With this new technology, hospitals and pharmacies could make vaccines and medicines onsite
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.

New research focuses on methods that could change medicine-making worldwide. The scientists propose bursting cells open, removing their DNA and using the cellular gears inside to make therapies.
Most modern biopharmaceutical medicines are produced by workhorse cells—typically bacterial but sometimes mammalian. The cells receive the synthesizing instructions on a snippet of a genetic code, which they incorporate into their DNA. The cellular machinery—ribosomes, RNAs, polymerases, and other compounds—read and use these instructions to build the medicinal molecules, which are harvested and administered to patients.
Although a staple of modern pharma, this process is complex and expensive. One must first insert the DNA instructions into the cells, which they may or may not uptake. One then must grow the cells, keeping them alive and well, so that they produce the required therapeutics, which then must be isolated and purified. To make this at scale requires massive bioreactors and big factories from where the drugs are distributed—and may take a while to arrive where they’re needed. “The pandemic showed us that this method is slow and cumbersome,” says Govind Rao, professor of biochemical engineering who directs the Center for Advanced Sensor Technology at the University of Maryland, Baltimore County (UMBC). “We need better methods that can work faster and can work locally where an outbreak is happening.”
Rao and his team of collaborators, which spans multiple research institutions, believe they have a better approach that may change medicine-making worldwide. They suggest forgoing the concept of using living cells as medicine-producers. Instead, they propose breaking the cells and using the remaining cellular gears for assembling the therapeutic compounds. Instead of inserting the DNA into living cells, the team burst them open, and removed their DNA altogether. Yet, the residual molecular machinery of ribosomes, polymerases and other cogwheels still functioned the way it would in a cell. “Now if you drop your DNA drug-making instructions into that soup, this machinery starts making what you need,” Rao explains. “And because you're no longer worrying about living cells, it becomes much simpler and more efficient.” The collaborators detail their cell-free protein synthesis or CFPS method in their recent paper published in preprint BioAxiv.
While CFPS does not use living cells, it still needs the basic building blocks to assemble proteins from—such as amino acids, nucleotides and certain types of enzymes. These are regularly added into this “soup” to keep the molecular factory chugging. “We just mix everything in as a batch and we let it integrate,” says James Robert Swartz, professor of chemical engineering and bioengineering at Stanford University and co-author of the paper. “And we make sure that we provide enough oxygen.” Rao likens the process to making milk from milk powder.
For a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology.
The idea of a cell-free protein synthesis is older than one might think. Swartz first experimented with it around 1997, when he was a chemical engineer at Genentech. While working on engineering bacteria to make pharmaceuticals, he discovered that there was a limit to what E. coli cells, the workhorse darling of pharma, could do. For example, it couldn’t grow and properly fold some complex proteins. “We tried many genetic engineering approaches, many fermentation, development, and environmental control approaches,” Swartz recalls—to no avail.
“The organism had its own agenda,” he quips. “And because everything was happening within the organism, we just couldn't really change those conditions very easily. Some of them we couldn’t change at all—we didn’t have control.”
It was out of frustration with the defiant bacteria that a new idea took hold. Could the cells be opened instead, so that the protein-forming reactions could be influenced more easily? “Obviously, we’d lose the ability for them to reproduce,” Swartz says. But that also meant that they no longer needed to keep the cells alive and could focus on making the specific reactions happen. “We could take the catalysts, the enzymes, and the more complex catalysts and activate them, make them work together, much as they would in a living cell, but the way we wanted.”
In 1998, Swartz joined Stanford, and began perfecting the biochemistry of the cell-free method, identifying the reactions he wanted to foster and stopping those he didn’t want. He managed to make the idea work, but for a variety of reasons—from the field’s general inertia to regulatory approval hurdles—the method hasn’t become mainstream. The pandemic rekindled interest in medicines that can be made quickly and easily, so it drew more attention to the technology. For their BioArxiv paper, the team tested the method by growing a specific antiviral protein called griffithsin.
First identified by Barry O’Keefe at National Cancer Institute over a decade ago, griffithsin is an antiviral known to interfere with many viruses’ ability to enter cells—including HIV, SARS, SARS-CoV-2, MERS and others. Originally isolated from the red algae Griffithsia, it works differently from antibodies and antibody cocktails.
