Meet the Scientists on the Frontlines of Protecting Humanity from a Man-Made Pathogen

From left: Jean Peccoud, Randall Murch, and Neeraj Rao.
Jean Peccoud wasn't expecting an email from the FBI. He definitely wasn't expecting the agency to invite him to a meeting. "My reaction was, 'What did I do wrong to be on the FBI watch list?'" he recalls.
You use those blueprints for white-hat research—which is, indeed, why the open blueprints exist—or you can do the same for a black-hat attack.
He didn't know what the feds could possibly want from him. "I was mostly scared at this point," he says. "I was deeply disturbed by the whole thing."
But he decided to go anyway, and when he traveled to San Francisco for the 2008 gathering, the reason for the e-vite became clear: The FBI was reaching out to researchers like him—scientists interested in synthetic biology—in anticipation of the potential nefarious uses of this technology. "The whole purpose of the meeting was, 'Let's start talking to each other before we actually need to talk to each other,'" says Peccoud, now a professor of chemical and biological engineering at Colorado State University. "'And let's make sure next time you get an email from the FBI, you don't freak out."
Synthetic biology—which Peccoud defines as "the application of engineering methods to biological systems"—holds great power, and with that (as always) comes great responsibility. When you can synthesize genetic material in a lab, you can create new ways of diagnosing and treating people, and even new food ingredients. But you can also "print" the genetic sequence of a virus or virulent bacterium.
And while it's not easy, it's also not as hard as it could be, in part because dangerous sequences have publicly available blueprints. You use those blueprints for white-hat research—which is, indeed, why the open blueprints exist—or you can do the same for a black-hat attack. You could synthesize a dangerous pathogen's code on purpose, or you could unwittingly do so because someone tampered with your digital instructions. Ordering synthetic genes for viral sequences, says Peccoud, would likely be more difficult today than it was a decade ago.
"There is more awareness of the industry, and they are taking this more seriously," he says. "There is no specific regulation, though."
Trying to lock down the interconnected machines that enable synthetic biology, secure its lab processes, and keep dangerous pathogens out of the hands of bad actors is part of a relatively new field: cyberbiosecurity, whose name Peccoud and colleagues introduced in a 2018 paper.
Biological threats feel especially acute right now, during the ongoing pandemic. COVID-19 is a natural pathogen -- not one engineered in a lab. But future outbreaks could start from a bug nature didn't build, if the wrong people get ahold of the right genetic sequences, and put them in the right sequence. Securing the equipment and processes that make synthetic biology possible -- so that doesn't happen -- is part of why the field of cyberbiosecurity was born.
The Origin Story
It is perhaps no coincidence that the FBI pinged Peccoud when it did: soon after a journalist ordered a sequence of smallpox DNA and wrote, for The Guardian, about how easy it was. "That was not good press for anybody," says Peccoud. Previously, in 2002, the Pentagon had funded SUNY Stonybrook researchers to try something similar: They ordered bits of polio DNA piecemeal and, over the course of three years, strung them together.
Although many years have passed since those early gotchas, the current patchwork of regulations still wouldn't necessarily prevent someone from pulling similar tricks now, and the technological systems that synthetic biology runs on are more intertwined — and so perhaps more hackable — than ever. Researchers like Peccoud are working to bring awareness to those potential problems, to promote accountability, and to provide early-detection tools that would catch the whiff of a rotten act before it became one.
Peccoud notes that if someone wants to get access to a specific pathogen, it is probably easier to collect it from the environment or take it from a biodefense lab than to whip it up synthetically. "However, people could use genetic databases to design a system that combines different genes in a way that would make them dangerous together without each of the components being dangerous on its own," he says. "This would be much more difficult to detect."
After his meeting with the FBI, Peccoud grew more interested in these sorts of security questions. So he was paying attention when, in 2010, the Department of Health and Human Services — now helping manage the response to COVID-19 — created guidance for how to screen synthetic biology orders, to make sure suppliers didn't accidentally send bad actors the sequences that make up bad genomes.
