oxygen, and the team saw similar results.
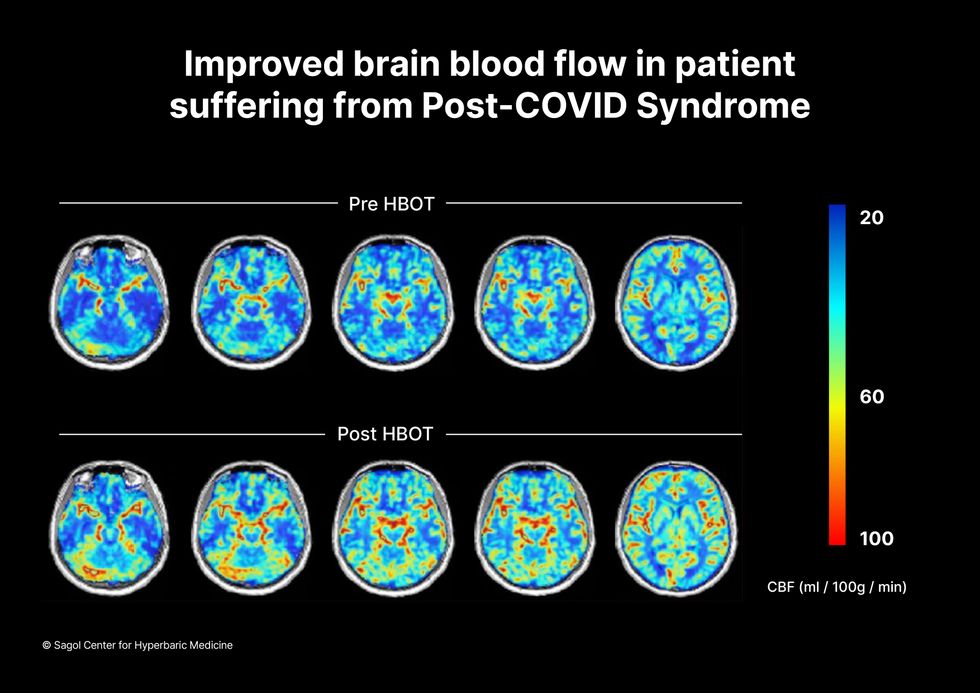
Scans show improved blood flow in a patient suffering from Long Covid.
Sagol Center for Hyperbaric Medicine
Efrati's use of HBOT is part of an emerging geroscience approach to diseases associated with aging. These researchers see systems dysfunctions that are common to several diseases, such as inflammation, which has been shown to play a role in micro infarcts, heart disease and Alzheimer’s disease. Preliminary research suggests that HBOT can retard some underlying mechanisms of aging, which might address several medical conditions. However, the drug approval process is set up to regulate individual disease, not conditions as broad as aging, and so they concentrate on treating the low hanging fruit: disorders where effective treatments currently are limited and success might be demonstrated.
The key to HBOT's effectiveness is something called the hyperoxic-hypoxic paradox where a body does not react to an increase in available oxygen, only to a decrease, regardless of the starting point. That danger signal has a powerful effect on gene expression, resulting in changes in metabolism, and the proliferation of stem cells. That occurs with each cycle of 20 minutes of pure oxygen followed by 5 minutes of regular air circulating through the masks, while the chamber remains pressurized. The high levels of oxygen in the blood provide the fuel necessary for tissue regeneration.
The hyperbaric chamber that Efrati has built can hold a dozen patients and attending medical staff. Think of it as a pressurized airplane cabin, only with much more space than even in first class. In the U.S., people think of HBOT as “a sack of air or some tube that you can buy on Amazon” or find at a health spa. “That is total bullshit,” Efrati says. “It has to be a medical class center where a physician can lose their license if they are not operating it properly.”