Most antiviral medicines tend to target the specific receptors that viruses use to gain entry to the cells they infect. For example, SARS-CoV-2 uses the infamous spike protein to latch onto the ACE2 receptor of mammalian cells. The antibodies or other antiviral molecules stick to the spike protein, shutting off its ability to cling onto the ACE2 receptors. Unfortunately, the spike proteins mutate very often, so the medicines lose their potency. On the contrary, griffithsin has the ability to cling to the different parts of viral shells called capsids—namely to the molecules of mannose, a type of sugar. That extra stuff, glued all around the capsid like dead weight, makes it impossible for the virus to squeeze into the cell.
“Every time we have a vaccine or an antibody against a specific SARS-CoV-2 strain, that strain then mutates and so you lose efficacy,” Rao explains. “But griffithsin molecules glom onto the viral capsid, so the capsid essentially becomes a sticky mess and can’t enter the cell.” Mannose molecules also don’t mutate as easily as viruses’ receptors, so griffithsin-based antivirals do not have to be constantly updated. And because mannose molecules are found on many viruses’ capsids, it makes griffithsin “a universal neutralizer,” Rao explains.
“When griffithsin was discovered, we recognized that it held a lot of promise as a potential antiviral agent,” O’Keefe says. In 2010, he published a paper about griffithsin efficacy in neutralizing viruses of the corona family—after the first SARS outbreak in the early 2000s, the scientific community was interested in such antivirals. Yet, griffithsin is still not available as an off-the-shelf product. So during the Covid pandemic, the team experimented with synthesizing griffithsin using the cell-free production method. They were able to generate potent griffithsin in less than 24 hours without having to grow living cells.
The antiviral protein isn't the only type of medicine that can be made cell-free. The proteins needed for vaccine production could also be made the same way. “Such portable, on-demand drug manufacturing platforms can produce antiviral proteins within hours, making them ideal for combating future pandemics,” Rao says. “We would be able to stop the pandemic before it spreads.”
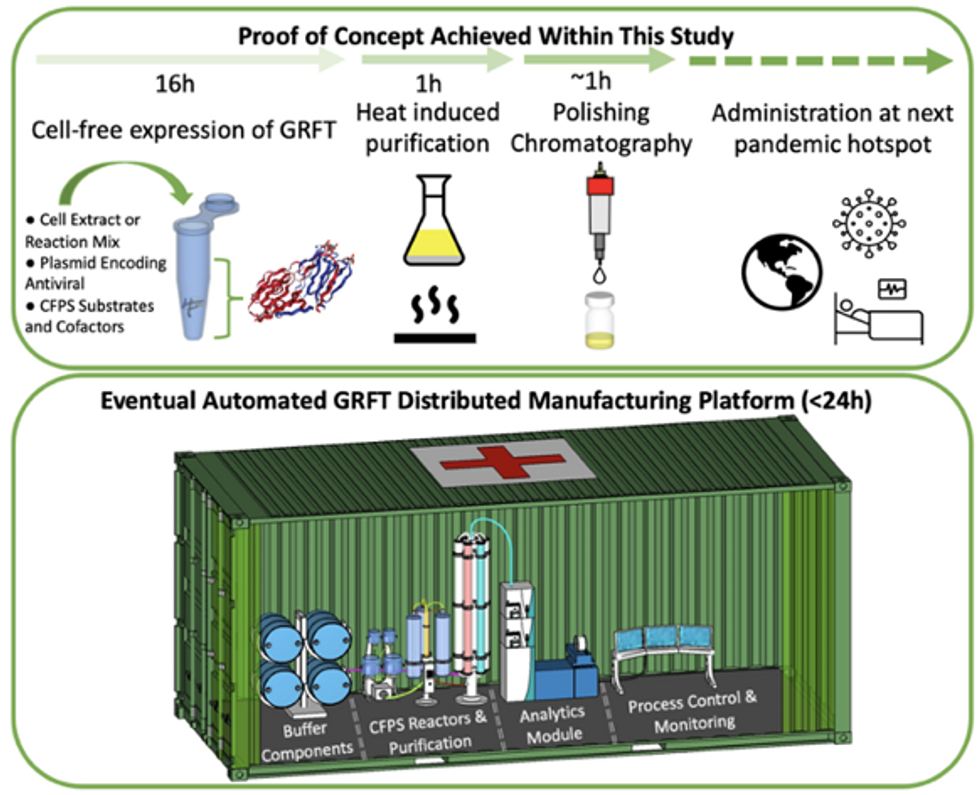
Top: Describes the process used in the study. Bottom: Describes how the new medicines and vaccines could be made at the site of a future viral outbreak.
