Meet the Scientists on the Frontlines of Protecting Humanity from a Man-Made Pathogen

From left: Jean Peccoud, Randall Murch, and Neeraj Rao.
Jean Peccoud wasn't expecting an email from the FBI. He definitely wasn't expecting the agency to invite him to a meeting. "My reaction was, 'What did I do wrong to be on the FBI watch list?'" he recalls.
You use those blueprints for white-hat research—which is, indeed, why the open blueprints exist—or you can do the same for a black-hat attack.
He didn't know what the feds could possibly want from him. "I was mostly scared at this point," he says. "I was deeply disturbed by the whole thing."
But he decided to go anyway, and when he traveled to San Francisco for the 2008 gathering, the reason for the e-vite became clear: The FBI was reaching out to researchers like him—scientists interested in synthetic biology—in anticipation of the potential nefarious uses of this technology. "The whole purpose of the meeting was, 'Let's start talking to each other before we actually need to talk to each other,'" says Peccoud, now a professor of chemical and biological engineering at Colorado State University. "'And let's make sure next time you get an email from the FBI, you don't freak out."
Synthetic biology—which Peccoud defines as "the application of engineering methods to biological systems"—holds great power, and with that (as always) comes great responsibility. When you can synthesize genetic material in a lab, you can create new ways of diagnosing and treating people, and even new food ingredients. But you can also "print" the genetic sequence of a virus or virulent bacterium.
And while it's not easy, it's also not as hard as it could be, in part because dangerous sequences have publicly available blueprints. You use those blueprints for white-hat research—which is, indeed, why the open blueprints exist—or you can do the same for a black-hat attack. You could synthesize a dangerous pathogen's code on purpose, or you could unwittingly do so because someone tampered with your digital instructions. Ordering synthetic genes for viral sequences, says Peccoud, would likely be more difficult today than it was a decade ago.
"There is more awareness of the industry, and they are taking this more seriously," he says. "There is no specific regulation, though."
Trying to lock down the interconnected machines that enable synthetic biology, secure its lab processes, and keep dangerous pathogens out of the hands of bad actors is part of a relatively new field: cyberbiosecurity, whose name Peccoud and colleagues introduced in a 2018 paper.
Biological threats feel especially acute right now, during the ongoing pandemic. COVID-19 is a natural pathogen -- not one engineered in a lab. But future outbreaks could start from a bug nature didn't build, if the wrong people get ahold of the right genetic sequences, and put them in the right sequence. Securing the equipment and processes that make synthetic biology possible -- so that doesn't happen -- is part of why the field of cyberbiosecurity was born.
The Origin Story
It is perhaps no coincidence that the FBI pinged Peccoud when it did: soon after a journalist ordered a sequence of smallpox DNA and wrote, for The Guardian, about how easy it was. "That was not good press for anybody," says Peccoud. Previously, in 2002, the Pentagon had funded SUNY Stonybrook researchers to try something similar: They ordered bits of polio DNA piecemeal and, over the course of three years, strung them together.
Although many years have passed since those early gotchas, the current patchwork of regulations still wouldn't necessarily prevent someone from pulling similar tricks now, and the technological systems that synthetic biology runs on are more intertwined — and so perhaps more hackable — than ever. Researchers like Peccoud are working to bring awareness to those potential problems, to promote accountability, and to provide early-detection tools that would catch the whiff of a rotten act before it became one.
Peccoud notes that if someone wants to get access to a specific pathogen, it is probably easier to collect it from the environment or take it from a biodefense lab than to whip it up synthetically. "However, people could use genetic databases to design a system that combines different genes in a way that would make them dangerous together without each of the components being dangerous on its own," he says. "This would be much more difficult to detect."
After his meeting with the FBI, Peccoud grew more interested in these sorts of security questions. So he was paying attention when, in 2010, the Department of Health and Human Services — now helping manage the response to COVID-19 — created guidance for how to screen synthetic biology orders, to make sure suppliers didn't accidentally send bad actors the sequences that make up bad genomes.
Guidance is nice, Peccoud thought, but it's just words. He wanted to turn those words into action: into a computer program. "I didn't know if it was something you can run on a desktop or if you need a supercomputer to run it," he says. So, one summer, he tasked a team of student researchers with poring over the sentences and turning them into scripts. "I let the FBI know," he says, having both learned his lesson and wanting to get in on the game.
