New drug for schizophrenia could meet desperate need for better treatments

The field of treating schizophrenia with drugs has been stuck in a long drought but, this month, a late-stage clinical trial found a new drug called KarXT could treat a range of symptoms.
Schizophrenia is a debilitating mental health condition that affects around 24 million people worldwide. Patients experience hallucinations and delusions when they develop schizophrenia, with experts referring to these new thoughts and behaviors as positive symptoms. They also suffer from negative symptoms in which they lose important functions, suffering from dulled emotions, lack of purpose and social withdrawal.
Currently available drugs can control only a portion of these symptoms but, on August 8th, Karuna Therapeutics announced its completion of a phase 3 clinical trial that found a new drug called KarXT could treat both positive and negative symptoms of schizophrenia. It could mean substantial progress against a problem that has stymied scientists for decades.
A long-standing problem
Since the 1950s, antipsychotics have been used to treat schizophrenia. People who suffer from it are thought to have too much of a brain chemical called dopamine, and antipsychotics work by blocking dopamine receptors in the brain. They can be effective in treating positive symptoms but have little impact on the negative ones, which can be devastating for a patient’s quality of life, making it difficult to maintain employment and have successful relationships. About 30 percent of schizophrenia patients don't actually respond to antipsychotics at all. Current drugs can also have adverse side effects including elevated cholesterol, high blood pressure, diabetes and movements that patients cannot control.
The recent clinical trial heralds a new treatment approach. “We believe it marks an important advancement for patients given its new and completely different mechanism of action from current therapies,” says Andrew Miller, COO of Karuna.
Scientists have been looking to develop alternatives. However, “the field of drug treatment of schizophrenia is currently in the doldrums,” says Peter McKenna, a senior researcher at FIDMAG Research Foundation in Spain which specialises in mental health.
In the 2000s there was a major push to target a brain receptor for a chemical called glutamate. Evidence suggested that this receptor is abnormal in the brains of schizophrenia patients, but attempts to try glutamate failed in clinical trials.
After that, many pharmaceutical companies dropped out of the race for a more useful treatment. But some companies continued to search, such as Karuna Therapeutics, led by founder and Chief Operating Officer Andrew Miller and CEO Steve Paul. The recent clinical trial suggests their persistence has led to an important breakthrough with their drug, KarXT. “We believe it marks an important advancement for patients given its new and completely different mechanism of action from current therapies,” Miller says.
How it works
Neurotransmitters are chemical messengers that pass signals between neurons. To work effectively, neurotransmitters need a receptor to bind to. A neurotransmitter called acetylcholine seems to be especially important in schizophrenia. It interacts with sites called muscarinic receptors, which are involved in the network of nerves that calm your body after a stressful event. Post mortem studies in people with schizophrenia have shown that two muscarinic receptors in the brain, the M1 and M4 receptors, are activated at unusually low levels because they don’t receive enough signals from acetylcholine.
The M4 receptor appears to play a role in psychosis. The M1 receptor is also associated with psychosis but is primarily thought to be involved in cognition. KarXT, taken orally, works by activating both of these receptors to signal properly. It is this twofold action that seems to explain its effectiveness. “[The drug’s] design enables the preferential stimulation of these muscarinic receptors in the brain,” Miller says.
How it developed
It all started in the early 1990s when Paul was at pharmaceutical company Eli Lilly. He discovered that Xanomeline, the drug they were testing on Alzheimer's patients, had antipsychotic effects. It worked by stimulating M1 and M4 receptors, so he and his colleagues decided to test Xanomeline on schizophrenia patients, supported by research on the connection between muscarinic receptors and psychosis. They found that Xanomeline reduced both positive and negative symptoms.
Unfortunately, it also caused significant side effects. The problem was that stimulating the M1 and M4 receptors in the brain also stimulated muscarinic receptors in the body that led to severe vomiting, diarrhea and even the temporary loss of consciousness.