Image courtesy of Rao and team, sourced from An approach to rapid distributed manufacturing of broad spectrumanti-viral griffithsin using cell-free systems to mitigate pandemics.
Rao’s idea is to perfect the technology to the point that any hospital or pharmacy can load up the media containing molecular factories, mix up the required amino acids, nucleotides and enzymes, and harvest the meds within hours. That will allow making medicines onsite and on demand. “That would be a self-contained production unit, so that you could just ship the production wherever the pandemic is breaking out,” says Swartz.
These units and the meds they produce, will, of course, have to undergo rigorous testing. “The biggest hurdles will be validating these against conventional technology,” Rao says. The biotech industry is risk-averse and prefers the familiar methods. But if this approach works, it may go beyond emergency situations and revolutionize the medicine-making paradigm even outside hospitals and pharmacies. Rao hopes that someday the method might become so mainstream that people may be able to buy and operate such reactors at home. “You can imagine a diabetic patient making insulin that way, or some other drugs,” Rao says. It would work not unlike making baby formula from the mere white powder. Just add water—and some oxygen, too.
Lina Zeldovich has written about science, medicine and technology for Popular Science, Smithsonian, National Geographic, Scientific American, Reader’s Digest, the New York Times and other major national and international publications. A Columbia J-School alumna, she has won several awards for her stories, including the ASJA Crisis Coverage Award for Covid reporting, and has been a contributing editor at Nautilus Magazine. In 2021, Zeldovich released her first book, The Other Dark Matter, published by the University of Chicago Press, about the science and business of turning waste into wealth and health. You can find her on http://linazeldovich.com/ and @linazeldovich.
A movie still from the 1966 film "Fantastic Voyage"
In the 1966 movie "Fantastic Voyage," actress Raquel Welch and her submarine were shrunk to the size of a cell in order to eliminate a blood clot in a scientist's brain. Now, 55 years later, the scenario is becoming closer to reality.
California-based startup Bionaut Labs has developed a nanobot about the size of a grain of rice that's designed to transport medication to the exact location in the body where it's needed. If you think about it, the conventional way to deliver medicine makes little sense: A painkiller affects the entire body instead of just the arm that's hurting, and chemotherapy is flushed through all the veins instead of precisely targeting the tumor.
"Chemotherapy is delivered systemically," Bionaut-founder and CEO Michael Shpigelmacher says. "Often only a small percentage arrives at the location where it is actually needed."
But what if it was possible to send a tiny robot through the body to attack a tumor or deliver a drug at exactly the right location?
Several startups and academic institutes worldwide are working to develop such a solution but Bionaut Labs seems the furthest along in advancing its invention. "You can think of the Bionaut as a tiny screw that moves through the veins as if steered by an invisible screwdriver until it arrives at the tumor," Shpigelmacher explains. Via Zoom, he shares the screen of an X-ray machine in his Culver City lab to demonstrate how the half-transparent, yellowish device winds its way along the spine in the body. The nanobot contains a tiny but powerful magnet. The "invisible screwdriver" is an external magnetic field that rotates that magnet inside the device and gets it to move and change directions.
The current model has a diameter of less than a millimeter. Shpigelmacher's engineers could build the miniature vehicle even smaller but the current size has the advantage of being big enough to see with bare eyes. It can also deliver more medicine than a tinier version. In the Zoom demonstration, the micorobot is injected into the spine, not unlike an epidural, and pulled along the spine through an outside magnet until the Bionaut reaches the brainstem. Depending which organ it needs to reach, it could be inserted elsewhere, for instance through a catheter.
"The hope is that we can develop a vehicle to transport medication deep into the body," says Max Planck scientist Tian Qiu.
Imagine moving a screw through a steak with a magnet — that's essentially how the device works. But of course, the Bionaut is considerably different from an ordinary screw: "At the right location, we give a magnetic signal, and it unloads its medicine package," Shpigelmacher says.
To start, Bionaut Labs wants to use its device to treat Parkinson's disease and brain stem gliomas, a type of cancer that largely affects children and teenagers. About 300 to 400 young people a year are diagnosed with this type of tumor. Radiation and brain surgery risk damaging sensitive brain tissue, and chemotherapy often doesn't work. Most children with these tumors live less than 18 months. A nanobot delivering targeted chemotherapy could be a gamechanger. "These patients really don't have any other hope," Shpigelmacher says.