Peccoud later joined forces with Randall Murch, a former FBI agent and current Virginia Tech professor, and a team of colleagues from both Virginia Tech and the University of Nebraska-Lincoln, on a prototype project for the Department of Defense. They went into a lab at the University of Nebraska at Lincoln and assessed all its cyberbio-vulnerabilities. The lab develops and produces prototype vaccines, therapeutics, and prophylactic components — exactly the kind of place that you always, and especially right now, want to keep secure.
"We were creating wiki of all these nasty things."
The team found dozens of Achilles' heels, and put them in a private report. Not long after that project, the two and their colleagues wrote the paper that first used the term "cyberbiosecurity." A second paper, led by Murch, came out five months later and provided a proposed definition and more comprehensive perspective on cyberbiosecurity. But although it's now a buzzword, it's the definition, not the jargon, that matters. "Frankly, I don't really care if they call it cyberbiosecurity," says Murch. Call it what you want: Just pay attention to its tenets.
A Database of Scary Sequences
Peccoud and Murch, of course, aren't the only ones working to screen sequences and secure devices. At the nonprofit Battelle Memorial Institute in Columbus, Ohio, for instance, scientists are working on solutions that balance the openness inherent to science and the closure that can stop bad stuff. "There's a challenge there that you want to enable research but you want to make sure that what people are ordering is safe," says the organization's Neeraj Rao.
Rao can't talk about the work Battelle does for the spy agency IARPA, the Intelligence Advanced Research Projects Activity, on a project called Fun GCAT, which aims to use computational tools to deep-screen gene-sequence orders to see if they pose a threat. It can, though, talk about a twin-type internal project: ThreatSEQ (pronounced, of course, "threat seek").
The project started when "a government customer" (as usual, no one will say which) asked Battelle to curate a list of dangerous toxins and pathogens, and their genetic sequences. The researchers even started tagging sequences according to their function — like whether a particular sequence is involved in a germ's virulence or toxicity. That helps if someone is trying to use synthetic biology not to gin up a yawn-inducing old bug but to engineer a totally new one. "How do you essentially predict what the function of a novel sequence is?" says Rao. You look at what other, similar bits of code do.
"We were creating wiki of all these nasty things," says Rao. As they were working, they realized that DNA manufacturers could potentially scan in sequences that people ordered, run them against the database, and see if anything scary matched up. Kind of like that plagiarism software your college professors used.
Battelle began offering their screening capability, as ThreatSEQ. When customers -- like, currently, Twist Bioscience -- throw their sequences in, and get a report back, the manufacturers make the final decision about whether to fulfill a flagged order — whether, in the analogy, to give an F for plagiarism. After all, legitimate researchers do legitimately need to have DNA from legitimately bad organisms.
"Maybe it's the CDC," says Rao. "If things check out, oftentimes [the manufacturers] will fulfill the order." If it's your aggrieved uncle seeking the virulent pathogen, maybe not. But ultimately, no one is stopping the manufacturers from doing so.
Beyond that kind of tampering, though, cyberbiosecurity also includes keeping a lockdown on the machines that make the genetic sequences. "Somebody now doesn't need physical access to infrastructure to tamper with it," says Rao. So it needs the same cyber protections as other internet-connected devices.
Scientists are also now using DNA to store data — encoding information in its bases, rather than into a hard drive. To download the data, you sequence the DNA and read it back into a computer. But if you think like a bad guy, you'd realize that a bad guy could then, for instance, insert a computer virus into the genetic code, and when the researcher went to nab her data, her desktop would crash or infect the others on the network.
Something like that actually happened in 2017 at the USENIX security symposium, an annual programming conference: Researchers from the University of Washington encoded malware into DNA, and when the gene sequencer assembled the DNA, it corrupted the sequencer's software, then the computer that controlled it.
"This vulnerability could be just the opening an adversary needs to compromise an organization's systems," Inspirion Biosciences' J. Craig Reed and Nicolas Dunaway wrote in a paper for Frontiers in Bioengineering and Biotechnology, included in an e-book that Murch edited called Mapping the Cyberbiosecurity Enterprise.
Where We Go From Here
So what to do about all this? That's hard to say, in part because we don't know how big a current problem any of it poses. As noted in Mapping the Cyberbiosecurity Enterprise, "Information about private sector infrastructure vulnerabilities or data breaches is protected from public release by the Protected Critical Infrastructure Information (PCII) Program," if the privateers share the information with the government. "Government sector vulnerabilities or data breaches," meanwhile, "are rarely shared with the public."