Guidance is nice, Peccoud thought, but it's just words. He wanted to turn those words into action: into a computer program. "I didn't know if it was something you can run on a desktop or if you need a supercomputer to run it," he says. So, one summer, he tasked a team of student researchers with poring over the sentences and turning them into scripts. "I let the FBI know," he says, having both learned his lesson and wanting to get in on the game.
Peccoud later joined forces with Randall Murch, a former FBI agent and current Virginia Tech professor, and a team of colleagues from both Virginia Tech and the University of Nebraska-Lincoln, on a prototype project for the Department of Defense. They went into a lab at the University of Nebraska at Lincoln and assessed all its cyberbio-vulnerabilities. The lab develops and produces prototype vaccines, therapeutics, and prophylactic components — exactly the kind of place that you always, and especially right now, want to keep secure.
"We were creating wiki of all these nasty things."
The team found dozens of Achilles' heels, and put them in a private report. Not long after that project, the two and their colleagues wrote the paper that first used the term "cyberbiosecurity." A second paper, led by Murch, came out five months later and provided a proposed definition and more comprehensive perspective on cyberbiosecurity. But although it's now a buzzword, it's the definition, not the jargon, that matters. "Frankly, I don't really care if they call it cyberbiosecurity," says Murch. Call it what you want: Just pay attention to its tenets.
A Database of Scary Sequences
Peccoud and Murch, of course, aren't the only ones working to screen sequences and secure devices. At the nonprofit Battelle Memorial Institute in Columbus, Ohio, for instance, scientists are working on solutions that balance the openness inherent to science and the closure that can stop bad stuff. "There's a challenge there that you want to enable research but you want to make sure that what people are ordering is safe," says the organization's Neeraj Rao.
Rao can't talk about the work Battelle does for the spy agency IARPA, the Intelligence Advanced Research Projects Activity, on a project called Fun GCAT, which aims to use computational tools to deep-screen gene-sequence orders to see if they pose a threat. It can, though, talk about a twin-type internal project: ThreatSEQ (pronounced, of course, "threat seek").
The project started when "a government customer" (as usual, no one will say which) asked Battelle to curate a list of dangerous toxins and pathogens, and their genetic sequences. The researchers even started tagging sequences according to their function — like whether a particular sequence is involved in a germ's virulence or toxicity. That helps if someone is trying to use synthetic biology not to gin up a yawn-inducing old bug but to engineer a totally new one. "How do you essentially predict what the function of a novel sequence is?" says Rao. You look at what other, similar bits of code do.
"We were creating wiki of all these nasty things," says Rao. As they were working, they realized that DNA manufacturers could potentially scan in sequences that people ordered, run them against the database, and see if anything scary matched up. Kind of like that plagiarism software your college professors used.
Battelle began offering their screening capability, as ThreatSEQ. When customers -- like, currently, Twist Bioscience -- throw their sequences in, and get a report back, the manufacturers make the final decision about whether to fulfill a flagged order — whether, in the analogy, to give an F for plagiarism. After all, legitimate researchers do legitimately need to have DNA from legitimately bad organisms.
"Maybe it's the CDC," says Rao. "If things check out, oftentimes [the manufacturers] will fulfill the order." If it's your aggrieved uncle seeking the virulent pathogen, maybe not. But ultimately, no one is stopping the manufacturers from doing so.
Beyond that kind of tampering, though, cyberbiosecurity also includes keeping a lockdown on the machines that make the genetic sequences. "Somebody now doesn't need physical access to infrastructure to tamper with it," says Rao. So it needs the same cyber protections as other internet-connected devices.
Scientists are also now using DNA to store data — encoding information in its bases, rather than into a hard drive. To download the data, you sequence the DNA and read it back into a computer. But if you think like a bad guy, you'd realize that a bad guy could then, for instance, insert a computer virus into the genetic code, and when the researcher went to nab her data, her desktop would crash or infect the others on the network.
Something like that actually happened in 2017 at the USENIX security symposium, an annual programming conference: Researchers from the University of Washington encoded malware into DNA, and when the gene sequencer assembled the DNA, it corrupted the sequencer's software, then the computer that controlled it.