Of course, the main challenge of the developing such a device is guaranteeing that it's safe. Because tissue is so sensitive, any mistake could risk disastrous results. In recent years, Bionaut has tested its technology in dozens of healthy sheep and pigs with no major adverse effects. Sheep make a good stand-in for humans because their brains and spines are similar to ours.
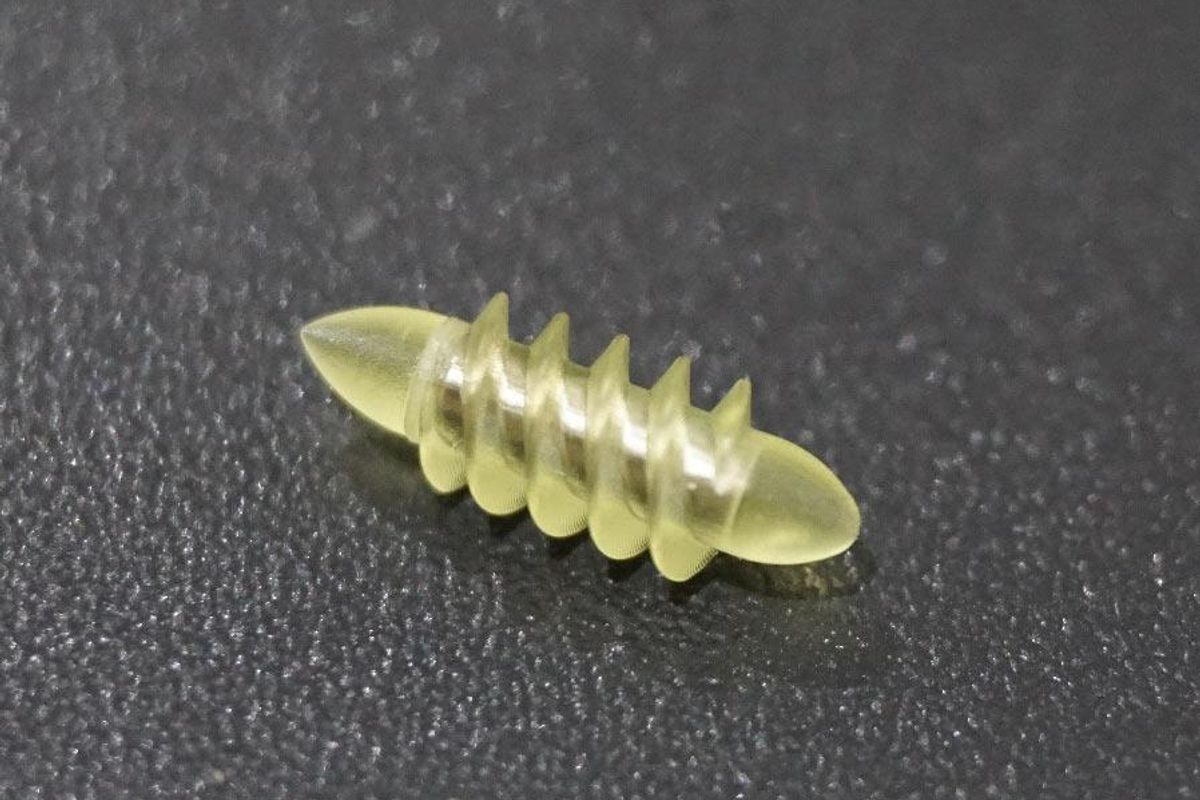

The Bionaut device is about the size of a grain of rice.
Bionaut Labs
"As the Bionaut moves through brain tissue, it creates a transient track that heals within a few weeks," Shpigelmacher says. The company is hoping to be the first to test a nanobot in humans. In December 2022, it announced that a recent round of funding drew $43.2 million, for a total of 63.2 million, enabling more research and, if all goes smoothly, human clinical trials by early next year.
Once the technique has been perfected, further applications could include addressing other kinds of brain disorders that are considered incurable now, such as Alzheimer's or Huntington's disease. "Microrobots could serve as a bridgehead, opening the gateway to the brain and facilitating precise access of deep brain structure – either to deliver medication, take cell samples or stimulate specific brain regions," Shpigelmacher says.