"What I think is encouraging right now is the fact that we're even having this discussion."
The regulations that could rein in problems aren't as robust as many would like them to be, and much good behavior is technically voluntary — although guidelines and best practices do exist from organizations like the International Gene Synthesis Consortium and the National Institute of Standards and Technology.
Rao thinks it would be smart if grant-giving agencies like the National Institutes of Health and the National Science Foundation required any scientists who took their money to work with manufacturing companies that screen sequences. But he also still thinks we're on our way to being ahead of the curve, in terms of preventing print-your-own bioproblems: "What I think is encouraging right now is the fact that we're even having this discussion," says Rao.
Peccoud, for his part, has worked to keep such conversations going, including by doing training for the FBI and planning a workshop for students in which they imagine and work to guard against the malicious use of their research. But actually, Peccoud believes that human error, flawed lab processes, and mislabeled samples might be bigger threats than the outside ones. "Way too often, I think that people think of security as, 'Oh, there is a bad guy going after me,' and the main thing you should be worried about is yourself and errors," he says.
Murch thinks we're only at the beginning of understanding where our weak points are, and how many times they've been bruised. Decreasing those contusions, though, won't just take more secure systems. "The answer won't be technical only," he says. It'll be social, political, policy-related, and economic — a cultural revolution all its own.
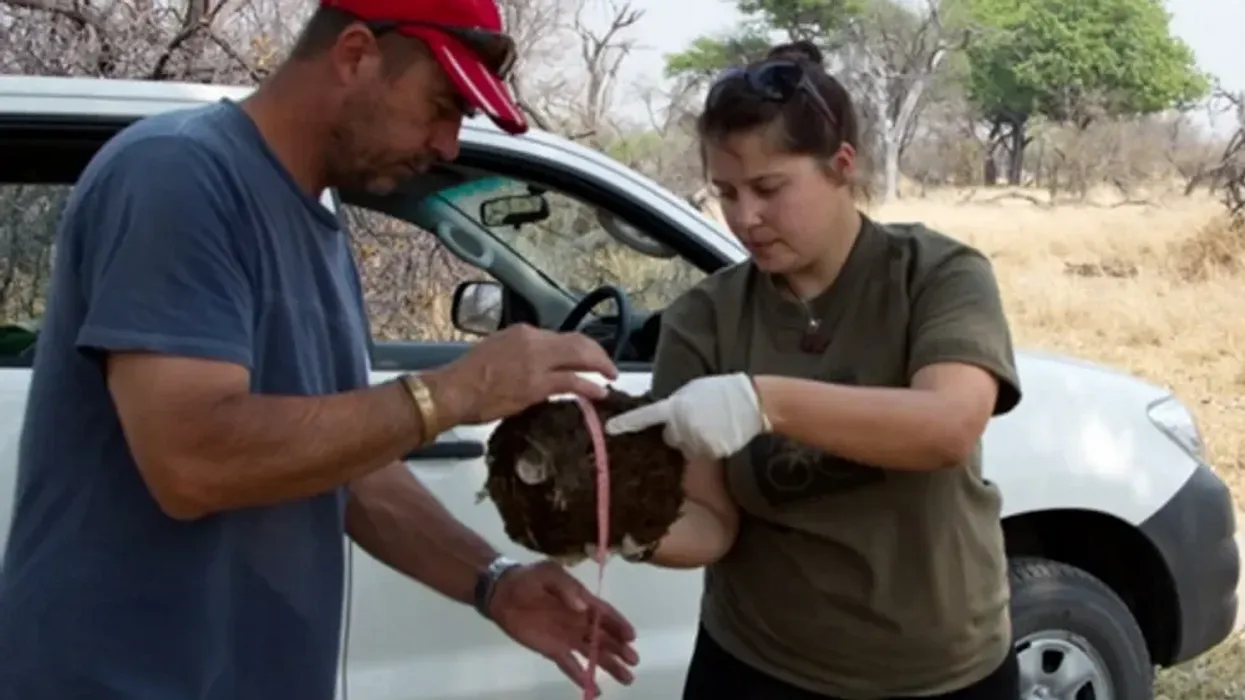
DNA gathered from animal poop helps protect wildlife
Alida de Flamingh and her team are collecting elephant dung. It holds a trove of information about animal health, diet and genetic diversity.
