When doctors couldn’t stop her daughter’s seizures, this mom earned a PhD and found a treatment herself.

Savannah Salazar (left) and her mother, Tracy Dixon-Salazaar, who earned a PhD in neurobiology in the quest for a treatment of her daughter's seizure disorder.
Twenty-eight years ago, Tracy Dixon-Salazaar woke to the sound of her daughter, two-year-old Savannah, in the midst of a medical emergency.
“I entered [Savannah’s room] to see her tiny little body jerking about violently in her bed,” Tracy said in an interview. “I thought she was choking.” When she and her husband frantically called 911, the paramedic told them it was likely that Savannah had had a seizure—a term neither Tracy nor her husband had ever heard before.
Over the next several years, Savannah’s seizures continued and worsened. By age five Savannah was having seizures dozens of times each day, and her parents noticed significant developmental delays. Savannah was unable to use the restroom and functioned more like a toddler than a five-year-old.
Doctors were mystified: Tracy and her husband had no family history of seizures, and there was no event—such as an injury or infection—that could have caused them. Doctors were also confused as to why Savannah’s seizures were happening so frequently despite trying different seizure medications.
Doctors eventually diagnosed Savannah with Lennox-Gaustaut Syndrome, or LGS, an epilepsy disorder with no cure and a poor prognosis. People with LGS are often resistant to several kinds of anti-seizure medications, and often suffer from developmental delays and behavioral problems. People with LGS also have a higher chance of injury as well as a higher chance of sudden unexpected death (SUDEP) due to the frequent seizures. In about 70 percent of cases, LGS has an identifiable cause such as a brain injury or genetic syndrome. In about 30 percent of cases, however, the cause is unknown.
Watching her daughter struggle through repeated seizures was devastating to Tracy and the rest of the family.
“This disease, it comes into your life. It’s uninvited. It’s unannounced and it takes over every aspect of your daily life,” said Tracy in an interview with Today.com. “Plus it’s attacking the thing that is most precious to you—your kid.”
Desperate to find some answers, Tracy began combing the medical literature for information about epilepsy and LGS. She enrolled in college courses to better understand the papers she was reading.
“Ironically, I thought I needed to go to college to take English classes to understand these papers—but soon learned it wasn’t English classes I needed, It was science,” Tracy said. When she took her first college science course, Tracy says, she “fell in love with the subject.”
Tracy was now a caregiver to Savannah, who continued to have hundreds of seizures a month, as well as a full-time student, studying late into the night and while her kids were at school, using classwork as “an outlet for the pain.”
“I couldn’t help my daughter,” Tracy said. “Studying was something I could do.”
Twelve years later, Tracy had earned a PhD in neurobiology.
After her post-doctoral training, Tracy started working at a lab that explored the genetics of epilepsy. Savannah’s doctors hadn’t found a genetic cause for her seizures, so Tracy decided to sequence her genome again to check for other abnormalities—and what she found was life-changing.
Tracy discovered that Savannah had a calcium channel mutation, meaning that too much calcium was passing through Savannah’s neural pathways, leading to seizures. The information made sense to Tracy: Anti-seizure medications often leech calcium from a person’s bones. When doctors had prescribed Savannah calcium supplements in the past to counteract these effects, her seizures had gotten worse every time she took the medication. Tracy took her discovery to Savannah’s doctor, who agreed to prescribe her a calcium blocker.
The change in Savannah was almost immediate.
Within two weeks, Savannah’s seizures had decreased by 95 percent. Once on a daily seven-drug regimen, she was soon weaned to just four, and then three. Amazingly, Tracy started to notice changes in Savannah’s personality and development, too.
“She just exploded in her personality and her talking and her walking and her potty training and oh my gosh she is just so sassy,” Tracy said in an interview.
Since starting the calcium blocker eleven years ago, Savannah has continued to make enormous strides. Though still unable to read or write, Savannah enjoys puzzles and social media. She’s “obsessed” with boys, says Tracy. And while Tracy suspects she’ll never be able to live independently, she and her daughter can now share more “normal” moments—something she never anticipated at the start of Savannah’s journey with LGS. While preparing for an event, Savannah helped Tracy get ready.
“We picked out a dress and it was the first time in our lives that we did something normal as a mother and a daughter,” she said. “It was pretty cool.”