Robot-assisted hybrid surgery with artificial intelligence is already used in state-of-the-art surgery centers, and many medical experts believe that nanorobotics will be the instrument of the future. In 2016, three scientists were awarded the Nobel Prize in Chemistry for their development of "the world's smallest machines," nano "elevators" and minuscule motors. Since then, the scientific experiments have progressed to the point where applicable devices are moving closer to actually being implemented.
Bionaut's technology was initially developed by a research team lead by Peer Fischer, head of the independent Micro Nano and Molecular Systems Lab at the Max Planck Institute for Intelligent Systems in Stuttgart, Germany. Fischer is considered a pioneer in the research of nano systems, which he began at Harvard University more than a decade ago. He and his team are advising Bionaut Labs and have licensed their technology to the company.
"The hope is that we can develop a vehicle to transport medication deep into the body," says Max Planck scientist Tian Qiu, who leads the cooperation with Bionaut Labs. He agrees with Shpigelmacher that the Bionaut's size is perfect for transporting medication loads and is researching potential applications for even smaller nanorobots, especially in the eye, where the tissue is extremely sensitive. "Nanorobots can sneak through very fine tissue without causing damage."
In "Fantastic Voyage," Raquel Welch's adventures inside the body of a dissident scientist let her swim through his veins into his brain, but her shrunken miniature submarine is attacked by antibodies; she has to flee through the nerves into the scientist's eye where she escapes into freedom on a tear drop. In reality, the exit in the lab is much more mundane. The Bionaut simply leaves the body through the same port where it entered. But apart from the dramatization, the "Fantastic Voyage" was almost prophetic, or, as Shpigelmacher says, "Science fiction becomes science reality."
This article was first published by Leaps.org on April 12, 2021.
How the Human Brain Project Built a Mind of its Own
In 2013, the Human Brain Project set out to build a realistic computer model of the brain over ten years. Now, experts are reflecting on HBP's achievements with an eye toward the future.
In 2009, neuroscientist Henry Markram gave an ambitious TED talk. “Our mission is to build a detailed, realistic computer model of the human brain,” he said, naming three reasons for this unmatched feat of engineering. One was because understanding the human brain was essential to get along in society. Another was because experimenting on animal brains could only get scientists so far in understanding the human ones. Third, medicines for mental disorders weren’t good enough. “There are two billion people on the planet that are affected by mental disorders, and the drugs that are used today are largely empirical,” Markram said. “I think that we can come up with very concrete solutions on how to treat disorders.”
Markram's arguments were very persuasive. In 2013, the European Commission launched the Human Brain Project, or HBP, as part of its Future and Emerging Technologies program. Viewed as Europe’s chance to try to win the “brain race” between the U.S., China, Japan, and other countries, the project received about a billion euros in funding with the goal to simulate the entire human brain on a supercomputer, or in silico, by 2023.
Now, after 10 years of dedicated neuroscience research, the HBP is coming to an end. As its many critics warned, it did not manage to build an entire human brain in silico. Instead, it achieved a multifaceted array of different goals, some of them unexpected.
Scholars have found that the project did help advance neuroscience more than some detractors initially expected, specifically in the area of brain simulations and virtual models. Using an interdisciplinary approach of combining technology, such as AI and digital simulations, with neuroscience, the HBP worked to gain a deeper understanding of the human brain’s complicated structure and functions, which in some cases led to novel treatments for brain disorders. Lastly, through online platforms, the HBP spearheaded a previously unmatched level of global neuroscience collaborations.
Simulating a human brain stirs up controversy
Right from the start, the project was plagued with controversy and condemnation. One of its prominent critics was Yves Fregnac, a professor in cognitive science at the Polytechnic Institute of Paris and research director at the French National Centre for Scientific Research. Fregnac argued in numerous articles that the HBP was overfunded based on proposals with unrealistic goals. “This new way of over-selling scientific targets, deeply aligned with what modern society expects from mega-sciences in the broad sense (big investment, big return), has been observed on several occasions in different scientific sub-fields,” he wrote in one of his articles, “before invading the field of brain sciences and neuromarketing.”
"A human brain model can simulate an experiment a million times for many different conditions, but the actual human experiment can be performed only once or a few times," said Viktor Jirsa, a professor at Aix-Marseille University.