In the end, Eli Lilly discontinued the clinical trials for the drug, but Miller set up Karuna Therapeutics to develop a solution. “I was determined to find a way to harness the therapeutic benefit demonstrated in studies of Xanomeline, while eliminating side effects that limited its development,” Miller says.
He analysed over 7,000 possible ways of mixing Xanomeline with other agents before settling on KarXT. It combines Xanomeline with a drug called Trospium Chloride, which blocks muscarinic receptors in the body – taking care of the side effects such as vomiting – but leaves them unblocked in the brain. Paul was so excited by Miller’s progress that he joined Karuna after leaving Eli Lilly and founding two previous startups.
“It's a very important approach,” says Rick Adams, Future Leaders Fellow in the Institute of Cognitive Neuroscience and Centre for Medical Image Computing at University College London. “We are in desperate need of alternative drug targets and this target is one of the best. There are other alternative targets, but not many are as close to being successful as the muscarinic receptor drug.”
Clinical Trial
Following a successful phase 2 clinical trial in 2019, the most recent trial involved 126 patients who were given KarXT, and 126 who were given a placebo. Compared to the placebo, patients taking KarXT had a significant 9.6 point reduction in the positive and negative syndrome scale (PANSS), the standard for rating schizophrenic symptoms.
KarXT also led to statistically significant declines in positive and negative symptoms compared to the placebo. “The results suggest that KarXT could be a potentially game-changing option in the management of both positive and negative symptoms of schizophrenia,” Miller says.
Robert McCutcheon, a psychiatrist and neuroscientist at Oxford University, is optimistic about the side effects but highlights the need for more safety trials.
McKenna, the researcher at FIDMAG Foundation, agrees about the drug’s potential. “The new [phase 3] study is positive,” he says. “It is reassuring that one is not dealing with a drug that works in one trial and then inexplicably fails in the next one.”
Robert McCutcheon, a psychiatrist and neuroscientist at Oxford University, said the drug is an unprecedented step forward. “KarXT is one of the first drugs with a novel mechanism of action to show promise in clinical trials.”
Even though the drug blocks muscarine receptors in the body, some patients still suffered from adverse side effects like vomiting, dizziness and diarrhea. But in general, these effects were mild to moderate, especially compared to dopamine-blocking antipsychotics or Xanomeline on its own.
McCutcheon is optimistic about the side effects but highlights the need for more safety trials. “The trial results suggest that gastrointestinal side effects appear to be manageable,” he says. “We know, however, from previous antipsychotic drugs that the full picture regarding the extent of side effects can sometimes take longer to become apparent to clinicians and patients. Careful ongoing assessment during a longer period of treatment will therefore be important.”
The Future
The team is currently conducting three other trials to evaluate the efficacy and long-term safety of KarXT. Their goal is to receive FDA approval next year.
Karuna is also conducting trials to evaluate the effectiveness of KarXT in treating psychosis in patients suffering from Alzheimer’s.
The big hope is that they will soon be able to provide a radically different drug to help many patients with schizophrenia. “We are another step closer to potentially providing the first new class of medicine in more than 50 years to the millions of people worldwide living with schizophrenia,” says Miller.