DNA- and RNA-based electronic implants may revolutionize healthcare
The test tubes contain tiny DNA/enzyme-based circuits, which comprise TRUMPET, a new type of electronic device, smaller than a cell.
Implantable electronic devices can significantly improve patients’ quality of life. A pacemaker can encourage the heart to beat more regularly. A neural implant, usually placed at the back of the skull, can help brain function and encourage higher neural activity. Current research on neural implants finds them helpful to patients with Parkinson’s disease, vision loss, hearing loss, and other nerve damage problems. Several of these implants, such as Elon Musk’s Neuralink, have already been approved by the FDA for human use.
Yet, pacemakers, neural implants, and other such electronic devices are not without problems. They require constant electricity, limited through batteries that need replacements. They also cause scarring. “The problem with doing this with electronics is that scar tissue forms,” explains Kate Adamala, an assistant professor of cell biology at the University of Minnesota Twin Cities. “Anytime you have something hard interacting with something soft [like muscle, skin, or tissue], the soft thing will scar. That's why there are no long-term neural implants right now.” To overcome these challenges, scientists are turning to biocomputing processes that use organic materials like DNA and RNA. Other promised benefits include “diagnostics and possibly therapeutic action, operating as nanorobots in living organisms,” writes Evgeny Katz, a professor of bioelectronics at Clarkson University, in his book DNA- And RNA-Based Computing Systems.
While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output.
Adamala’s research focuses on developing such biocomputing systems using DNA, RNA, proteins, and lipids. Using these molecules in the biocomputing systems allows the latter to be biocompatible with the human body, resulting in a natural healing process. In a recent Nature Communications study, Adamala and her team created a new biocomputing platform called TRUMPET (Transcriptional RNA Universal Multi-Purpose GatE PlaTform) which acts like a DNA-powered computer chip. “These biological systems can heal if you design them correctly,” adds Adamala. “So you can imagine a computer that will eventually heal itself.”
The basics of biocomputing
Biocomputing and regular computing have many similarities. Like regular computing, biocomputing works by running information through a series of gates, usually logic gates. A logic gate works as a fork in the road for an electronic circuit. The input will travel one way or another, giving two different outputs. An example logic gate is the AND gate, which has two inputs (A and B) and two different results. If both A and B are 1, the AND gate output will be 1. If only A is 1 and B is 0, the output will be 0 and vice versa. If both A and B are 0, the result will be 0. While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output. In this case, the DNA enters the logic gate as a single or double strand.
If the DNA is double-stranded, the system “digests” the DNA or destroys it, which results in non-fluorescence or “0” output. Conversely, if the DNA is single-stranded, it won’t be digested and instead will be copied by several enzymes in the biocomputing system, resulting in fluorescent RNA or a “1” output. And the output for this type of binary system can be expanded beyond fluorescence or not. For example, a “1” output might be the production of the enzyme insulin, while a “0” may be that no insulin is produced. “This kind of synergy between biology and computation is the essence of biocomputing,” says Stephanie Forrest, a professor and the director of the Biodesign Center for Biocomputing, Security and Society at Arizona State University.
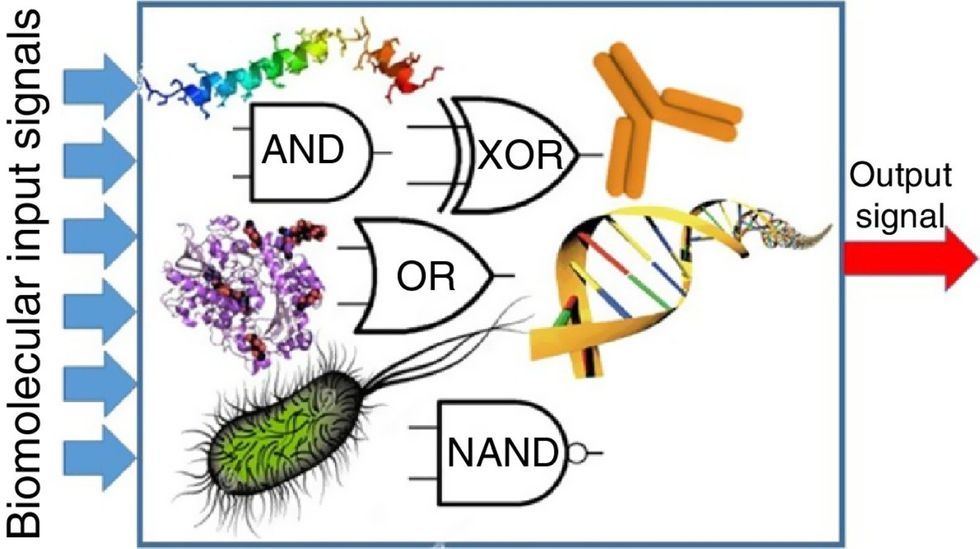
Biocomputing circles are made of DNA, RNA, proteins and even bacteria.