Responding to such critiques, the HBP worked to restructure the effort in its early days with new leadership, organization, and goals that were more flexible and attainable. “The HBP got a more versatile, pluralistic approach,” said Viktor Jirsa, a professor at Aix-Marseille University and one of the HBP lead scientists. He believes that these changes fixed at least some of HBP’s issues. “The project has been on a very productive and scientifically fruitful course since then.”
After restructuring, the HBP became a European hub on brain research, with hundreds of scientists joining its growing network. The HBP created projects focused on various brain topics, from consciousness to neurodegenerative diseases. HBP scientists worked on complex subjects, such as mapping out the brain, combining neuroscience and robotics, and experimenting with neuromorphic computing, a computational technique inspired by the human brain structure and function—to name just a few.
Simulations advance knowledge and treatment options
In 2013, it seemed that bringing neuroscience into a digital age would be farfetched, but research within the HBP has made this achievable. The virtual maps and simulations various HBP teams create through brain imaging data make it easier for neuroscientists to understand brain developments and functions. The teams publish these models on the HBP’s EBRAINS online platform—one of the first to offer access to such data to neuroscientists worldwide via an open-source online site. “This digital infrastructure is backed by high-performance computers, with large datasets and various computational tools,” said Lucy Xiaolu Wang, an assistant professor in the Resource Economics Department at the University of Massachusetts Amherst, who studies the economics of the HBP. That means it can be used in place of many different types of human experimentation.
Jirsa’s team is one of many within the project that works on virtual brain models and brain simulations. Compiling patient data, Jirsa and his team can create digital simulations of different brain activities—and repeat these experiments many times, which isn’t often possible in surgeries on real brains. “A human brain model can simulate an experiment a million times for many different conditions,” Jirsa explained, “but the actual human experiment can be performed only once or a few times.” Using simulations also saves scientists and doctors time and money when looking at ways to diagnose and treat patients with brain disorders.
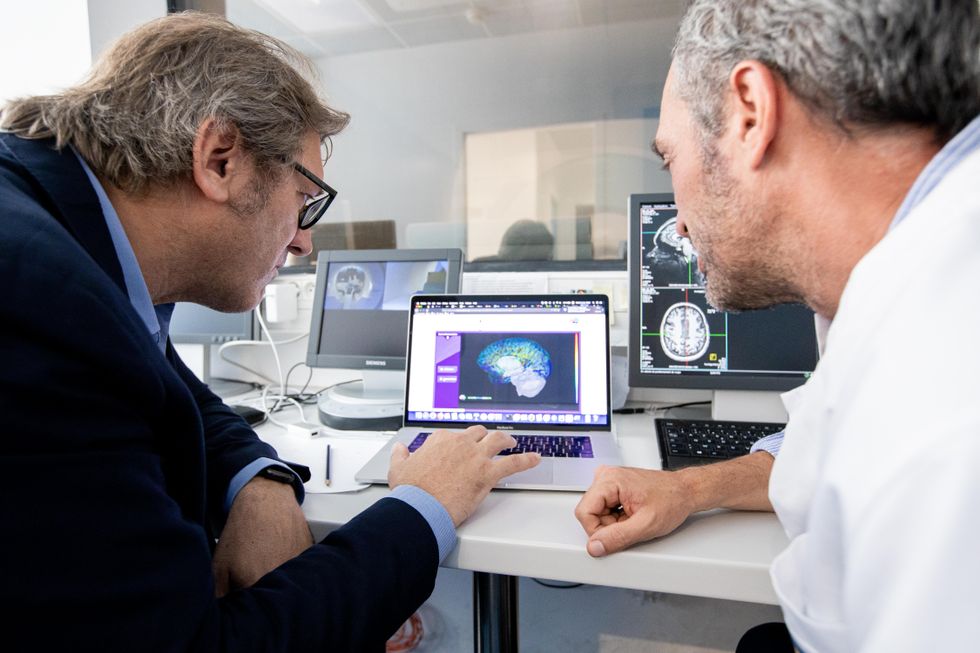

Compiling patient data, scientists can create digital simulations of different brain activities—and repeat these experiments many times.
The Human Brain Project
Simulations can help scientists get a full picture that otherwise is unattainable. “Another benefit is data completion,” added Jirsa, “in which incomplete data can be complemented by the model. In clinical settings, we can often measure only certain brain areas, but when linked to the brain model, we can enlarge the range of accessible brain regions and make better diagnostic predictions.”