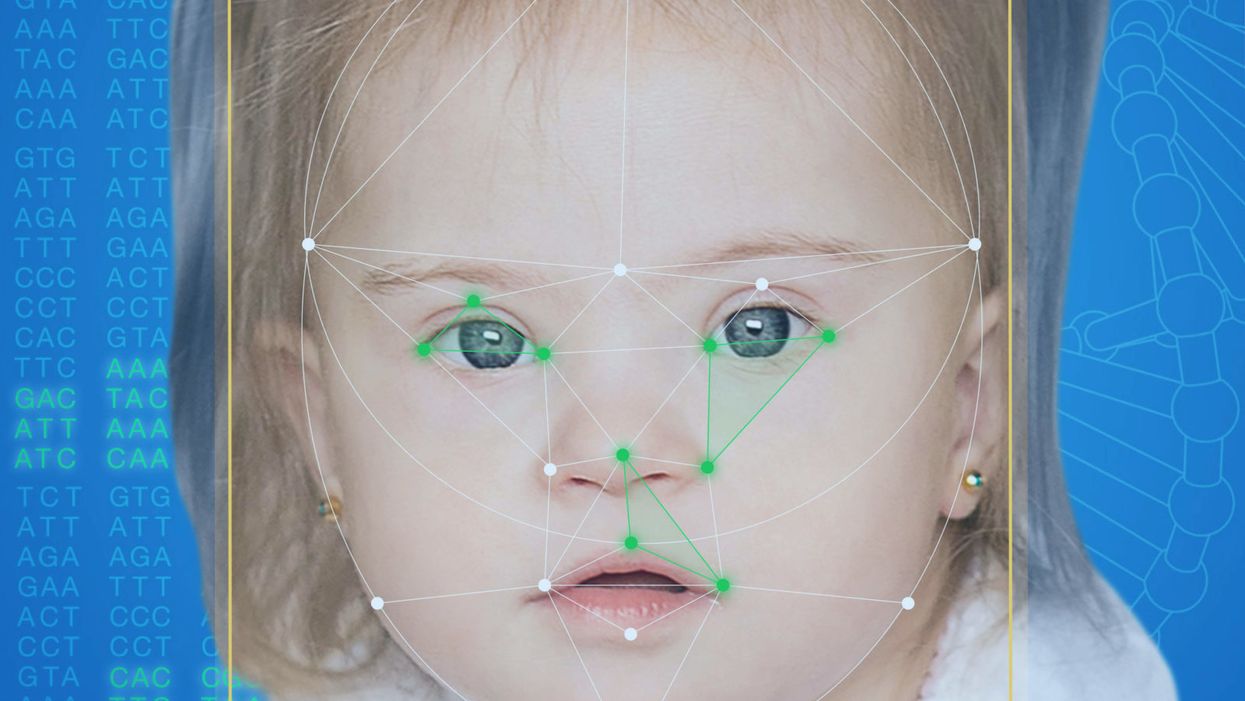
This App Helps Diagnose Rare Genetic Disorders from a Picture
FDNA's Face2Gene technology analyzes patient biometric data using artificial intelligence, identifying correlations with disease-causing genetic variations.
Medical geneticist Omar Abdul-Rahman had a hunch. He thought that the three-year-old boy with deep-set eyes, a rounded nose, and uplifted earlobes might have Mowat-Wilson syndrome, but he'd never seen a patient with the rare disorder before.
"If it weren't for the app I'm not sure I would have had the confidence to say 'yes you should spend $1000 on this test."
Rahman had already ordered genetic tests for three different conditions without any luck, and he didn't want to cost the family any more money—or hope—if he wasn't sure of the diagnosis. So he took a picture of the boy and uploaded the photo to Face2Gene, a diagnostic aid for rare genetic disorders. Sure enough, Mowat-Wilson came up as a potential match. The family agreed to one final genetic test, which was positive for the syndrome.
"If it weren't for the app I'm not sure I would have had the confidence to say 'yes you should spend $1000 on this test,'" says Rahman, who is now the director of Genetic Medicine at the University of Nebraska Medical Center, but saw the boy when he was in the Department of Pediatrics at the University of Mississippi Medical Center in 2012.
"Families who are dealing with undiagnosed diseases never know what's going to come around the corner, what other organ system might be a problem next week," Rahman says. With a diagnosis, "You don't have to wait for the other shoe to drop because now you know the extent of the condition."
A diagnosis is the first and most important step for patients to attain medical care. Disease prognosis, treatment plans, and emotional coping all stem from this critical phase. But diagnosis can also be the trickiest part of the process, particularly for rare disorders. According to one European survey, 40 percent of rare diseases are initially misdiagnosed.