Evgeny Katz
The TRUMPET’s promise
Depending on whether the biocomputing system is placed directly inside a cell within the human body, or run in a test-tube, different environmental factors play a role. When an output is produced inside a cell, the cell's natural processes can amplify this output (for example, a specific protein or DNA strand), creating a solid signal. However, these cells can also be very leaky. “You want the cells to do the thing you ask them to do before they finish whatever their businesses, which is to grow, replicate, metabolize,” Adamala explains. “However, often the gate may be triggered without the right inputs, creating a false positive signal. So that's why natural logic gates are often leaky." While biocomputing outside a cell in a test tube can allow for tighter control over the logic gates, the outputs or signals cannot be amplified by a cell and are less potent.
TRUMPET, which is smaller than a cell, taps into both cellular and non-cellular biocomputing benefits. “At its core, it is a nonliving logic gate system,” Adamala states, “It's a DNA-based logic gate system. But because we use enzymes, and the readout is enzymatic [where an enzyme replicates the fluorescent RNA], we end up with signal amplification." This readout means that the output from the TRUMPET system, a fluorescent RNA strand, can be replicated by nearby enzymes in the platform, making the light signal stronger. "So it combines the best of both worlds,” Adamala adds.
These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body.
The TRUMPET biocomputing process is relatively straightforward. “If the DNA [input] shows up as single-stranded, it will not be digested [by the logic gate], and you get this nice fluorescent output as the RNA is made from the single-stranded DNA, and that's a 1,” Adamala explains. "And if the DNA input is double-stranded, it gets digested by the enzymes in the logic gate, and there is no RNA created from the DNA, so there is no fluorescence, and the output is 0." On the story's leading image above, if the tube is "lit" with a purple color, that is a binary 1 signal for computing. If it's "off" it is a 0.
While still in research, TRUMPET and other biocomputing systems promise significant benefits to personalized healthcare and medicine. These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body. The study’s lead author and graduate student Judee Sharon is already beginning to research TRUMPET's ability for earlier cancer diagnoses. Because the inputs for TRUMPET are single or double-stranded DNA, any mutated or cancerous DNA could theoretically be detected from the platform through the biocomputing process. Theoretically, devices like TRUMPET could be used to detect cancer and other diseases earlier.
Adamala sees TRUMPET not only as a detection system but also as a potential cancer drug delivery system. “Ideally, you would like the drug only to turn on when it senses the presence of a cancer cell. And that's how we use the logic gates, which work in response to inputs like cancerous DNA. Then the output can be the production of a small molecule or the release of a small molecule that can then go and kill what needs killing, in this case, a cancer cell. So we would like to develop applications that use this technology to control the logic gate response of a drug’s delivery to a cell.”
Although platforms like TRUMPET are making progress, a lot more work must be done before they can be used commercially. “The process of translating mechanisms and architecture from biology to computing and vice versa is still an art rather than a science,” says Forrest. “It requires deep computer science and biology knowledge,” she adds. “Some people have compared interdisciplinary science to fusion restaurants—not all combinations are successful, but when they are, the results are remarkable.”