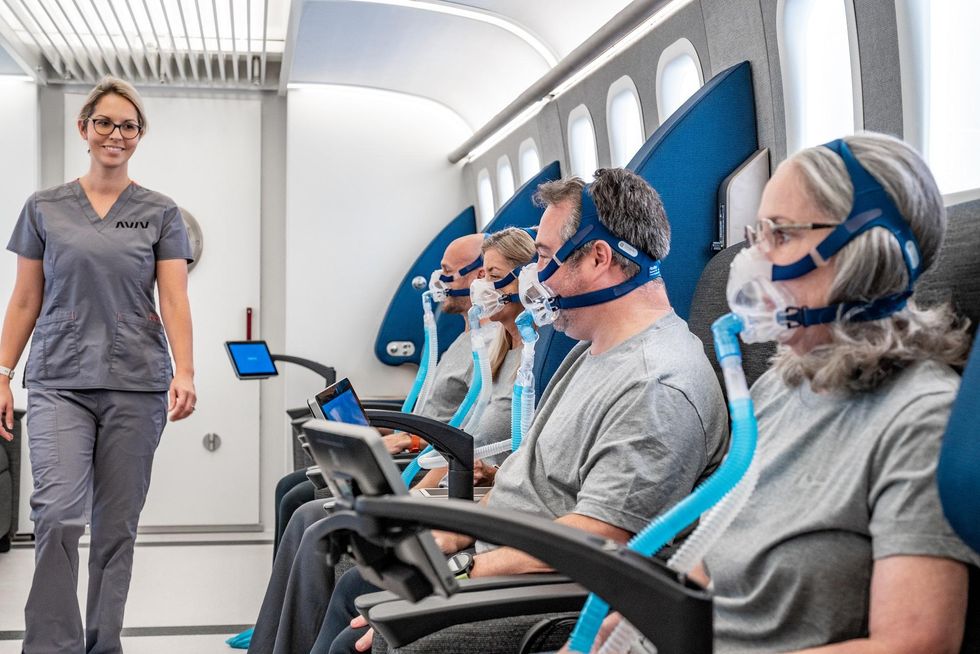

Shai Efrati
Alexander Charney, a research psychiatrist at the Icahn School of Medicine at Mount Sinai in New York City, calls Efrati’s study thoughtful and well designed. But it demands a lot from patients with its intense number of sessions. Those types of regimens have proven difficult to roll out to large numbers of patients. Still, the results are intriguing enough to merit additional trials.
John J. Miller, a physician and editor in chief of Psychiatric Times, has seen “many physicians that use hyperbaric oxygen for various brain disorders such as TBI.” He is intrigued by Efrati's work and believes the approach “has great potential to help patients with long COVID whose symptoms are related to brain tissue changes.”
Efrati believes so much in the power of the hyperoxic-hypoxic paradox to heal a variety of tissue injuries that he is leading the medical advisory board at Aviv Clinic, an international network of clinics that are delivering HBOT treatments based on research conducted in Israel. His goal is to silence doubters by quickly opening about 50 such clinics worldwide, based on the model of standalone dialysis clinics in the United States. Sagol Center is treating 300 patients per day, and clinics have opened in Florida and Dubai. There are plans to open another in Manhattan.
A blood test may catch colorectal cancer before it's too late
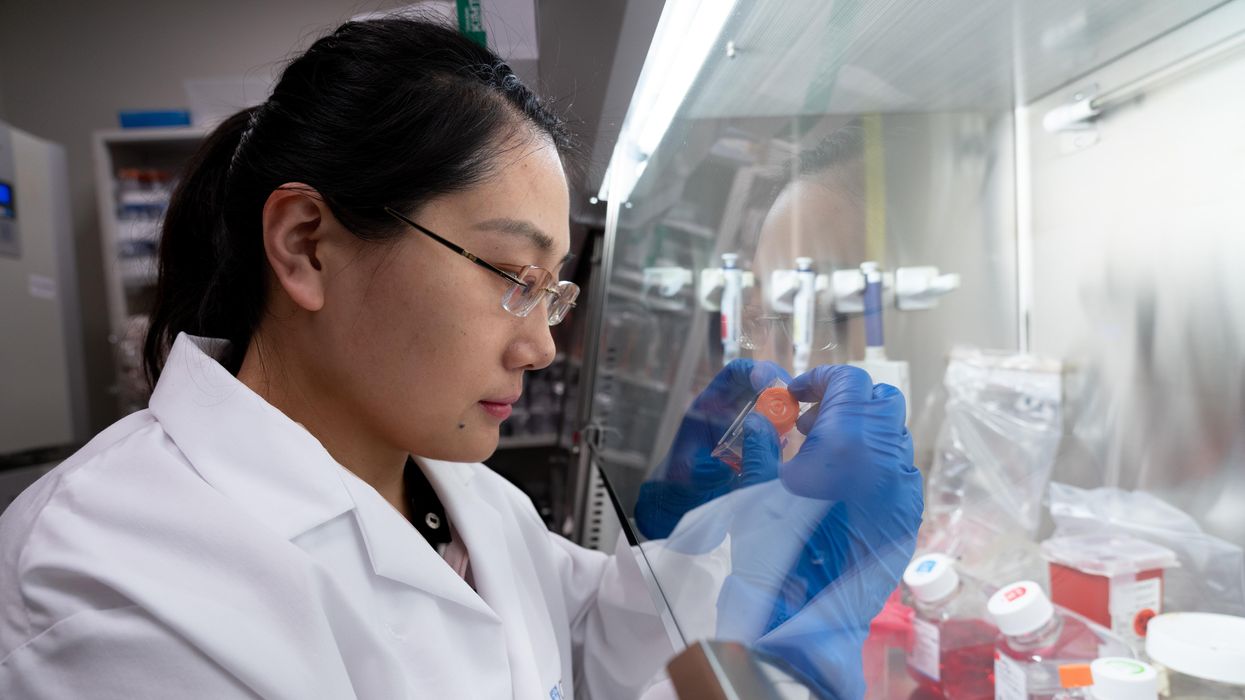
A scientist works on a blood test in the Ajay Goel Lab, one of many labs that are developing blood tests to screen for different types of cancer.
Soon it may be possible to find different types of cancer earlier than ever through a simple blood test.
Among the many blood tests in development, researchers announced in July that they have developed one that may screen for early-onset colorectal cancer. The new potential screening tool, detailed in a study in the journal Gastroenterology, represents a major step in noninvasively and inexpensively detecting nonhereditary colorectal cancer at an earlier and more treatable stage.
In recent years, this type of cancer has been on the upswing in adults under age 50 and in those without a family history. In 2021, the American Cancer Society's revised guidelines began recommending that colorectal cancer screenings with colonoscopy begin at age 45. But that still wouldn’t catch many early-onset cases among people in their 20s and 30s, says Ajay Goel, professor and chair of molecular diagnostics and experimental therapeutics at City of Hope, a Los Angeles-based nonprofit cancer research and treatment center that developed the new blood test.