With time, Jirsa’s team was able to move into patient-specific simulations. “We advanced from generic brain models to the ability to use a specific patient’s brain data, from measurements like MRI and others, to create individualized predictive models and simulations,” Jirsa explained. He and his team are working on this personalization technique to treat patients with epilepsy. According to the World Health Organization, about 50 million people worldwide suffer from epilepsy, a disorder that causes recurring seizures. While some epilepsy causes are known others remain an enigma, and many are hard to treat. For some patients whose epilepsy doesn’t respond to medications, removing part of the brain where seizures occur may be the only option. Understanding where in the patients’ brains seizures arise can give scientists a better idea of how to treat them and whether to use surgery versus medications.
“We apply such personalized models…to precisely identify where in a patient’s brain seizures emerge,” Jirsa explained. “This guides individual surgery decisions for patients for which surgery is the only treatment option.” He credits the HBP for the opportunity to develop this novel approach. “The personalization of our epilepsy models was only made possible by the Human Brain Project, in which all the necessary tools have been developed. Without the HBP, the technology would not be in clinical trials today.”
Personalized simulations can significantly advance treatments, predict the outcome of specific medical procedures and optimize them before actually treating patients. Jirsa is watching this happen firsthand in his ongoing research. “Our technology for creating personalized brain models is now used in a large clinical trial for epilepsy, funded by the French state, where we collaborate with clinicians in hospitals,” he explained. “We have also founded a spinoff company called VB Tech (Virtual Brain Technologies) to commercialize our personalized brain model technology and make it available to all patients.”
The Human Brain Project created a level of interconnectedness within the neuroscience research community that never existed before—a network not unlike the brain’s own.
Other experts believe it’s too soon to tell whether brain simulations could change epilepsy treatments. “The life cycle of developing treatments applicable to patients often runs over a decade,” Wang stated. “It is still too early to draw a clear link between HBP’s various project areas with patient care.” However, she admits that some studies built on the HBP-collected knowledge are already showing promise. “Researchers have used neuroscientific atlases and computational tools to develop activity-specific stimulation programs that enabled paraplegic patients to move again in a small-size clinical trial,” Wang said. Another intriguing study looked at simulations of Alzheimer’s in the brain to understand how it evolves over time.
Some challenges remain hard to overcome even with computer simulations. “The major challenge has always been the parameter explosion, which means that many different model parameters can lead to the same result,” Jirsa explained. An example of this parameter explosion could be two different types of neurodegenerative conditions, such as Parkinson’s and Huntington’s diseases. Both afflict the same area of the brain, the basal ganglia, which can affect movement, but are caused by two different underlying mechanisms. “We face the same situation in the living brain, in which a large range of diverse mechanisms can produce the same behavior,” Jirsa said. The simulations still have to overcome the same challenge.
Understanding where in the patients’ brains seizures arise can give scientists a better idea of how to treat them and whether to use surgery versus medications.
The Human Brain Project
A network not unlike the brain’s own
Though the HBP will be closing this year, its legacy continues in various studies, spin-off companies, and its online platform, EBRAINS. “The HBP is one of the earliest brain initiatives in the world, and the 10-year long-term goal has united many researchers to collaborate on brain sciences with advanced computational tools,” Wang said. “Beyond the many research articles and projects collaborated on during the HBP, the online neuroscience research infrastructure EBRAINS will be left as a legacy even after the project ends.”
Those who worked within the HBP see the end of this project as the next step in neuroscience research. “Neuroscience has come closer to very meaningful applications through the systematic link with new digital technologies and collaborative work,” Jirsa stated. “In that way, the project really had a pioneering role.” It also created a level of interconnectedness within the neuroscience research community that never existed before—a network not unlike the brain’s own. “Interconnectedness is an important advance and prerequisite for progress,” Jirsa said. “The neuroscience community has in the past been rather fragmented and this has dramatically changed in recent years thanks to the Human Brain Project.”
According to its website, by 2023 HBP’s network counted over 500 scientists from over 123 institutions and 16 different countries, creating one of the largest multi-national research groups in the world. Even though the project hasn’t produced the in-silico brain as Markram envisioned it, the HBP created a communal mind with immense potential. “It has challenged us to think beyond the boundaries of our own laboratories,” Jirsa said, “and enabled us to go much further together than we could have ever conceived going by ourselves.”