"This vulnerability could be just the opening an adversary needs to compromise an organization's systems," Inspirion Biosciences' J. Craig Reed and Nicolas Dunaway wrote in a paper for Frontiers in Bioengineering and Biotechnology, included in an e-book that Murch edited called Mapping the Cyberbiosecurity Enterprise.
Where We Go From Here
So what to do about all this? That's hard to say, in part because we don't know how big a current problem any of it poses. As noted in Mapping the Cyberbiosecurity Enterprise, "Information about private sector infrastructure vulnerabilities or data breaches is protected from public release by the Protected Critical Infrastructure Information (PCII) Program," if the privateers share the information with the government. "Government sector vulnerabilities or data breaches," meanwhile, "are rarely shared with the public."
"What I think is encouraging right now is the fact that we're even having this discussion."
The regulations that could rein in problems aren't as robust as many would like them to be, and much good behavior is technically voluntary — although guidelines and best practices do exist from organizations like the International Gene Synthesis Consortium and the National Institute of Standards and Technology.
Rao thinks it would be smart if grant-giving agencies like the National Institutes of Health and the National Science Foundation required any scientists who took their money to work with manufacturing companies that screen sequences. But he also still thinks we're on our way to being ahead of the curve, in terms of preventing print-your-own bioproblems: "What I think is encouraging right now is the fact that we're even having this discussion," says Rao.
Peccoud, for his part, has worked to keep such conversations going, including by doing training for the FBI and planning a workshop for students in which they imagine and work to guard against the malicious use of their research. But actually, Peccoud believes that human error, flawed lab processes, and mislabeled samples might be bigger threats than the outside ones. "Way too often, I think that people think of security as, 'Oh, there is a bad guy going after me,' and the main thing you should be worried about is yourself and errors," he says.
Murch thinks we're only at the beginning of understanding where our weak points are, and how many times they've been bruised. Decreasing those contusions, though, won't just take more secure systems. "The answer won't be technical only," he says. It'll be social, political, policy-related, and economic — a cultural revolution all its own.
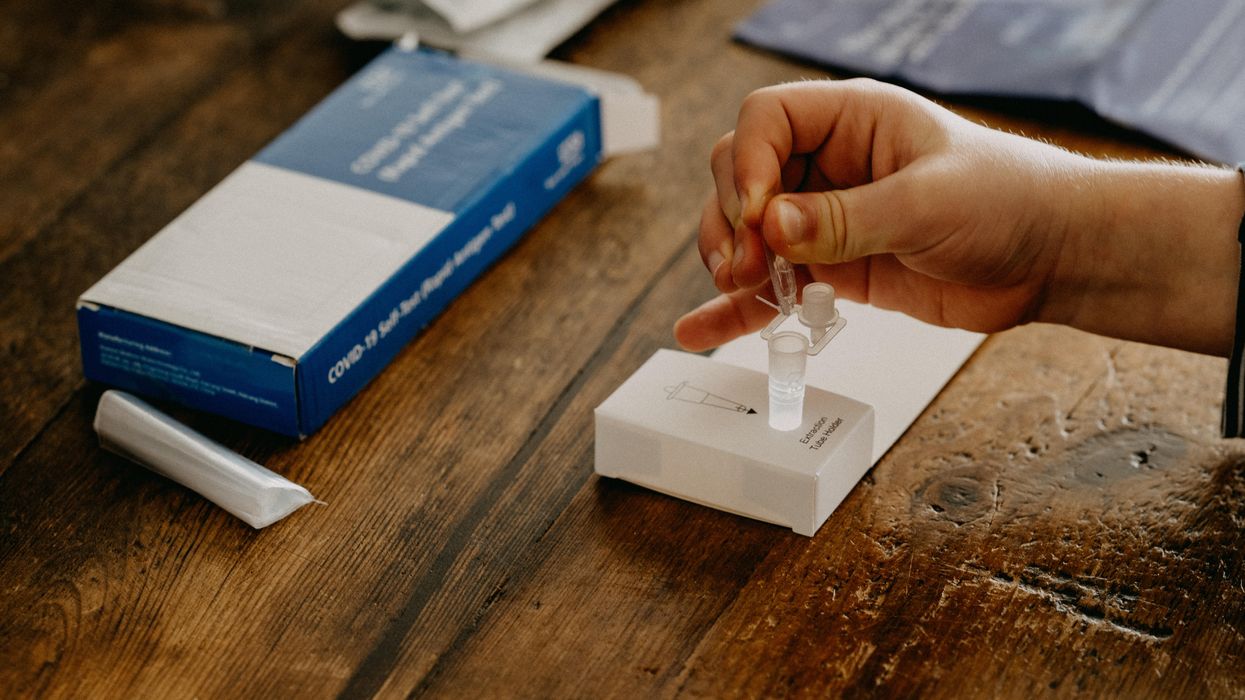
Biohackers Made a Cheap and Effective Home Covid Test -- But No One Is Allowed to Use It
A stock image of a home test for COVID-19.