Healthcare professionals and medical technology companies hope that facial recognition software will help prevent families from facing difficult disruptions due to misdiagnoses.
"Patients with rare diseases or genetic disorders go through a long period of diagnostic odyssey, and just putting a name to a syndrome or finding a diagnosis can be very helpful and relieve a lot of tension for the family," says Dekel Gelbman, CEO of FDNA.
Consequently, a misdiagnosis can be devastating for families. Money and time may have been wasted on fruitless treatments, while opportunities for potentially helpful therapies or clinical trials were missed. Parents led down the wrong path must change their expectations of their child's long-term prognosis and care. In addition, they may be misinformed regarding future decisions about family planning.
Healthcare professionals and medical technology companies hope that facial recognition software will help prevent families from facing these difficult disruptions by improving the accuracy and ease of diagnosing genetic disorders. Traditionally, doctors diagnose these types of conditions by identifying unique patterns of facial features, a practice called dysmorphology. Trained physicians can read a child's face like a map and detect any abnormal ridges or plateaus—wide-set eyes, broad forehead, flat nose, rotated ears—that, combined with other symptoms such as intellectual disability or abnormal height and weight, signify a specific genetic disorder.
These morphological changes can be subtle, though, and often only specialized medical geneticists are able to detect and interpret these facial clues. What's more, some genetic disorders are so rare that even a specialist may not have encountered it before, much less a general practitioner. Diagnosing rare conditions has improved thanks to genomic testing that can confirm (or refute) a doctor's suspicion. Yet with thousands of variants in each person's genome, identifying the culprit mutation or deletion can be extremely difficult if you don't know what you're looking for.
Facial recognition technology is trying to take some of the guesswork out of this process. Software such as the Face2Gene app use machine learning to compare a picture of a patient against images of thousands of disorders and come back with suggestions of possible diagnoses.
"This is a classic field for artificial intelligence because no human being can really have enough knowledge and enough experience to be able to do this for thousands of different disorders."
"When we met a geneticist for the first time we were pretty blown away with the fact that they actually use their own human pattern recognition" to diagnose patients, says Gelbman. "This is a classic field for AI [artificial intelligence], for machine learning because no human being can really have enough knowledge and enough experience to be able to do this for thousands of different disorders."
When a physician uploads a photo to the app, they are given a list of different diagnostic suggestions, each with a heat map to indicate how similar the facial features are to a classic representation of the syndrome. The physician can hone the suggestions by adding in other symptoms or family history. Gelbman emphasized that the app is a "search and reference tool" and should not "be used to diagnose or treat medical conditions." It is not approved by the FDA as a diagnostic.
"As a tool, we've all been waiting for this, something that can help everyone," says Julian Martinez-Agosto, an associate professor in human genetics and pediatrics at UCLA. He sees the greatest benefit of facial recognition technology in its ability to empower non-specialists to make a diagnosis. Many areas, including rural communities or resource-poor countries, do not have access to either medical geneticists trained in these types of diagnostics or genomic screens. Apps like Face2Gene can help guide a general practitioner or flag diseases they might not be familiar with.
One concern is that most textbook images of genetic disorders come from the West, so the "classic" face of a condition is often a child of European descent.
Maximilian Muenke, a senior investigator at the National Human Genome Research Institute (NHGRI), agrees that in many countries, facial recognition programs could be the only way for a doctor to make a diagnosis.
"There are only geneticists in countries like the U.S., Canada, Europe, Japan. In most countries, geneticists don't exist at all," Muenke says. "In Nigeria, the most populous country in all of Africa with 160 million people, there's not a single clinical geneticist. So in a country like that, facial recognition programs will be sought after and will be extremely useful to help make a diagnosis to the non-geneticists."
One concern about providing this type of technology to a global population is that most textbook images of genetic disorders come from the West, so the "classic" face of a condition is often a child of European descent. However, the defining facial features of some of these disorders manifest differently across ethnicities, leaving clinicians from other geographic regions at a disadvantage.