“These people will mostly be missed because they will never be screened for it,” Goel says. Overall, colorectal cancer is the fourth most common malignancy, according to the U.S. Centers for Disease Control and Prevention.
Goel is far from the only one working on this. Dozens of companies are in the process of developing blood tests to screen for different types of malignancies.
Some estimates indicate that between one-fourth and one-third of all newly diagnosed colorectal cancers are early-onset. These patients generally present with more aggressive and advanced disease at diagnosis compared to late-onset colorectal cancer detected in people 50 years or older.
To develop his test, Goel examined publicly available datasets and figured out that changes in novel microRNAs, or miRNAs, which regulate the expression of genes, occurred in people with early-onset colorectal cancer. He confirmed these biomarkers by looking for them in the blood of 149 patients who had the early-onset form of the disease. In particular, Goel and his team of researchers were able to pick out four miRNAs that serve as a telltale sign of this cancer when they’re found in combination with each other.
The blood test is being validated by following another group of patients with early-onset colorectal cancer. “We have filed for intellectual property on this invention and are currently seeking biotech/pharma partners to license and commercialize this invention,” Goel says.
He’s far from the only one working on this. Dozens of companies are in the process of developing blood tests to screen for different types of malignancies, says Timothy Rebbeck, a professor of cancer prevention at the Harvard T.H. Chan School of Public Health and the Dana-Farber Cancer Institute. But, he adds, “It’s still very early, and the technology still needs a lot of work before it will revolutionize early detection.”
The accuracy of the early detection blood tests for cancer isn’t yet where researchers would like it to be. To use these tests widely in people without cancer, a very high degree of precision is needed, says David VanderWeele, interim director of the OncoSET Molecular Tumor Board at Northwestern University’s Lurie Cancer Center in Chicago.