On the savannah near the Botswana-Zimbabwe border, elephants grazed contentedly. Nearby, postdoctoral researcher Alida de Flamingh watched and waited. As the herd moved away, she went into action, collecting samples of elephant dung that she and other wildlife conservationists would study in the months to come. She pulled on gloves, took a swab, and ran it all over the still-warm, round blob of elephant poop.
Sequencing DNA from fecal matter is a safe, non-invasive way to track and ultimately help protect over 42,000 species currently threatened by extinction. Scientists are using this DNA to gain insights into wildlife health, genetic diversity and even the broader environment. Applied to elephants, chimpanzees, toucans and other species, it helps scientists determine the genetic diversity of groups and linkages with other groups. Such analysis can show changes in rates of inbreeding. Populations with greater genetic diversity adapt better to changes and environmental stressors than those with less diversity, thus reducing their risks of extinction, explains de Flamingh, a postdoctoral researcher at the University of Illinois Urbana-Champaign.
Analyzing fecal DNA also reveals information about an animal’s diet and health, and even nearby flora that is eaten. That information gives scientists broader insights into the ecosystem, and the findings are informing conservation initiatives. Examples include restoring or maintaining genetic connections among groups, ensuring access to certain foraging areas or increasing diversity in captive breeding programs.
Approximately 27 percent of mammals and 28 percent of all assessed species are close to dying out. The IUCN Red List of threatened species, simply called the Red List, is the world’s most comprehensive record of animals’ risk of extinction status. The more information scientists gather, the better their chances of reducing those risks. In Africa, populations of vertebrates declined 69 percent between 1970 and 2022, according to the World Wildlife Fund (WWF).
“We put on sterile gloves and use a sterile swab to collect wet mucus and materials from the outside of the dung ball,” says Alida de Flamingh, a postdoctoral researcher at the University of Illinois Urbana-Champaign.
“When people talk about species, they often talk about ecosystems, but they often overlook genetic diversity,” says Christina Hvilsom, senior geneticist at the Copenhagen Zoo. “It’s easy to count (individuals) to assess whether the population size is increasing or decreasing, but diversity isn’t something we can see with our bare eyes. Yet, it’s actually the foundation for the species and populations.” DNA analysis can provide this critical information.
Assessing elephants’ health
“Africa’s elephant populations are facing unprecedented threats,” says de Flamingh, the postdoc, who has studied them since 2009. Challenges include ivory poaching, habitat destruction and smaller, more fragmented habitats that result in smaller mating pools with less genetic diversity. Additionally, de Flamingh studies the microbial communities living on and in elephants – their microbiomes – looking for parasites or dangerous microbes.
Approximately 415,000 elephants inhabit Africa today, but de Flamingh says the number would be four times higher without these challenges. The IUCN Red List reports African savannah elephants are endangered and African forest elephants are critically endangered. Elephants support ecosystem biodiversity by clearing paths that help other species travel. Their very footprints create small puddles that can host smaller organisms such as tadpoles. Elephants are often described as ecosystems’ engineers, so if they disappear, the rest of the ecosystem will suffer too.
There’s a process to collecting elephant feces. “We put on sterile gloves (which we change for each sample) and use a sterile swab to collect wet mucus and materials from the outside of the dung ball,” says de Flamingh. They rub a sample about the size of a U.S. quarter onto a paper card embedded with DNA preservation technology. Each card is air dried and stored in a packet of desiccant to prevent mold growth. This way, samples can be stored at room temperature indefinitely without the DNA degrading.
Earlier methods required collecting dung in bags, which needed either refrigeration or the addition of preservatives, or the riskier alternative of tranquilizing the animals before approaching them to draw blood samples. The ability to collect and sequence the DNA made things much easier and safer.
“Our research provides a way to assess elephant health without having to physically interact with elephants,” de Flamingh emphasizes. “We also keep track of the GPS coordinates of each sample so that we can create a map of the sampling locations,” she adds. That helps researchers correlate elephants’ health with geographic areas and their conditions.
Although de Flamingh works with elephants in the wild, the contributions of zoos in the United States and collaborations in South Africa (notably the late Professor Rudi van Aarde and the Conservation Ecology Research Unit at the University of Pretoria) were key in studying this method to ensure it worked, she points out.