"Every syndrome is either more easy or more difficult to detect in people from different geographic backgrounds," explains Muenke. For example, "in some countries of Southeast Asia, the eyes are slanted upward, and that happens to be one of the findings that occurs mostly with children with Down Syndrome. So then it might be more difficult for some individuals to recognize Down Syndrome in children from Southeast Asia."
There is a risk that providing this type of diagnostic information online will lead to parents trying to classify their own children.
To combat this issue, Muenke helped develop the Atlas of Human Malformation Syndromes, a database that incorporates descriptions and pictures of patients from every continent. By providing examples of rare genetic disorders in children from outside of the United States and Europe, Muenke hopes to provide clinicians with a better understanding of what to look for in each condition, regardless of where they practice.
There is a risk that providing this type of diagnostic information online will lead to parents trying to classify their own children. Face2Gene is free to download in the app store, although users must be authenticated by the company as a healthcare professional before they can access the database. The NHGRI Atlas can be accessed by anyone through their website. However, Martinez and Muenke say parents already use Google and WebMD to look up their child's symptoms; facial recognition programs and databases are just an extension of that trend. In fact, Martinez says, "Empowering families is another way to facilitate access to care. Some families live in rural areas and have no access to geneticists. If they can use software to get a diagnosis and then contact someone at a large hospital, it can help facilitate the process."
Martinez also says the app could go further by providing greater transparency about how the program makes its assessments. Giving clinicians feedback about why a diagnosis fits certain facial features would offer a valuable teaching opportunity in addition to a diagnostic aid.
Both Martinez and Muenke think the technology is an innovation that could vastly benefit patients. "In the beginning, I was quite skeptical and I could not believe that a machine could replace a human," says Muenke. "However, I am a convert that it actually can help tremendously in making a diagnosis. I think there is a place for facial recognition programs, and I am a firm believer that this will spread over the next five years."
A thinking person.
"A world where people are slotted according to their inborn ability – well, that is Gattaca. That is eugenics."
This was the assessment of Dr. Catherine Bliss, a sociologist who wrote a new book on social science genetics, when asked by MIT Technology Review about polygenic scores that can predict a person's intelligence or performance in school. Like a credit score, a polygenic score is statistical tool that combines a lot of information about a person's genome into a single number. Fears about using polygenic scores for genetic discrimination are understandable, given this country's ugly history of using the science of heredity to justify atrocities like forcible sterilization. But polygenic scores are not the new eugenics. And, rushing to discuss polygenic scores in dystopian terms only contributes to widespread public misunderstanding about genetics.
Can we start genotyping toddlers to identify the budding geniuses among them? The short answer is no.
Let's begin with some background on how polygenic scores are developed. In a genome wide-association study, researchers conduct millions of statistical tests to identify small differences in people's DNA sequence that are correlated with differences in a target outcome (beyond what can attributed to chance or ancestry differences). Successful studies of this sort require enormous sample sizes, but companies like 23andMe are now contributing genetic data from their consumers to research studies, and national biorepositories like U.K. Biobank have put genetic information from hundreds of thousands of people online. When applied to studying blood lipids or myopia, this kind of study strikes people as a straightforward and uncontroversial scientific tool. But it can also be conducted for cognitive and behavioral outcomes, like how many years of school a person has completed. When researchers have finished a genome-wide association study, they are left with a dataset with millions of rows (one for each genetic variant analyzed) and one column with the correlations between each variant and the outcome being studied.
The trick to polygenic scoring is to use these results and apply them to people who weren't participants in the original study. Measure the genes of a new person, weight each one of her millions of genetic variants by its correlation with educational attainment from a genome-wide association study, and then simply add everything up into a single number. Voila! -- you've created a polygenic score for educational attainment. On its face, the idea of "scoring" a person's genotype does immediately suggest Gattaca-type applications. Can we now start screening embryos for their "inborn ability," as Bliss called it? Can we start genotyping toddlers to identify the budding geniuses among them?