Last summer, when fast and cheap Covid tests were in high demand and governments were struggling to manufacture and distribute them, a group of independent scientists working together had a bit of a breakthrough.
Working on the Just One Giant Lab platform, an online community that serves as a kind of clearing house for open science researchers to find each other and work together, they managed to create a simple, one-hour Covid test that anyone could take at home with just a cup of hot water. The group tested it across a network of home and professional laboratories before being listed as a semi-finalist team for the XPrize, a competition that rewards innovative solutions-based projects. Then, the group hit a wall: they couldn't commercialize the test.
They wanted to keep their project open source, making it accessible to people around the world, so they decided to forgo traditional means of intellectual property protection and didn't seek patents. (They couldn't afford lawyers anyway). And, as a loose-knit group that was not supported by a traditional scientific institution, working in community labs and homes around the world, they had no access to resources or financial support for manufacturing or distributing their test at scale.
But without ethical and regulatory approval for clinical testing, manufacture, and distribution, they were legally unable to create field tests for real people, leaving their inexpensive, $16-per-test, innovative product languishing behind, while other, more expensive over-the-counter tests made their way onto the market.
Who Are These Radical Scientists?
Independent, decentralized biomedical research has come of age. Also sometimes called DIYbio, biohacking, or community biology, depending on whom you ask, open research is today a global movement with thousands of members, from scientists with advanced degrees to middle-grade students. Their motivations and interests vary across a wide spectrum, but transparency and accessibility are key to the ethos of the movement. Teams are agile, focused on shoestring-budget R&D, and aim to disrupt business as usual in the ivory towers of the scientific establishment.
Ethics oversight is critical to ensuring that research is conducted responsibly, even by biohackers.
Initiatives developed within the community, such as Open Insulin, which hopes to engineer processes for affordable, small-batch insulin production, "Slybera," a provocative attempt to reverse engineer a $1 million dollar gene therapy, and the hundreds of projects posted on the collaboration platform Just One Giant Lab during the pandemic, all have one thing in common: to pursue testing in humans, they need an ethics oversight mechanism.
These groups, most of which operate collaboratively in community labs, homes, and online, recognize that some sort of oversight or guidance is useful—and that it's the right thing to do.
But also, and perhaps more immediately, they need it because federal rules require ethics oversight of any biomedical research that's headed in the direction of the consumer market. In addition, some individuals engaged in this work do want to publish their research in traditional scientific journals, which—you guessed it—also require that research has undergone an ethics evaluation. Ethics oversight is critical to ensuring that research is conducted responsibly, even by biohackers.
Bridging the Ethics Gap
The problem is that traditional oversight mechanisms, such as institutional review boards at government or academic research institutions, as well as the private boards utilized by pharmaceutical companies, are not accessible to most independent researchers. Traditional review boards are either closed to the public, or charge fees that are out of reach for many citizen science initiatives. This has created an "ethics gap" in nontraditional scientific research.
Biohackers are seen in some ways as the direct descendents of "white hat" computer hackers, or those focused on calling out security holes and contributing solutions to technical problems within self-regulating communities. In the case of health and biotechnology, those problems include both the absence of treatments and the availability of only expensive treatments for certain conditions. As the DIYbio community grows, there needs to be a way to provide assurance that, when the work is successful, the public is able to benefit from it eventually. The team that developed the one-hour Covid test found a potential commercial partner and so might well overcome the oversight hurdle, but it's been 14 months since they developed the test--and counting.