Protecting chimpanzees
Genetic work with chimpanzees began about a decade ago. Hvilsom and her group at the Copenhagen Zoo analyzed DNA from nearly 1,000 fecal samples collected between 2003 and 2018 by a team of international researchers. The goal was to assess the status of the West African subspecies, which is critically endangered after rapid population declines. Of the four subspecies of chimpanzees, the West African subspecies is considered the most at-risk.
In total, the WWF estimates the numbers of chimpanzees inhabiting Africa’s forests and savannah woodlands at between 173,000 and 300,000. Poaching, disease and human-caused changes to their lands are their major risks.
By analyzing genetics obtained from fecal samples, Hvilsom estimated the chimpanzees’ population, ascertained their family relationships and mapped their migration routes.
“One of the threats is mining near the Nimba Mountains in Guinea,” a stronghold for the West African subspecies, Hvilsom says. The Nimba Mountains are a UNESCO World Heritage Site, but they are rich in iron ore, which is used to make the steel that is vital to the Asian construction boom. As she and colleagues wrote in a recent paper, “Many extractive industries are currently developing projects in chimpanzee habitat.”
Analyzing DNA allows researchers to identify individual chimpanzees more accurately than simply observing them, she says. Normally, field researchers would install cameras and manually inspect each picture to determine how many chimpanzees were in an area. But, Hvilsom says, “That’s very tricky. Chimpanzees move a lot and are fast, so it’s difficult to get clear pictures. Often, they find and destroy the cameras. Also, they live in large areas, so you need a lot of cameras.”
By analyzing genetics obtained from fecal samples, Hvilsom estimated the chimpanzees’ population, ascertained their family relationships and mapped their migration routes based upon DNA comparisons with other chimpanzee groups. The mining companies and builders are using this information to locate future roads where they won’t disrupt migration – a more effective solution than trying to build artificial corridors for wildlife.
“The current route cuts off communities of chimpanzees,” Hvilsom elaborates. That effectively prevents young adult chimps from joining other groups when the time comes, eventually reducing the currently-high levels of genetic diversity.
“The mining company helped pay for the genetics work,” Hvilsom says, “as part of its obligation to assess and monitor biodiversity and the effect of the mining in the area.”
Of 50 toucan subspecies, 11 are threatened or near-threatened with extinction because of deforestation and poaching.
Identifying toucan families
Feces aren't the only substance researchers draw DNA samples from. Jeffrey Coleman, a Ph.D. candidate at the University of Texas at Austin relies on blood tests for studying the genetic diversity of toucans---birds species native to Central America and nearby regions. They live in the jungles, where they hop among branches, snip fruit from trees, toss it in the air and catch it with their large beaks. “Toucans are beautiful, charismatic birds that are really important to the ecosystem,” says Coleman.
Of their 50 subspecies, 11 are threatened or near-threatened with extinction because of deforestation and poaching. “When people see these aesthetically pleasing birds, they’re motivated to care about conservation practices,” he points out.
Coleman works with the Dallas World Aquarium and its partner zoos to analyze DNA from blood draws, using it to identify which toucans are related and how closely. His goal is to use science to improve the genetic diversity among toucan offspring.
Specifically, he’s looking at sections of the genome of captive birds in which the nucleotides repeat multiple times, such as AGATAGATAGAT. Called microsatellites, these consecutively-repeating sections can be passed from parents to children, helping scientists identify parent-child and sibling-sibling relationships. “That allows you to make strategic decisions about how to pair (captive) individuals for mating...to avoid inbreeding,” Coleman says.

Jeffrey Coleman is studying the microsatellites inside the toucan genomes.
Courtesy Jeffrey Coleman
The alternative is to use a type of analysis that looks for a single DNA building block – a nucleotide – that differs in a given sequence. Called single nucleotide polymorphisms (SNPs, pronounced “snips”), they are very common and very accurate. Coleman says they are better than microsatellites for some uses. But scientists have already developed a large body of microsatellite data from multiple species, so microsatellites can shed more insights on relations.
Regardless of whether conservation programs use SNPs or microsatellites to guide captive breeding efforts, the goal is to help them build genetically diverse populations that eventually may supplement endangered populations in the wild. “The hope is that the ecosystem will be stable enough and that the populations (once reintroduced into the wild) will be able to survive and thrive,” says Coleman. History knows some good examples of captive breeding success.
The California condor, which had a total population of 27 in 1987, when the last wild birds were captured, is one of them. A captive breeding program boosted their numbers to 561 by the end of 2022. Of those, 347 of those are in the wild, according to the National Park Service.