The short answer is no. Here are four reasons why dystopian projections about polygenic scores are out of touch with the current science:
The phrase "DNA tests for IQ" makes for an attention-grabbing headline, but it's scientifically meaningless.
First, a polygenic score currently predicts the life outcomes of an individual child with a great deal of uncertainty. The amount of uncertainty around polygenic predictions will decrease in the future, as genetic discovery samples get bigger and genetic studies include more of the variation in the genome, including rare variants that are particular to a few families. But for now, knowing a child's polygenic score predicts his ultimate educational attainment about as well as knowing his family's income, and slightly worse than knowing how far his mother went in school. These pieces of information are also readily available about children before they are born, but no one is writing breathless think-pieces about the dystopian outcomes that will result from knowing whether a pregnant woman graduated from college.
Second, using polygenic scoring for embryo selection requires parents to create embryos using reproductive technology, rather than conceiving them by having sex. The prediction that many women will endure medically-unnecessary IVF, in order to select the embryo with the highest polygenic score, glosses over the invasiveness, indignity, pain, and heartbreak that these hormonal and surgical procedures can entail.
Third, and counterintuitively, a polygenic score might be using DNA to measure aspects of the child's environment. Remember, a child inherits her DNA from her parents, who typically also shape the environment she grows up in. And, children's environments respond to their unique personalities and temperaments. One Icelandic study found that parents' polygenic scores predicted their children's educational attainment, even if the score was constructed using only the half of the parental genome that the child didn't inherit. For example, imagine mom has genetic variant X that makes her more likely to smoke during her pregnancy. Prenatal exposure to nicotine, in turn, affects the child's neurodevelopment, leading to behavior problems in school. The school responds to his behavioral problems with suspension, causing him to miss out on instructional content. A genome-wide association study will collapse this long and winding causal path into a simple correlation -- "genetic variant X is correlated with academic achievement." But, a child's polygenic score, which includes variant X, will partly reflect his likelihood of being exposed to adverse prenatal and school environments.
Finally, the phrase "DNA tests for IQ" makes for an attention-grabbing headline, but it's scientifically meaningless. As I've written previously, it makes sense to talk about a bacterial test for strep throat, because strep throat is a medical condition defined as having streptococcal bacteria growing in the back of your throat. If your strep test is positive, you have strep throat, no matter how serious your symptoms are. But a polygenic score is not a test "for" IQ, because intelligence is not defined at the level of someone's DNA. It doesn't matter how high your polygenic score is, if you can't reason abstractly or learn from experience. Equating your intelligence, a cognitive capacity that is tested behaviorally, with your polygenic score, a number that is a weighted sum of genetic variants discovered to be statistically associated with educational attainment in a hypothesis-free data mining exercise, is misleading about what intelligence is and is not.
The task for many scientists like me, who are interested in understanding why some children do better in school than other children, is to disentangle correlations from causation.
So, if we're not going to build a Gattaca-style genetic hierarchy, what are polygenic scores good for? They are not useless. In fact, they give scientists a valuable new tool for studying how to improve children's lives. The task for many scientists like me, who are interested in understanding why some children do better in school than other children, is to disentangle correlations from causation. The best way to do that is to run an experiment where children are randomized to environments, but often a true experiment is unethical or impractical. You can't randomize children to be born to a teenage mother or to go to school with inexperienced teachers. By statistically controlling for some of the relevant genetic differences between people using a polygenic score, scientists are better able to identify potential environmental causes of differences in children's life outcomes. As we have seen with other methods from genetics, like twin studies, understanding genes illuminates the environment.
Research that examines genetics in relation to social inequality, such as differences in higher education outcomes, will obviously remind people of the horrors of the eugenics movement. Wariness regarding how genetic science will be applied is certainly warranted. But, polygenic scores are not pure measures of "inborn ability," and genome-wide association studies of human intelligence and educational attainment are not inevitably ushering in a new eugenics age.