Otherwise, “you’re going to cause a lot of anxiety unnecessarily if people have false-positive tests,” VanderWeele says. So far, “these tests are better at finding cancer when there’s a higher burden of cancer present,” although the goal is to detect cancer at the earliest stages. Even so, “we are making progress,” he adds.
While early detection is known to improve outcomes, most cancers are detected too late, often after they metastasize and people develop symptoms. Only five cancer types have recommended standard screenings, none of which involve blood tests—breast, cervical, colorectal, lung (smokers considered at risk) and prostate cancers, says Trish Rowland, vice president of corporate communications at GRAIL, a biotechnology company in Menlo Park, Calif., which developed a multi-cancer early detection blood test.
These recommended screenings check for individual cancers rather than looking for any form of cancer someone may have. The devil lies in the fact that cancers without widespread screening recommendations represent the vast majority of cancer diagnoses and most cancer deaths.
GRAIL’s Galleri multi-cancer early detection test is designed to find more cancers at earlier stages by analyzing DNA shed into the bloodstream by cells—with as few false positives as possible, she says. The test is currently available by prescription only for those with an elevated risk of cancer. Consumers can request it from their healthcare or telemedicine provider. “Galleri can detect a shared cancer signal across more than 50 types of cancers through a simple blood draw,” Rowland says, adding that it can be integrated into annual health checks and routine blood work.
Cancer patients—even those with early and curable disease—often have tumor cells circulating in their blood. “These tumor cells act as a biomarker and can be used for cancer detection and diagnosis,” says Andrew Wang, a radiation oncologist and professor at the University of Texas Southwestern Medical Center in Dallas. “Our research goal is to be able to detect these tumor cells to help with cancer management.” Collaborating with Seungpyo Hong, the Milton J. Henrichs Chair and Professor at the University of Wisconsin-Madison School of Pharmacy, “we have developed a highly sensitive assay to capture these circulating tumor cells.”
Even if the quality of a blood test is superior, finding cancer early doesn’t always mean it’s absolutely best to treat it. For example, prostate cancer treatment’s potential side effects—the inability to control urine or have sex—may be worse than living with a slow-growing tumor that is unlikely to be fatal. “[The test] needs to tell me, am I going to die of that cancer? And, if I intervene, will I live longer?” says John Marshall, chief of hematology and oncology at Medstar Georgetown University Hospital in Washington, D.C.

Ajay Goel Lab
A blood test developed at the University of Texas MD Anderson Cancer Center in Houston helps predict who may benefit from lung cancer screening when it is combined with a risk model based on an individual’s smoking history, according to a study published in January in the Journal of Clinical Oncology. The personalized lung cancer risk assessment was more sensitive and specific than the 2021 and 2013 U.S. Preventive Services Task Force criteria.
The study involved participants from the Prostate, Lung, Colorectal, and Ovarian Cancer Screening Trial with a minimum of a 10 pack-year smoking history, meaning they smoked 20 cigarettes per day for ten years. If implemented, the blood test plus model would have found 9.2 percent more lung cancer cases for screening and decreased referral to screening among non-cases by 13.7 percent compared to the 2021 task force criteria, according to Oncology Times.
The conventional type of screening for lung cancer is an annual low-dose CT scan, but only a small percentage of people who are eligible will actually get these scans, says Sam Hanash, professor of clinical cancer prevention and director of MD Anderson’s Center for Global Cancer Early Detection. Such screening is not readily available in most countries.
In methodically searching for blood-based biomarkers for lung cancer screening, MD Anderson researchers developed a simple test consisting of four proteins. These proteins circulating in the blood were at high levels in individuals who had lung cancer or later developed it, Hanash says.
“The interest in blood tests for cancer early detection has skyrocketed in the past few years,” he notes, “due in part to advances in technology and a better understanding of cancer causation, cancer drivers and molecular changes that occur with cancer development.”
However, at the present time, none of the blood tests being considered eliminate the need for screening of eligible subjects using established methods, such as colonoscopy for colorectal cancer. Yet, Hanash says, “they have the potential to complement these modalities.”