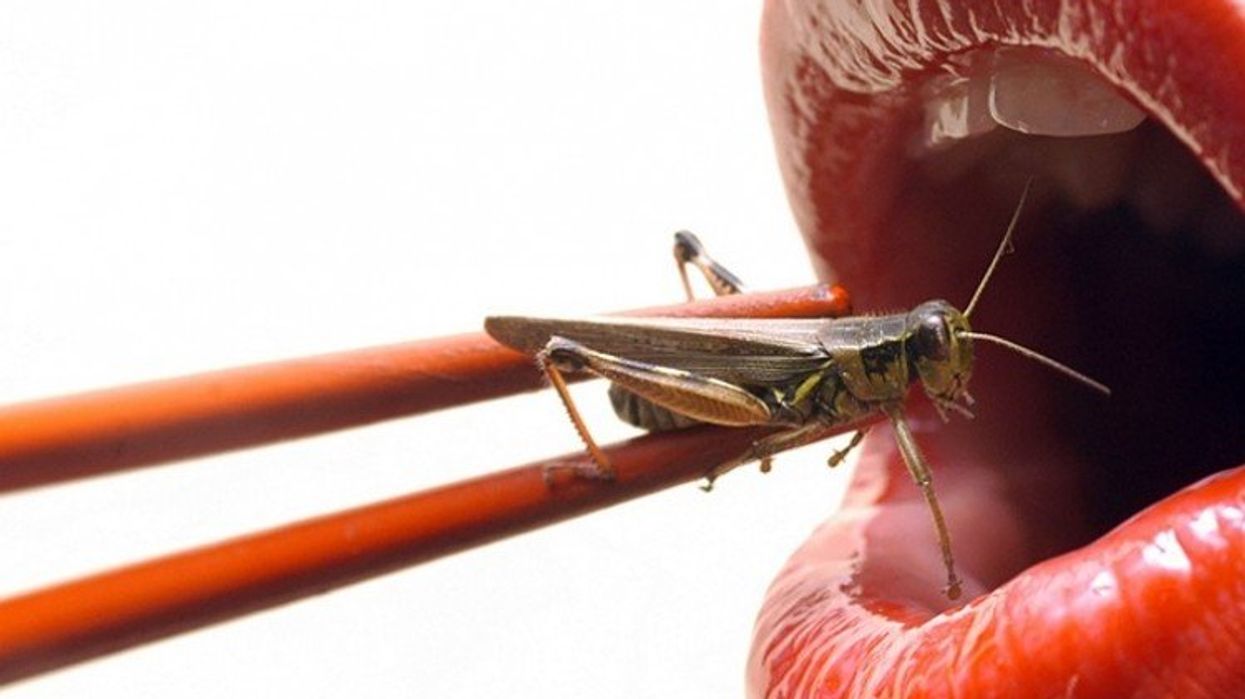
Crickets are low on fat, high on protein, and can be farmed sustainably. They are also crunchy.
In today’s podcast episode, Leaps.org Deputy Editor Lina Zeldovich speaks about the health and ecological benefits of farming crickets for human consumption with Bicky Nguyen, who joins Lina from Vietnam. Bicky and her business partner Nam Dang operate an insect farm named CricketOne. Motivated by the idea of sustainable and healthy protein production, they started their unconventional endeavor a few years ago, despite numerous naysayers who didn’t believe that humans would ever consider munching on bugs.
Yet, making creepy crawlers part of our diet offers many health and planetary advantages. Food production needs to match the rise in global population, estimated to reach 10 billion by 2050. One challenge is that some of our current practices are inefficient, polluting and wasteful. According to nonprofit EarthSave.org, it takes 2,500 gallons of water, 12 pounds of grain, 35 pounds of topsoil and the energy equivalent of one gallon of gasoline to produce one pound of feedlot beef, although exact statistics vary between sources.
Meanwhile, insects are easy to grow, high on protein and low on fat. When roasted with salt, they make crunchy snacks. When chopped up, they transform into delicious pâtes, says Bicky, who invents her own cricket recipes and serves them at industry and public events. Maybe that’s why some research predicts that edible insects market may grow to almost $10 billion by 2030. Tune in for a delectable chat on this alternative and sustainable protein.
Listen on Apple | Listen on Spotify | Listen on Stitcher | Listen on Amazon | Listen on Google
Further reading:
More info on Bicky Nguyen
https://yseali.fulbright.edu.vn/en/faculty/bicky-n...
The environmental footprint of beef production
https://www.earthsave.org/environment.htm
https://www.watercalculator.org/news/articles/beef-king-big-water-footprints/
https://www.frontiersin.org/articles/10.3389/fsufs.2019.00005/full
https://ourworldindata.org/carbon-footprint-food-methane
Insect farming as a source of sustainable protein
https://www.insectgourmet.com/insect-farming-growing-bugs-for-protein/
https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/insect-farming
Cricket flour is taking the world by storm
https://www.cricketflours.com/
https://talk-commerce.com/blog/what-brands-use-cricket-flour-and-why/

Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.