In short, without some kind of oversight mechanism for the work of independent biomedical researchers, the solutions they innovate will never have the opportunity to reach consumers.
In a new paper in the journal Citizen Science: Theory & Practice, we consider the issue of the ethics gap and ask whether ethics oversight is something nontraditional researchers want, and if so, what forms it might take. Given that individuals within these communities sometimes vehemently disagree with each other, is consensus on these questions even possible?
We learned that there is no "one size fits all" solution for ethics oversight of nontraditional research. Rather, the appropriateness of any oversight model will depend on each initiative's objectives, needs, risks, and constraints.
We also learned that nontraditional researchers are generally willing (and in some cases eager) to engage with traditional scientific, legal, and bioethics experts on ethics, safety, and related questions.
We suggest that these experts make themselves available to help nontraditional researchers build infrastructure for ethics self-governance and identify when it might be necessary to seek outside assistance.
Independent biomedical research has promise, but like any emerging science, it poses novel ethical questions and challenges. Existing research ethics and oversight frameworks may not be well-suited to answer them in every context, so we need to think outside the box about what we can create for the future. That process should begin by talking to independent biomedical researchers about their activities, priorities, and concerns with an eye to understanding how best to support them.
Christi Guerrini, JD, MPH studies biomedical citizen science and is an Associate Professor at Baylor College of Medicine. Alex Pearlman, MA, is a science journalist and bioethicist who writes about emerging issues in biotechnology. They have recently launched outlawbio.org, a place for discussion about nontraditional research.
Sept. 13th Event: Delta, Vaccines, and Breakthrough Infections
A visualization of the virus that causes COVID-19.
This virtual event will convene leading scientific and medical experts to address the public's questions and concerns about COVID-19 vaccines, Delta, and breakthrough infections. Audience Q&A will follow the panel discussion. Your questions can be submitted in advance at the registration link.
DATE:
Monday, September 13th, 2021
12:30 p.m. - 1:45 p.m. EDT
REGISTER:

Dr. Amesh Adalja, M.D., FIDSA, Senior Scholar, Johns Hopkins Center for Health Security; Adjunct Assistant Professor, Johns Hopkins Bloomberg School of Public Health; Affiliate of the Johns Hopkins Center for Global Health. His work is focused on emerging infectious disease, pandemic preparedness, and biosecurity.

Dr. Nahid Bhadelia, M.D., MALD, Founding Director, Boston University Center for Emerging Infectious Diseases Policy and Research (CEID); Associate Director, National Emerging Infectious Diseases Laboratories (NEIDL), Boston University; Associate Professor, Section of Infectious Diseases, Boston University School of Medicine. She is an internationally recognized leader in highly communicable and emerging infectious diseases (EIDs) with clinical, field, academic, and policy experience in pandemic preparedness.

Dr. Akiko Iwasaki, Ph.D., Waldemar Von Zedtwitz Professor of Immunobiology and Molecular, Cellular and Developmental Biology and Professor of Epidemiology (Microbial Diseases), Yale School of Medicine; Investigator, Howard Hughes Medical Institute. Her laboratory researches how innate recognition of viral infections lead to the generation of adaptive immunity, and how adaptive immunity mediates protection against subsequent viral challenge.
 Dr. Marion Pepper, Ph.D., Associate Professor, Department of Immunology, University of Washington. Her lab studies how cells of the adaptive immune system, called CD4+ T cells and B cells, form immunological memory by visualizing their differentiation, retention, and function.
Dr. Marion Pepper, Ph.D., Associate Professor, Department of Immunology, University of Washington. Her lab studies how cells of the adaptive immune system, called CD4+ T cells and B cells, form immunological memory by visualizing their differentiation, retention, and function.
This event is the third of a four-part series co-hosted by Leaps.org, the Aspen Institute Science & Society Program, and the Sabin–Aspen Vaccine Science & Policy Group, with generous support from the Gordon and Betty Moore Foundation and the Howard Hughes Medical Institute.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.