Conservationists hope that their work on animals’ genetic diversity will help preserve and restore endangered species in captivity and the wild. DNA analysis is crucial to both types of efforts. The ability to apply genome sequencing to wildlife conservation brings a new level of accuracy that helps protect species and gives fresh insights that observation alone can’t provide.
“A lot of species are threatened,” Coleman says. “I hope this research will be a resource people can use to get more information on longer-term genealogies and different populations.”
DNA- and RNA-based electronic implants may revolutionize healthcare
The test tubes contain tiny DNA/enzyme-based circuits, which comprise TRUMPET, a new type of electronic device, smaller than a cell.
Implantable electronic devices can significantly improve patients’ quality of life. A pacemaker can encourage the heart to beat more regularly. A neural implant, usually placed at the back of the skull, can help brain function and encourage higher neural activity. Current research on neural implants finds them helpful to patients with Parkinson’s disease, vision loss, hearing loss, and other nerve damage problems. Several of these implants, such as Elon Musk’s Neuralink, have already been approved by the FDA for human use.
Yet, pacemakers, neural implants, and other such electronic devices are not without problems. They require constant electricity, limited through batteries that need replacements. They also cause scarring. “The problem with doing this with electronics is that scar tissue forms,” explains Kate Adamala, an assistant professor of cell biology at the University of Minnesota Twin Cities. “Anytime you have something hard interacting with something soft [like muscle, skin, or tissue], the soft thing will scar. That's why there are no long-term neural implants right now.” To overcome these challenges, scientists are turning to biocomputing processes that use organic materials like DNA and RNA. Other promised benefits include “diagnostics and possibly therapeutic action, operating as nanorobots in living organisms,” writes Evgeny Katz, a professor of bioelectronics at Clarkson University, in his book DNA- And RNA-Based Computing Systems.
While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output.
Adamala’s research focuses on developing such biocomputing systems using DNA, RNA, proteins, and lipids. Using these molecules in the biocomputing systems allows the latter to be biocompatible with the human body, resulting in a natural healing process. In a recent Nature Communications study, Adamala and her team created a new biocomputing platform called TRUMPET (Transcriptional RNA Universal Multi-Purpose GatE PlaTform) which acts like a DNA-powered computer chip. “These biological systems can heal if you design them correctly,” adds Adamala. “So you can imagine a computer that will eventually heal itself.”
The basics of biocomputing
Biocomputing and regular computing have many similarities. Like regular computing, biocomputing works by running information through a series of gates, usually logic gates. A logic gate works as a fork in the road for an electronic circuit. The input will travel one way or another, giving two different outputs. An example logic gate is the AND gate, which has two inputs (A and B) and two different results. If both A and B are 1, the AND gate output will be 1. If only A is 1 and B is 0, the output will be 0 and vice versa. If both A and B are 0, the result will be 0. While a computer gives these inputs in binary code or "bits," such as a 0 or 1, biocomputing uses DNA strands as inputs, whether double or single-stranded, and often uses fluorescent RNA as an output. In this case, the DNA enters the logic gate as a single or double strand.
If the DNA is double-stranded, the system “digests” the DNA or destroys it, which results in non-fluorescence or “0” output. Conversely, if the DNA is single-stranded, it won’t be digested and instead will be copied by several enzymes in the biocomputing system, resulting in fluorescent RNA or a “1” output. And the output for this type of binary system can be expanded beyond fluorescence or not. For example, a “1” output might be the production of the enzyme insulin, while a “0” may be that no insulin is produced. “This kind of synergy between biology and computation is the essence of biocomputing,” says Stephanie Forrest, a professor and the director of the Biodesign Center for Biocomputing, Security and Society at Arizona State University.
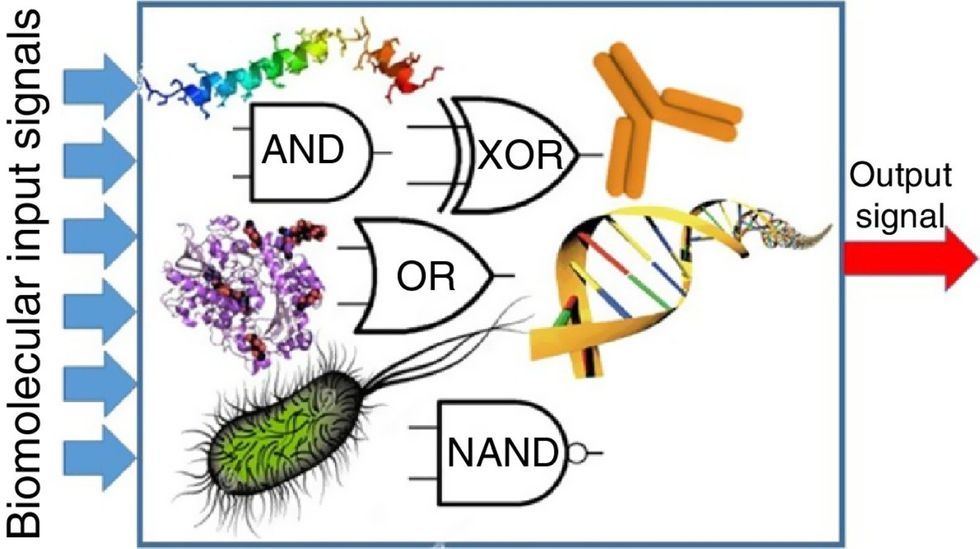
Biocomputing circles are made of DNA, RNA, proteins and even bacteria.
Evgeny Katz
The TRUMPET’s promise
Depending on whether the biocomputing system is placed directly inside a cell within the human body, or run in a test-tube, different environmental factors play a role. When an output is produced inside a cell, the cell's natural processes can amplify this output (for example, a specific protein or DNA strand), creating a solid signal. However, these cells can also be very leaky. “You want the cells to do the thing you ask them to do before they finish whatever their businesses, which is to grow, replicate, metabolize,” Adamala explains. “However, often the gate may be triggered without the right inputs, creating a false positive signal. So that's why natural logic gates are often leaky." While biocomputing outside a cell in a test tube can allow for tighter control over the logic gates, the outputs or signals cannot be amplified by a cell and are less potent.
TRUMPET, which is smaller than a cell, taps into both cellular and non-cellular biocomputing benefits. “At its core, it is a nonliving logic gate system,” Adamala states, “It's a DNA-based logic gate system. But because we use enzymes, and the readout is enzymatic [where an enzyme replicates the fluorescent RNA], we end up with signal amplification." This readout means that the output from the TRUMPET system, a fluorescent RNA strand, can be replicated by nearby enzymes in the platform, making the light signal stronger. "So it combines the best of both worlds,” Adamala adds.
These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body.
The TRUMPET biocomputing process is relatively straightforward. “If the DNA [input] shows up as single-stranded, it will not be digested [by the logic gate], and you get this nice fluorescent output as the RNA is made from the single-stranded DNA, and that's a 1,” Adamala explains. "And if the DNA input is double-stranded, it gets digested by the enzymes in the logic gate, and there is no RNA created from the DNA, so there is no fluorescence, and the output is 0." On the story's leading image above, if the tube is "lit" with a purple color, that is a binary 1 signal for computing. If it's "off" it is a 0.
While still in research, TRUMPET and other biocomputing systems promise significant benefits to personalized healthcare and medicine. These organic-based systems could detect cancer cells or low insulin levels inside a patient’s body. The study’s lead author and graduate student Judee Sharon is already beginning to research TRUMPET's ability for earlier cancer diagnoses. Because the inputs for TRUMPET are single or double-stranded DNA, any mutated or cancerous DNA could theoretically be detected from the platform through the biocomputing process. Theoretically, devices like TRUMPET could be used to detect cancer and other diseases earlier.
Adamala sees TRUMPET not only as a detection system but also as a potential cancer drug delivery system. “Ideally, you would like the drug only to turn on when it senses the presence of a cancer cell. And that's how we use the logic gates, which work in response to inputs like cancerous DNA. Then the output can be the production of a small molecule or the release of a small molecule that can then go and kill what needs killing, in this case, a cancer cell. So we would like to develop applications that use this technology to control the logic gate response of a drug’s delivery to a cell.”
Although platforms like TRUMPET are making progress, a lot more work must be done before they can be used commercially. “The process of translating mechanisms and architecture from biology to computing and vice versa is still an art rather than a science,” says Forrest. “It requires deep computer science and biology knowledge,” she adds. “Some people have compared interdisciplinary science to fusion restaurants—not all combinations are successful, but when they are, the results are remarkable.”